t(4;22)(q12;q11) BCR/PDGFRA
2004-09-01 Barbara K Goodman , Anne Michele Safely Affiliation1.Clinical Cytogenetics, Molecular Diagnostics Laboratories, Duke University Health System, Box 3631, Durham, NC 27710, UK
Clinics and Pathology
Disease
Reported in 3 cases; myeloproliferative disorder : atypical chronic myeloid leukemia (aCML) (Philadelphia chromosome-negative) .
Etiology
One case was post-treatment for lymphoma, thus suspected to be a secondary MPD.
Clinics
3 cases shared characteristics of splenic enlargement, eosinophilia and male predominance.
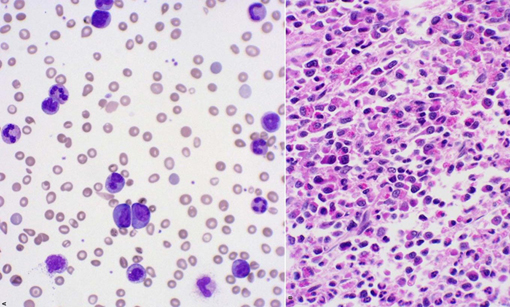
Peripheral blood smear showing anemia, leukocytosis with increased granulocytes and precursors, and eosinophilia. Bone marrow showing hypercellularity with marked myeloid hyperplasia, mild eosinophilia, no increase in blasts.
Prognosis
One patient on treatment with good response to imatinib mesylate.
Cytogenetics
Note
t(4;22)(q12;q11.2), easily distinguished cytogenetically.
Cytogenetics molecular
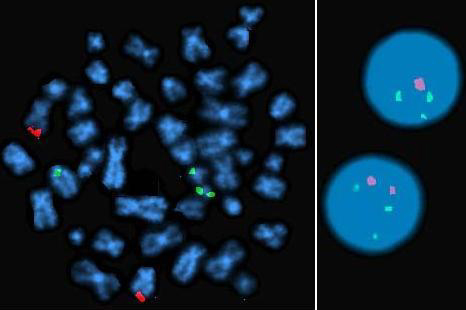
Interphase FISH showing 3 signals for BCR (green), suggesting rearrangement. Two normal signals for ABL1 (red).
Additional anomalies
The t(4;22) was observed as the sole anomaly in the few reported cases to date.
Genes Involved and Proteins
Gene name
PDGFRA (platelet-derived growth factor receptor, alpha polypeptide)
Location
4q12
Note
Dna rna description
Protein description
170-kD transmembrane glycoprotein that normally binds all PDGF isoforms, AA, AB, and BB at its extracellular Ig domain.
Gene name
BCR (Breakpoint cluster region)
Location
22q11.23
Dna rna description
23 exons; alternate splicing.
Protein description
160-kDa protein; contains a unique serine/threonine kinase activity and at least two SH2 binding sites encoded in its first exon and a C-terminal domain that functions as a GTPase activating protein for p21(rac).
Result of the Chromosomal Anomaly
Note
5BCR-3PDGFRA fusion
Description
Fusion of BCR exon 7, 12 or 17 (in 3 cases described) with PDGFRA exon 12, in frame, containing intronic sequence from BCR in two cases.169 kDa protein in one case, somewhat smaller in two others.
Expression localisation
Predicted to be localized intracellularly
Oncogenesis
Alteration of tyrosine kinase activity secondary to loss of regulatory and PDGF binding domains; also, BCR domains may significantly affect BCR-PDGFRA downstream signaling pathways as seen with BCR-ABL fusion.
Highly cited references
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 19366796 | 2009 | Ligand-dependent platelet-derived growth factor receptor (PDGFR)-alpha activation sensitizes rare lung cancer and sarcoma cells to PDGFR kinase inhibitors. | 31 |
| 35713428 | 2022 | A rare cause of persistent leukocytosis with massive splenomegaly: Myeloid neoplasm with BCR-PDGFRA rearrangement-Case report and literature review. | 15 |
| 26095243 | 2015 | BCR-PDGFRA fusion in a T lymphoblastic leukemia/lymphoma. | 0 |
| 12023981 | 2002 | The t(4;22)(q12;q11) in atypical chronic myeloid leukaemia fuses BCR to PDGFRA. | 0 |
| 15034867 | 2004 | Molecular and cytogenetic characterization of a novel translocation t(4;22) involving the breakpoint cluster region and platelet-derived growth factor receptor-alpha genes in a patient with atypical chronic myeloid leukemia. | 0 |
| 25941032 | 2015 | Novel fusion between the breakpoint cluster region and platelet-derived growth factor receptor-alpha genes in a patient with chronic myeloid leukemia-like neoplasm: undetectable residual disease after imatinib therapy. | 0 |
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 12023981 | 2002 | The t(4;22)(q12;q11) in atypical chronic myeloid leukaemia fuses BCR to PDGFRA. | Baxter EJ et al |
| 12542482 | 2003 | Novel translocations that disrupt the platelet-derived growth factor receptor beta (PDGFRB) gene in BCR-ABL-negative chronic myeloproliferative disorders. | Baxter EJ et al |
| 12353319 | 2002 | CML after treatment for lymphoid malignancy: Therapy-related CML or coincidence? | Bolaños-Meade J et al |
| 2536372 | 1989 | Identification and structural analysis of the A type receptor for platelet-derived growth factor. Similarities with the B type receptor. | Claesson-Welsh L et al |
| 12522257 | 2003 | PDGFRA activating mutations in gastrointestinal stromal tumors. | Heinrich MC et al |
| 8586421 | 1995 | Structure, organization, and transcription units of the human alpha-platelet-derived growth factor receptor gene, PDGFRA. | Kawagishi J et al |
| 9407086 | 1997 | Functional importance of platelet-derived growth factor (PDGF) receptor extracellular immunoglobulin-like domains. Identification of PDGF binding site and neutralizing monoclonal antibodies. | Lokker NA et al |
| 10748292 | 2000 | A FISH study of variant Philadelphia rearrangements. | Reddy KS et al |
| 15034867 | 2004 | Molecular and cytogenetic characterization of a novel translocation t(4;22) involving the breakpoint cluster region and platelet-derived growth factor receptor-alpha genes in a patient with atypical chronic myeloid leukemia. | Safley AM et al |
| 12944919 | 2003 | Chronic myeloproliferative disorders with rearrangement of the platelet-derived growth factor alpha receptor: a new clinical target for STI571/Glivec. | Trempat P et al |
Summary
Fusion gene
BCR/PDGFRA BCR (22q11.23) PDGFRA (4q12) M t(4;22)(q12;q11)|BCR/PDGFRA BCR (22q11.23) PDGFRA (4q12) TIC
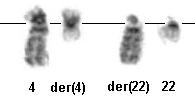
Partial karyotype showing chromosomes 4 and 22.
Citation
Barbara K Goodman ; Anne Michele Safely
t(4;22)(q12;q11) BCR/PDGFRA
Atlas Genet Cytogenet Oncol Haematol. 2004-09-01
Online version: http://atlasgeneticsoncology.org/haematological/1153/t(4;22)(q12;q11)-bcr-pdgfra
