t(3;8)(q26;q24) PVT1/MECOM
2007-11-01 Pei Lin Affiliation1.Department of Hematopathology, Box 72, The University of Texas M. D. Anderson Cancer Center, 1515 Holcombe Boulevard, Houston, TX, 77030, USA
Clinics and Pathology
Disease
Acute myeloid leukemia, de novo myelodysplastic syndrome or therapy related myelodysplastic syndrome.
Phenotype stem cell origin
Mostly AML FAB-M2 or FAB-M-4 subtype.
Etiology
Unclear, may be secondary to chemotherapy.
Epidemiology
10 cases reported so far in the literature, less than 1% of AML cases.
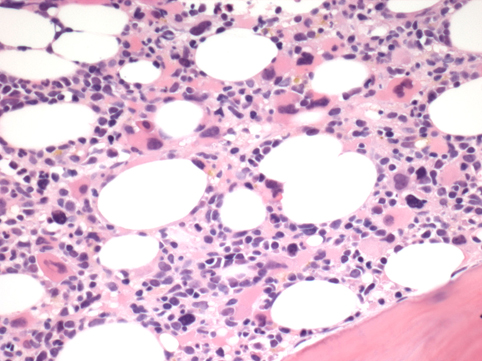
Dysplastic myeloid elements.
Cytology
Acute myeloid leukemia of mostly M2, M4 or M5 FAB subtype or high grade MDS. Marked trilineage dysplasia and megakaryocytic hyperplasia, may be associated with peripheral blood thrombocytosis giving the so-call 3q21q26 syndrome.
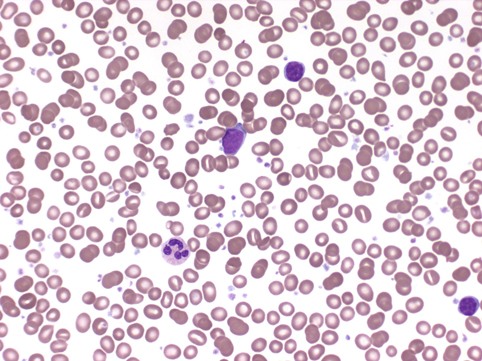
Increased dysplastic megakaryocytes and increased blasts in the interstitium.
Treatment
Chemotherapy; May responds to thalidomide or arsenic better than conventional chemotherapy.
Evolution
Myelodysplastic syndrome progress to acute myeloid leukemia.
Prognosis
Poor.
Genes Involved and Proteins
Gene name
MECOM (Ecotropic Viral Integration Site 1 (EVI1) and Myelodysplastic Syndrome 1 (MDS1-EVI1)
Location
3q26.2
Note
Aberrant EVI1 expression usually occurs in AML, MDS or CML-BC as a result of translocation involving 3q26. The most common ones are inv(3)(q21q26), t(3;3) and t(3;21)(q26;q22). The partner genes of EVI1 are identified as Ribophorin I in inv(3)(q21q26) and t(3;3), AML/MDS1/EAP in t(3;21), and ETV6 in t(3;12), respectively. Others involving t(3;12), t(2;3)(p13;q26), t(3;17)(q26;q22) and t(3;13)(q26;q13-14) are uncommon. Aberrant EVI1 expression also occurs in 10% of acute myeloid leukemia without involving 3q26 and is also correlated with an adverse outcome.
Dna rna description
16 exons spanning 64.2 Kb. Transcriptional orientation is from telomere to centromere. EVI1 gene may be transcribed in different isoform which may have different oncogenic effect.
Protein description
1051 amino acids; 118335 Da. Nuclear location, contains 10 C2H2-type zinc fingers.
Gene name
PVT1 (Pvt1 oncogene (non-protein coding))
Location
8q24.21
Note
The RNA function of pvt1 is unknown.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 12393383 | 2003 | High EVI1 expression predicts poor survival in acute myeloid leukemia: a study of 319 de novo AML patients. | Barjesteh van Waalwijk van Doorn-Khosrovani S et al |
| 17693189 | 2007 | Aberrant EVI1 expression in acute myeloid leukemias associated with the t(3;8)(q26;q24). | Lennon PA et al |
| 16342172 | 2006 | EVI1 is consistently expressed as principal transcript in common and rare recurrent 3q26 rearrangements. | Poppe B et al |
Summary
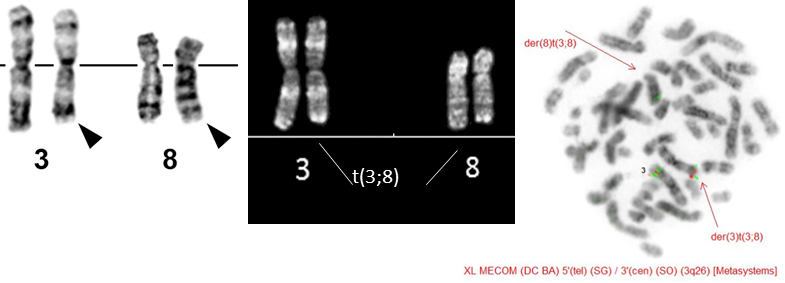
t(3;8)(q26;q24) PVT1/MECOM G-banding (left) u2013 Courtesy Pei Lin; R-banded karyotype (middle) and FISH using dual color break apart probe XL MECOM (Metasystems)(right) - Courtesy Karolien Beel, Peter Meeus, Geneviu00e8ve Ameye and Lucienne Michaux
Citation
Pei Lin
t(3;8)(q26;q24) PVT1/MECOM
Atlas Genet Cytogenet Oncol Haematol. 2007-11-01
Online version: http://atlasgeneticsoncology.org/haematological/1463/t(3;8)(q26;q24)-pvt1-mecom
