t(1;1)(p36;q21) in Non Hodgkin Lymphoma
2006-06-01 Valia S Lestou Affiliation1.The Hospital for Sick Kids, Dept. Paediatric Laboratory Medicine (DPLM), 555 University Ave, Toronto, ON M5G 1X8, Canada
Clinics and Pathology
Disease
Non Hodgkin Lymphoma (NHL). Aberrations of chromosomal bands 1p36 and 1q11-q23 are among the most common chromosomal alterations in NHL.
Phenotype stem cell origin
Lymphocytes (B-cell and T-cell).
Etiology
The exact etiology of NHL is still unknown, risk increases with exposed to ionizing radiation, chemicals such as pesticides or solvents, Epstein-Barr Virus infection, family history of NHL (although no hereditary pattern has been established, Human Immunodeficiency Virus (HIV) infection, immunosuppression or immunodeficiency, genetics.
Epidemiology
NHL is the 5th most frequently diagnosed cancer overall for both males and females, males are slightly more often affected than females, increasing over time.
Clinics
At diagnosis, painful swelling of lymph nodes located in the neck, underarm and groin, unexplained fever, night sweats, constant fatigue, unexplained weight loss, itchy skin.
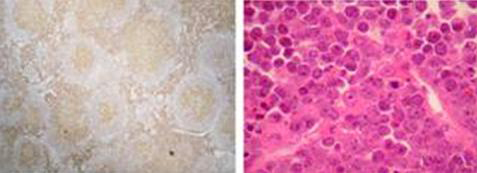
Transformed follicular lymphoma (courtesy, Dr. R.D. Gascoyne, BC Cancer Agency, Vancouver, Canada).
Cytology
Anti-B-cell antibodies (e.g. CD19, CD20, CD10, CD23); anti-T-cell antibodies (e.g. CD3, CD4, CD2/HLADR); other antibodies (e.g. CD45 for total lymphocytes, CD10 for monocytes).
Pathology
t(1;1)(p36;q21) has been seen in following NHL types as characterized by pathology; follicular lymphoma (FL) grades 1-3; diffuse large B-cell lymphoma; T-cell lymphoma and peripheral T-cell lymphoma.
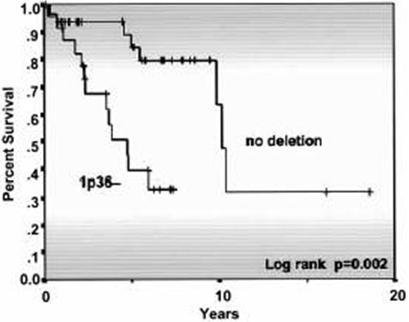
Univariate analyses using the Kaplan-Meier method for 1p36-, demonstrating the significance of this chromosomal change for overall survival. In multivariate analysis using the Cox regression model controlling for IPI, the significance remained intact.
Treatment
Depend on the stage and type and genetics of NHL; "watch-and-wait" approach in case of indolent follicular lymphomas; radiotherapy to site of problem; systemic chemotherapy; oral agents; IV agents; antibody against CD20; stem cell or bone marrow transplant.
Evolution
Initial genomic aberration (such as t(14;18)(q32;q21) in follicular lymphoma) may or may not be sufficient for the initiation of the malignant phenotype. Additional genomic rearrangements are required for disease progression.
Prognosis
Depend on the stage, type and genetics of NHL; in general, highly treatable and some times curable. However, a number of karyotype parameters have been reported to influence prognosis in NHL. It has been demonstrated that the cytogenetic abnormality 1p36-, as a result of t(1;1)(p36;q21) or another rearrangement involving chromosome 1, was found to be a significant predictors of adverse overall survival for FL (univariate and multivariate analysis).
Note
Initial cytogenetic changes often seen in e.g. FL and t(14;18)(q32;q21) (/BCL2); DLBL and t(3;14)(q27;q21)(BCL6/IGH). Additional acquired mutations are necessary to generate a fully malignant clonal proliferation. Many of these secondary genetic alterations (including chromosome 1) are visible in the clonal karyotype; it is now possible to identify the sequence by which they arise and their influence on clinical behavior by using computational methods to manipulate complex chromosomal data in large number of cases.
Cytogenetics
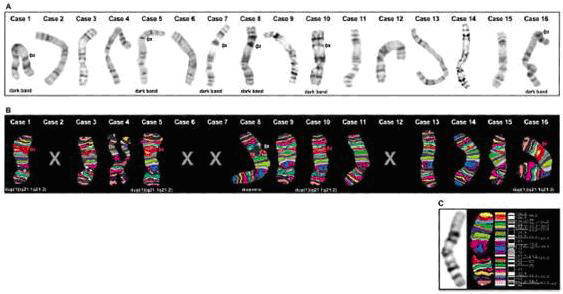
A: The derivative chromosome 1 in all 16 NHL cases as seen by G-band analysis. Arrows indicate the additional unidentified dark band.
