t(3;7)(q27;q32) MIR29A/BCL6
2008-12-01 Björn Schneider , Stefan Nagel , Roderick AF MacLeod Affiliation1.DSMZ, German Collection of Microorganisms, Cell Cultures, Department of Human, Animal Cell Cultures, Inhoffenstr. 7b, 38124 Braunschweig, Germany
Clinics and Pathology
Disease
Etiology
SMZL is quite rare, comprising about 1% of lymphomas. Patients typically present with splenomegaly often involving peripheral blood, liver, and bone marrow. About a third of patients have a slight monoclonal gammopathy, thus SMZL may overlap Waldenstroms macroglobulinemia.
Epidemiology
Though rare overall, SMZL is still one of the most common small B-cell lymphoma of the spleen. It mainly affects those over 50 years.
Pathology
The histologic, immunohistochemical, and molecular heterogeneity of SMZL suggests it originates from different (centrocytic, monocytoid, lymphoplasmacytic) B-cell populations residing within normal SMZ.
Treatment
Splenectomy. Responds poorly to chemotherapy.
Evolution
May develop into large cell lymphoma.
Prognosis
Favorable: SMZL displays an indolent course: 10 year survival ≅ 70%.
Cytogenetics
Note
t(3;7)(q27;q32) may be a variant of del(7)(q32) - the main recurrent abnormality reported in SMZL.
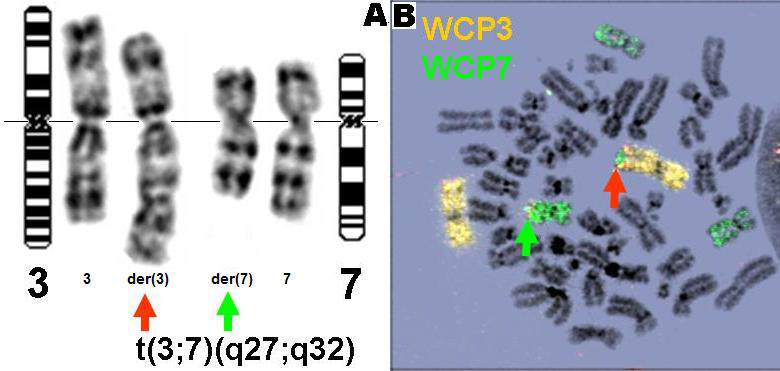
Figure 1: Cytogenetic analysis of t(3;7) in a DLBCL cell line (RC-K8). G-banding (A) and FISH (B) images show t(3;7)(q27;q32) in a DLBCL cell line RC-K8 established from a patient with DLBCL. Expression profiling shows this cell line to express a related but significantly different set of genes from other DLBCL derived cell lines.
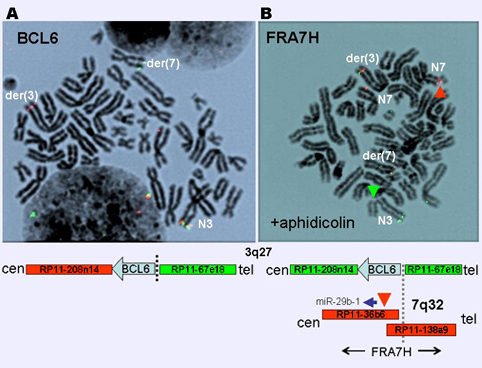
Figure 2: BCL6 and FRA7H Breakpoints in RC-K8 cells. FISH analysis showed rearrangement of BCL6 (A) with FRA7H (B). Treatment of RC-K8 cells with aphidicolin (APC) to induce expression of fragile sites revealed chromatid breaks (ctb) at FRA7H (red arrowhead) as well as elsewhere, e.g. at FRA3D (green). The break at FRA7H induced by APC (B) lies close to the translocation breakpoint present in t(3;7) as determined by LDI-PCR (see below). Interestingly, clastogenesis at FRA7H favored normal chr. 7 homologs over t(3;7) implying stabilization of FRA7H by the latter.
