t(11;17)(q23;q12-21) KMT2A/MLLT6
2005-08-01 Sabine Strehl Affiliation1.Childrens Cancer Research Institute, Kinderspitalgasse 6, A-1090 Vienna, Austria
Cytogenetics
Note
so far three MLL fusion partners, namely LASP1, AF17 (alias MLLT6), and ACACA have been identified in 17q12-21; these translocations cannot be distinguished cytogenetically and the accurate detection of the specific fusion gene requires RT-PCR or refined FISH analysis
Cytogenetics molecular
chromosomes (arrows).
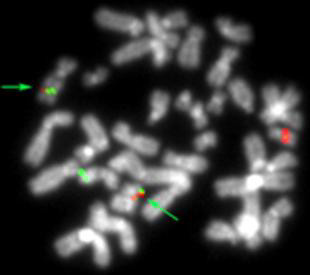
FISH using a combination of the MLL-specific PACs 217a21 and 167k13 (green signals) and the AF17-specific BAC RP11-25H10 (red signals) results in two fusion signals on the der(11) and the der(17)
Additional anomalies
+8
Genes Involved and Proteins
Gene name
KMT2A (myeloid/lymphoid or mixed lineage leukemia)
Location
11q23.3
Dna rna description
37 exons, spanning over 100 kb; transcription in a centromeric to telomeric direction; 13 -15 kb mRNA; coding sequence 11.9 kb
Protein description
431 kDa; contains two DNA binding motifs (an AT hook, and Zinc fingers), a DNA methyl transferase motif, and a bromodomain; transcriptional regulatory factor; nuclear localization
Gene name
MLLT6 (myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 6)
Location
17q12
Note
previously LASP1 and AF17 (alias MLLT6) were mapped to 17q21, but according to the most recent genome assembly built by the Genome Bioinformatics Group of the University of California Santa Cruz and recent FISH data both genes are localized in 17q12 and proximal to RARA
Dna rna description
encompasses 19.97 kb of genomic DNA; 4914 bp mRNA; 20 exons, 3282 bp coding sequence
Protein description
1023 amino acids, 112 kDa; MLLT6 (alias AF17), MLLT10 (alias AF10), and BRPF1 (alias BR140) belong to a small evolutionary highly conserved family of putative nuclear transcription factors, which contain amino-terminal PHD fingers, a highly conserved octapeptide, and a classical leucine zipper dimerization motif; nuclear localization
Result of the Chromosomal Anomaly
Transcript
5 MLL - 3 AF17
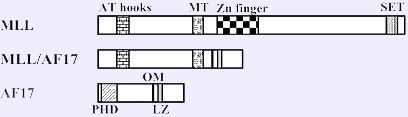
Schematic representation of MLL, AF17 (alias MLLT6), and the putative MLL-AF17 fusion protein. MT, methyltransferase domain; Zn finger, Zinc finger domain; SET-domain; PHD, Zinc finger PHD-type; OM, octapeptide motif; LZ, leucine-zipper dimerization motif.
Description
the AT-hook DNA-binding domain and the methyltransferase motif including the CXXC zinc-finger (Zn) domain of MLL are fused to the highly conserved octapeptide (OM) and the leucine-zipper (LZ) dimerization motif of AF17 (alias MLLT6).
Highly cited references
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 33832921 | 2021 | Clinical impact of genomic characterization of 15 patients with acute megakaryoblastic leukemia-related malignancies. | 24 |
| 30277115 | 2019 | Panoramic view of common fusion genes in a large cohort of Chinese de novo acute myeloid leukemia patients. | 0 |
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15322560 | 2004 | A diagnostic biochip for the comprehensive analysis of MLL translocations in acute leukemia. | Maroc N et al |
| 15676155 | 2005 | Acute myelocytic leukemia with t(11;17)(q23;q12-q21) involves a fusion of MLL and AF17. | Moore SD et al |
| 8058765 | 1994 | Leucine-zipper dimerization motif encoded by the AF17 gene fused to ALL-1 (MLL) in acute leukemia. | Prasad R et al |
| 12805060 | 2003 | AML with 11q23/MLL abnormalities as defined by the WHO classification: incidence, partner chromosomes, FAB subtype, age distribution, and prognostic impact in an unselected series of 1897 cytogenetically analyzed AML cases. | Schoch C et al |
Summary
Note
not to be confused with the t(11;17)(q23;q12-21) involving MLL and LASP1 or the t(11;17)(q23;q12-21) involving MLL and ACACA
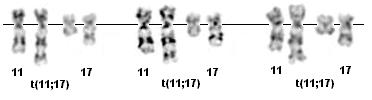
t(11;17)(q23;q12) G-banding - Courtesy Cytogenetics Laboratory of the CCRI, Childrens Cancer Research Institute, Vienna.
Citation
Sabine Strehl
t(11;17)(q23;q12-21) KMT2A/MLLT6
Atlas Genet Cytogenet Oncol Haematol. 2005-08-01
Online version: http://atlasgeneticsoncology.org/haematological/1027/t(11;17)(q23;q12-21)-kmt2a-mllt6
