t(1;11)(q24;p15) NUP98/PRRX1
2018-01-01 Adriana Zamecnikova Affiliation1.Kuwait Cancer Control Center, Kuwait [email protected]
2.Genetics, Dept Medical Information, University of Poitiers; CHU Poitiers Hospital, F-86021 Poitiers, France
Abstract
NUP98 is considered as one of the most promiscuous genes in hematologic malignancies due to its participation in chromosomal translocations with up to 30 different partner genes, including homeodomain transcription factors.
Clinics and Pathology
Disease
Etiology
Epidemiology
Evolution
Cytogenetics
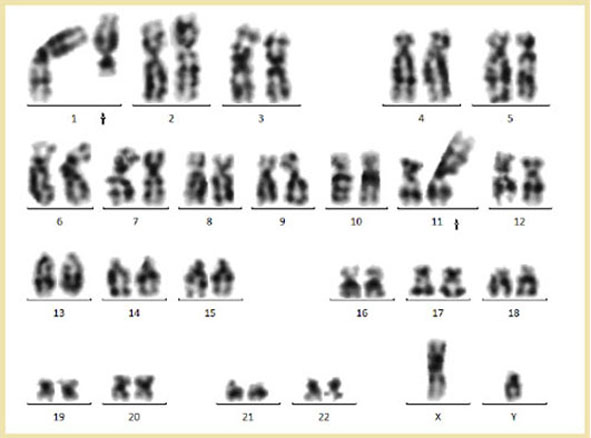
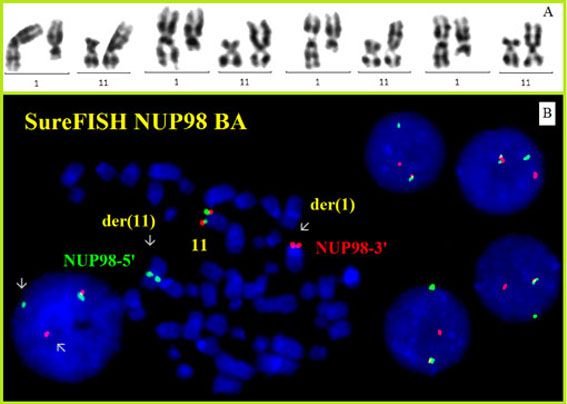
Cytogenetics morphological
Additional anomalies
Genes Involved and Proteins
Result of the Chromosomal Anomaly
Description
Oncogenesis
Highly cited references
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 34903620 | 2022 | Phase Separation Mediates NUP98 Fusion Oncoprotein Leukemic Transformation. | 56 |
| 38493578 | 2024 | Rare NUP98::PRRX1 fusion transcript in a therapy-related acute myeloid leukemia associated with del(7q) following chemotherapy for diffuse large B-cell lymphoma. | 0 |
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 16651408 | 2006 | Trans-repressive effect of NUP98-PMX1 on PMX1-regulated c-FOS gene through recruitment of histone deacetylase 1 by FG repeats. | Bai XT et al |
| 10700860 | 2000 | Translocation (1;11)(q23;p15), a novel simple variant of translocation (7;11)(p15;p15), in a patient with AML (M2) accompanied by non-Hodgkin lymphoma and gastric cancer. | Hatano Y et al |
| 15390187 | 2004 | Analysis of translocations that involve the NUP98 gene in patients with 11p15 chromosomal rearrangements. | Kobzev YN et al |
| 10397741 | 1999 | NUP98 is fused to PMX1 homeobox gene in human acute myelogenous leukemia with chromosome translocation t(1;11)(q23;p15). | Nakamura T et al |
| 16875945 | 2006 | Secondary acute myeloid leukemia with a t(1;11)(q23;p15) in an adolescent treated for testicular sarcoma. | Soares EM et al |
| 17889707 | 2007 | Rare t(1;11)(q23;p15) in therapy-related myelodysplastic syndrome evolving into acute myelomonocytic leukemia: a case report and review of the literature. | Zhang L et al |
Summary
Fusion gene
Note
Citation
Adriana Zamecnikova
t(1;11)(q24;p15) NUP98/PRRX1
Atlas Genet Cytogenet Oncol Haematol. 2018-01-01
Online version: http://atlasgeneticsoncology.org/haematological/1169/t(1;11)(q24;p15)-nup98-prrx1
Historical Card
2005-09-01 t(1;11)(q24;p15) NUP98/PRRX1 by Jean-Loup Huret Affiliation
