B3GNT6 (UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 6 (core 3 synthase))
2009-05-01 Neeru M Sharma , Prakash Radhakrishnan , Shuhua Tan , Pi-Wan Cheng AffiliationDepartment of Biochemistry, Molecular Biology, College of Medicine, University of Nebraska Medical Center, 985870 Nebraska Medical Center, Omaha, NE 68198-5870, USA
DNA/RNA
Note
Human B3GNT6 is located on chromosome 11 in the region of q13.4.
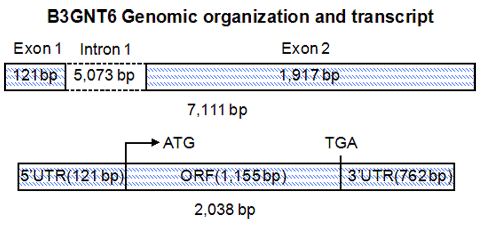
The schematic representation of human B3GNT6 gene and its transcript (ATG, translation start codon; TGA, translation stop codon; UTR, Untranslated region; ORF, Open reading frame).
Description
Human B3GNT6 gene is 7,111 bp in length, composed of 2 exons and 1 intron, and located at chromosome 11q13.4.
Transcription
B3GNT6 transcript contains two exons. Exon 1 is 121 bp and exon 2 is 1,917 bp. The exon 2 contains 1,155 bp ORF and 762 bp 3UTR.
Pseudogene
-NA-
Proteins
Note
Human UDP-GlcNAc:GalNAca1-Ser/Thr beta-1,3-N-acetylglucosaminyltransferase 6 (core 3 synthase) has 384 amino acids and 43 KDa molecular weight.

The predicted B3GNT6 structure contains a short N-terminal cytoplasmic tail (CT) (12 aa), a transmembrane domain (TM) (19 aa), a long stem region and catalytic domain (353 aa) at the C-terminal region.
Description
B3GNT6/C3S is single-pass type II membrane protein belonging to the glycosyltransferase 31 family.
Expression
B3GNT6/C3S gene expression is restricted to mucus-secretory tissues. The level of beta3GnT6 transcript expressed in various human tissues as measured by the real time PCR revealed that the expression level was highest in the stomach, followed by the colon and small intestine. Skeletal muscle and testis expressed the beta3GnT6 transcript at a moderate level. The expression levels in the remaining tissues were very low or not detectable. Its expression was markedly down-regulated in gastric and colorectal carcinomas, which include both tumor tissues and cell lines-derived from these carcinomas.
Localisation
Golgi Membrane.
Function
B3GNT6/C3S enzyme catalyzes the transfer of GlcNAc from UDP-GlcNAc to GalNAcalpha1-Ser/Thr (Tn antigen) to form the core 3 structure (GlcNAcbeta1-3GalNAcalpha1-Ser/Thr). Core 3 can be extended by the addition of galactose and then other sugars to generate biologically important epitopes or serves as the precursor for the formation of core 4, which in turn can be further elaborated to form more complex structure. Core 3-containing O-glycans are found in the secreted mucins produced in the mucus-secretory tissues. Loss of core 3 synthase results in the loss of not only core 3 glycans but also core 4 glycans. Loss of core 3 could lead to the production of secreted mucins with compromised mucus protection function. As a result, mucus would be more dehydrated, bacteria would be inefficiently cleared from the system, and chronic inflammation would be developed, which eventually would result in development of cancer. A mouse model devoid of core 3 synthase gene has been shown to develop colon cancer. Because the loss of this gene leads to development of colon cancer, B3GNT6/C3S gene is a tumor suppressor gene.
Homology
An alignment of the amino acid sequences of five B3GnTs made using ClustalW showed 41, 54, 42, and 35% sequence identity between B3GnT6 and B3GnT2, B3GnT3, B3GnT4, and B3GnT5, respectively, and this sequence similarity was limited to the putative catalytic domains. Five cysteine residues were conserved among these five B3GnTs. However, only B3GNT6/C3S exhibits significant core 3 synthase activity.