Genes Involved and Proteins
Gene name
BCL6 (B-Cell Lymphoma 6)
Location
3q27.3
Note
Breakpoint lies outwith MBR and ABR.
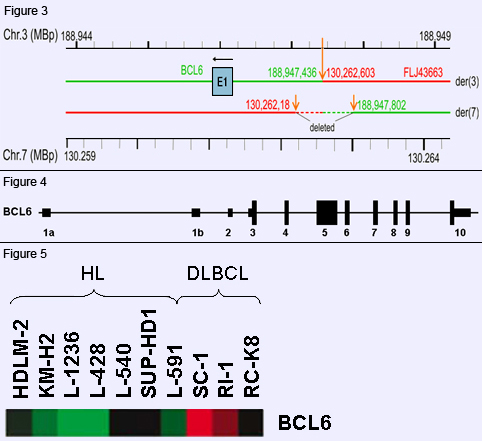
Figure 3: Molecular breakpoint analysis at 3q27 by LDI-PCR. Results of molecular breakpoint analysis by long-distance inverse (LDI)-PCR of the BCL6 and FRA7H junctions on der(3) and der(7) (arrows). Note deletions of 365 bp from chromosome 3 and 416 bp from chromosome 7 (broken lines) on der(7).
Figure 4: BCL6 protein. The BCL6 gene comprizes 10 exons. There are two alternative exons 1 (a or b). Only exons 3-10 harbor protein coding sequences.
Figure 5: Expression of BCL6 in Hodgkin lymphoma and DLBCL. Note preferential expression of BCL6 in DLBCL. RC-K8 t(3;7) cells display moderately upregulated BCL6 expression typical of non-IGH BCL6 rearrangements. Heatmap shows upregulation (red), inconspicuous expression (black) and downregulation (green).
Gene name
MIR29A (microRNA 29a)
Location
7q32.3
Note
miR-29-a and/or miR 29b1 (7q32). miR-29-a/b1 resides inside common fragile site FRA7H (APC inducible, cloned).
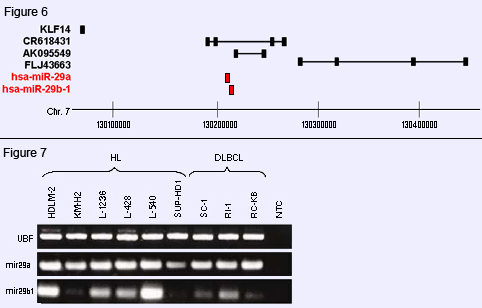
Figure 6: Putative gene targets at 7q32 in t(3;7)(q27;q32). The miR-29 sequences (miR-29a and miR-29b1) are located on chromosome 7q32 upstream of Ref.Seq. gene KLF14 which lies outside FRA7H, and within the intron of a putative uncharacterized gene CR618431.
Figure 7: Expression of miR-29-a/b1 in Hodgkin lymphoma and DLBCL. Expression analysis of miR-29a and miR-29b1 was performed by RT-PCR analysis in HL and DLBCL cell lines in comparison to the control gene UBF. Data indicate low level expression of miR-29-b1 in DLBCL cell lines as compared to HL cell lines.
Result of the Chromosomal Anomaly
Note
t(3;7)(q27;q32) belongs to the emerging class of non-fusogenic BCL6 translocations. These carry upstream BCL6 breakpoints which lie closer to the transcription unit than ABR breakpoints at ≅ 250 KBp. While BCL6 is undoubtedly upregulated in such cases, expression levels lie below those carrying IGH-BCL6 translocations. In the case of t(3;7) the chromosome 7 breakpoint lies within FRA7H, the first FRA firmly associated with an hematopoietic malignant translocation (as opposed to deletion). Physiological BCL6 expression occurs in germinal centers where it is thought to permit immunological DNA breakage by suppressing apoptosis induced by the p53 damage pathway. It is tempting to suppose that BCL6 expression might also incur the risk of untoward breakage at fragile sites.
