AIFM2 (apoptosis-inducing factor, mitochondrion-associated, 2)
2009-06-01 Miroslav Varecha AffiliationCentre for Biomedical Image Analysis, Faculty of Informatics, Masaryk University, Botanicka 68a, Brno 60200, Czech Republic
Identity
HGNC
LOCATION
10q22.1
LOCUSID
ALIAS
AMID,FSP1,PRG3
FUSION GENES
DNA/RNA
Description
The gene spans approximately 20.66kb. Number of exons: 14, minus strand.
Transcription
The length of AIFM2 transcript is 3240 bp.
Proteins
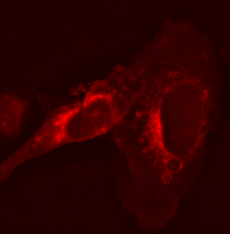
Stably transfected (by lipofection) living cells U-2 OS (human osteosarcoma) cell line with plasmid producing red fluorescent fusion protein AIFM2-tHcRed.
Description
Expression
AIFM2 was detected in most healthy tissues in form of two transcripts (1.8 and 4.0 kb). It is highly expressed in heart, moderately in liver and skeletal muscles, and expressed at low levels in placenta, lung, kidney, and pancreas.
Localisation
Cytoplasmic side of cellular membranes.
Function
Oxidoreductase, that may be important in mediating a TP53/p53-dependent apoptotic response. Predicted to be caspase-independent effector of apoptotic cell death, but not shown by other authors. Function of this protein is thus unknown.
Homology
Homologous to AIFM1 (AIF, PDCD8). They share 22% aminoacid identity. It belongs to the FAD-dependent oxidoreductase family.
Implicated in
Entity name
Apoptosis and Cancer
Note
AIFM2 expression was found to be activated by overexpression of p53, which leads to cell cycle arrest or apoptosis. Inactivation of p53 was observed in many human cancers.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 18368494 | 2008 | AMID: new insights on its intracellular localization and expression at apoptosis. | Bilyy R et al |
| 17711848 | 2007 | DNA binding suppresses human AIF-M2 activity and provides a connection between redox chemistry, reactive oxygen species, and apoptosis. | Gong M et al |
| 15958387 | 2005 | The human apoptosis-inducing protein AMID is an oxidoreductase with a modified flavin cofactor and DNA binding activity. | Marshall KR et al |
| 16186796 | 2006 | The p53-inducible apoptotic protein AMID is not required for normal development and tumor suppression. | Mei J et al |
| 12135761 | 2002 | A novel p53-inducible apoptogenic gene, PRG3, encodes a homologue of the apoptosis-inducing factor (AIF). | Ohiro Y et al |
| 17347867 | 2007 | Bioinformatic and image analyses of the cellular localization of the apoptotic proteins endonuclease G, AIF, and AMID during apoptosis in human cells. | Varecha M et al |
| 15273740 | 2004 | AMID is a p53-inducible gene downregulated in tumors. | Wu M et al |
Other Information
Locus ID:
NCBI: 84883
MIM: 605159
HGNC: 21411
Ensembl: ENSG00000042286
Variants:
dbSNP: 84883
ClinVar: 84883
TCGA: ENSG00000042286
COSMIC: AIFM2
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000042286 | ENST00000307864 | Q9BRQ8 |
| ENSG00000042286 | ENST00000373248 | Q9BRQ8 |
| ENSG00000042286 | ENST00000613322 | Q9BRQ8 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38244593 | 2024 | Circ0060467 sponges miR-6805 to promote hepatocellular carcinoma progression through regulating AIFM2 and GPX4 expression. | 1 |
| 38244593 | 2024 | Circ0060467 sponges miR-6805 to promote hepatocellular carcinoma progression through regulating AIFM2 and GPX4 expression. | 1 |
| 36443441 | 2023 | Elevated FSP1 protects KRAS-mutated cells from ferroptosis during tumor initiation. | 28 |
| 36484954 | 2023 | Fear stress promotes glioma progression through inhibition of ferroptosis by enhancing FSP1 stability. | 6 |
| 36788244 | 2023 | A genome-wide CRISPR-Cas9 knockout screen identifies FSP1 as the warfarin-resistant vitamin K reductase. | 22 |
| 37080964 | 2023 | Molecular characterization of AIFM2/FSP1 inhibition by iFSP1-like molecules. | 12 |
| 37380771 | 2023 | Phase separation of FSP1 promotes ferroptosis. | 39 |
| 37587773 | 2023 | TRIM21-Promoted FSP1 Plasma Membrane Translocation Confers Ferroptosis Resistance in Human Cancers. | 7 |
| 37619958 | 2023 | Ferroptosis of macrophages facilitates bone loss in apical periodontitis via NRF2/FSP1/ROS pathway. | 2 |
| 37695069 | 2023 | Sorafenib induces ferroptosis by promoting TRIM54-mediated FSP1 ubiquitination and degradation in hepatocellular carcinoma. | 5 |
| 37903990 | 2023 | YTHDC1 as a tumor progression suppressor through modulating FSP1-dependent ferroptosis suppression in lung cancer. | 6 |
| 36443441 | 2023 | Elevated FSP1 protects KRAS-mutated cells from ferroptosis during tumor initiation. | 28 |
| 36484954 | 2023 | Fear stress promotes glioma progression through inhibition of ferroptosis by enhancing FSP1 stability. | 6 |
| 36788244 | 2023 | A genome-wide CRISPR-Cas9 knockout screen identifies FSP1 as the warfarin-resistant vitamin K reductase. | 22 |
| 37080964 | 2023 | Molecular characterization of AIFM2/FSP1 inhibition by iFSP1-like molecules. | 12 |
Citation
Miroslav Varecha
AIFM2 (apoptosis-inducing factor, mitochondrion-associated, 2)
Atlas Genet Cytogenet Oncol Haematol. 2009-06-01
Online version: http://atlasgeneticsoncology.org/gene/41842/aifm2