B: The corresponding derivative chromosome 1 as seen by M-BAND1 analysis. Arrows indicate the dup(1)(q21.1q21.2) at the p/q-arm interface (broad orange/ pink bands). X indicates cases where no material was available for M-BAND1 analysis.
C: Normal chromosome 1 as seen by G-band and M-BAND1 analysis, color classifier, and ISCN 550-band level ideogram.
Cytogenetics morphological
Normal chromosome 1 with derivative chromosomes 1; Breakpoints are at chromosomal positions 1p36.3 and 1q21.1-2; duplication of the 1q21 to 1q44; adverse prognosis (?as a result of 1p36 suppressor genes deletions and/or duplication of 1q21q44 oncogenes); additional secondary abnormalities to t(1;1) of various complexity as usually seen in NHL.
Cytogenetics molecular
LS-FISH identification of 1p36.3 and 1q21.1-2 breakpoints on der(1)t(1;1); Two distinct types of 1p36.3 rearrangements were observed: One type involved deletions of SKI, MEL1, and TP73, and retained CASP9 the other type showed breakpoints telomeric to TP73; Four distinct types of 1q21.1-2 rearrangements were observed: The first type involved breakpoints at IRTA1 and IRTA2 with duplications of IRTA1, IRTA2, BCL9, AF1Q, JTB, and MUC1; the second type involved a breakpoint at BCL9 with duplications of BCL9, AF1Q, JTB, and MUC1; the third type involved a breakpoint at AF1Q with duplications of AF1Q, JTB, and MUC1; the fourth type involved an undefined breakpoint telomeric to MUC1.
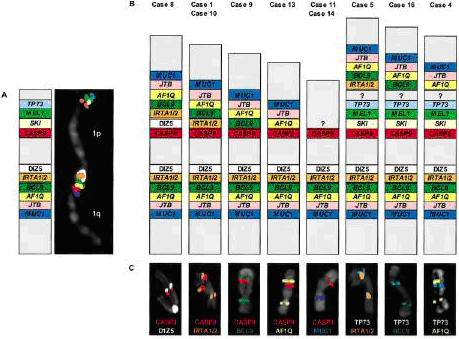
Composite picture of all LS-FISH patterns observed in this study with representative examples.
A: Normal color-coded chromosome 1 LS-FISH pattern, demonstrating the relative localization of all BAC probes.
B: All der(1)t(1;1) combinations seen by LS-FISH.
C: Two color-coded representative images corresponding to B and demonstrating the p/q-arm breakpoint interfaces.
Genes Involved and Proteins
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 14732921 | 2004 | Identification of cytogenetic subgroups and karyotypic pathways of clonal evolution in follicular lymphomas. | Höglund M et al |
| 12580956 | 2003 | Follicular lymphoma lacking the t(14;18)(q32;q21): identification of two disease subtypes. | Horsman DE et al |
| 12930384 | 2003 | Multicolour fluorescence in situ hybridization analysis of t(14;18)-positive follicular lymphoma and correlation with gene expression data and clinical outcome. | Lestou VS et al |
| 12619161 | 2003 | Characterization of the recurrent translocation t(1;1)(p36.3;q21.1-2) in non-Hodgkin lymphoma by multicolor banding and fluorescence in situ hybridization analysis. | Lestou VS et al |
| 15545733 | 2005 | New insights into the evolution of chromosome 1. | Weise A et al |
Summary
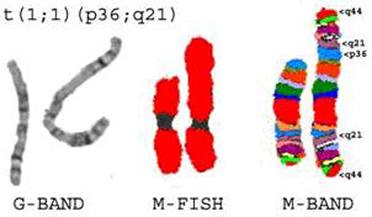
.Unbalanced t(1;1)(p36;q21) in NHL with dup(q21q44) as observed by, G-band, M-FISH and M-BAND [ISCN2005: der(1)t(p36.3;q21.1-2)dup(1)(q21.1-2q44)].
Citation
Valia S Lestou
t(1;1)(p36;q21) in Non Hodgkin Lymphoma
Atlas Genet Cytogenet Oncol Haematol. 2006-06-01
Online version: http://atlasgeneticsoncology.org/haematological/1431/t(1;1)(p36;q21)-in-non-hodgkin-lymphoma