FRA7H is bereft of RefSeq genes. Apart from putative mRNA transcripts of dubious provenance (several including CR618431 shown in Figure 6 have been inadequately annotated and may be pseudogenes), miR-29-a/b1 are the only verified genes mapped to FRA7H. Deletions affecting the miR-29-a/b1 cluster have been recently linked to SMZL and previously to CLL. Interestingly, a key target of miR-29-a/b1 is TCL1 known to be upregulated in SMZL.
FRA7H is bereft of RefSeq genes. Apart from putative mRNA transcripts of dubious provenance (several including CR618431 shown in Figure 6 have been inadequately annotated and may be pseudogenes), miR-29-a/b1 are the only verified genes mapped to FRA7H. Deletions affecting the miR-29-a/b1 cluster have been recently linked to SMZL and previously to CLL. Interestingly, a key target of miR-29-a/b1 is TCL1 known to be upregulated in SMZL.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15128417 | 2004 | A narrow deletion of 7q is common to HCL, and SMZL, but not CLL. | Andersen CL et al |
| 18023866 | 2008 | New chromosomal alterations in a series of 23 splenic marginal zone lymphoma patients revealed by Spectral Karyotyping (SKY). | Baró C et al |
| 14681321 | 2003 | Splenic marginal zone lymphomas presenting with splenomegaly and typical immunophenotype are characterized by allelic loss in 7q31-32. | Boonstra R et al |
| 12176884 | 2002 | Splenic marginal zone lymphoma: clinical characteristics and prognostic factors in a series of 60 patients. | Chacón JI et al |
| 12446449 | 2003 | Splenic marginal zone lymphoma. | Franco V et al |
| 10086800 | 1999 | The incidence of trisomy 3 in splenic lymphoma with villous lymphocytes: a study by FISH. | Gruszka-Westwood AM et al |
| 11337382 | 2001 | Novel genomic imbalances in B-cell splenic marginal zone lymphomas revealed by comparative genomic hybridization and cytogenetics. | Hernández JM et al |
| 18094718 | 2008 | Splenic marginal zone lymphoma proposals for a revision of diagnostic, staging and therapeutic criteria. | Matutes E et al |
| 9653154 | 1998 | Molecular characterization of a common fragile site (FRA7H) on human chromosome 7 by the cloning of a simian virus 40 integration site. | Mishmar D et al |
| 8136270 | 1993 | Cytogenetic studies in splenic lymphoma with villous lymphocytes. | Oscier DG et al |
| 10862046 | 2000 | Marginal zone B-cell lymphomas (MZBL) arising at different sites represent different biological entities. | Ott MM et al |
| 17625607 | 2007 | MicroRNA losses in the frequently deleted region of 7q in SMZL. | Ruiz-Ballesteros E et al |
| 17989715 | 2008 | T(3;7)(q27;q32) fuses BCL6 to a non-coding region at FRA7H near miR-29. | Schneider B et al |
| 11146574 | 2001 | Splenic marginal zone B-cell lymphomas: two cytogenetic subtypes, one with gain of 3q and the other with loss of 7q. | Solé F et al |
| 18492102 | 2008 | Splenic marginal zone lymphomas are characterized by loss of interstitial regions of chromosome 7q, 7q31.32 and 7q36.2 that include the protection of telomere 1 (POT1) and sonic hedgehog (SHH) genes. | Vega F et al |
Citation
Björn Schneider ; Stefan Nagel ; Roderick AF MacLeod
t(3;7)(q27;q32) MIR29A/BCL6
Atlas Genet Cytogenet Oncol Haematol. 2008-12-01
Online version: http://atlasgeneticsoncology.org/haematological/1515/t(3;7)(q27;q32)-mir29a-bcl6