Implicated in
Entity name
Gastric and Colorectal carcinomas
Note
Colorectal cancer, which is also called colon cancer or large bowel cancer, includes cancerous growths in the colon, rectum and appendix. Loss of function of the mucus layer is one major cause of this cancer. Globally, cancer of the colon and rectum is the third leading cause of cancer in males and the fourth leading cause of cancer in females. The frequency of colorectal cancer varies around the world. It is common in the Western world and is rare in Asia and Africa. In countries where the people have adopted western diets, the incidence of colorectal cancer is increasing. Colorectal cancer can take many years to develop and early detection of colorectal cancer greatly improves the chances of a cure. Therefore, screening for the disease is recommended in individuals who are at increased risks. Prevention and early detection are key factors in controlling and curing colorectal cancer. Indeed, colorectal cancer is the second most preventable cancer, after lung cancer.
Prognosis
B3GNT6/C3S is down-regulated in gastric and colorectal carcinomas, suggesting that it may be used as a marker for distinguishing between benign adenomas and premalignant lesions.
Oncogenesis
O-linked oligosaccharides (O-glycans) are the primary components of the intestinal mucus layer that covers the gastrointestinal epithelium. This layer is a dense, carbohydrate-rich matrix that consists primarily of mucins containing multiple serine and threonine residues, which have been modified by O-glycans and account for 80-90% of the mucin mass. The mucus layer and epithelial cells comprise an intestinal barrier that protects epithelial and intestinal mucosal immune cells from potentially harmful luminal microflora and food components. Among all mucin glycan core structures, Core 3 and core 4 are unique to secreted mucins, which may play important roles in protecting the molecular integrity of these mucins and enable them to perform their functions under extreme harsh conditions, such as gastric and colonic environment. Loss of these functions resulted from the loss of core 3 synthase is thought to initiate oncogenesis in the gastrointestinal tract.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 17517967 | 2007 | Increased susceptibility to colitis and colorectal tumors in mice lacking core 3-derived O-glycans. | An G et al |
| 16741504 | 2006 | Mucin-type O-glycans in human colon and breast cancer: glycodynamics and functions. | Brockhausen I et al |
| 11821425 | 2002 | Molecular cloning and characterization of a novel UDP-GlcNAc:GalNAc-peptide beta1,3-N-acetylglucosaminyltransferase (beta 3Gn-T6), an enzyme synthesizing the core 3 structure of O-glycans. | Iwai T et al |
| 15755813 | 2005 | Core 3 synthase is down-regulated in colon carcinoma and profoundly suppresses the metastatic potential of carcinoma cells. | Iwai T et al |
Other Information
Locus ID:
NCBI: 192134
MIM: 615315
HGNC: 24141
Ensembl: ENSG00000198488
Variants:
dbSNP: 192134
ClinVar: 192134
TCGA: ENSG00000198488
COSMIC: B3GNT6
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000198488 | ENST00000528622 | E9PJ79 |
| ENSG00000198488 | ENST00000622824 | Q6ZMB0 |
| ENSG00000198488 | ENST00000622824 | A8K9Q8 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 35387659 | 2022 | Downregulation of B3GNT6 is a predictor of poor outcomes in patients with colorectal cancer. | 1 |
| 35387659 | 2022 | Downregulation of B3GNT6 is a predictor of poor outcomes in patients with colorectal cancer. | 1 |
| 33253272 | 2020 | Clinicopathological significance of core 3 O-glycan synthetic enzyme, β1,3-N-acetylglucosaminyltransferase 6 in pancreatic ductal adenocarcinoma. | 5 |
| 33253272 | 2020 | Clinicopathological significance of core 3 O-glycan synthetic enzyme, β1,3-N-acetylglucosaminyltransferase 6 in pancreatic ductal adenocarcinoma. | 5 |
| 19395705 | 2009 | Core3 O-glycan synthase suppresses tumor formation and metastasis of prostate carcinoma PC3 and LNCaP cells through down-regulation of alpha2beta1 integrin complex. | 34 |
| 19395705 | 2009 | Core3 O-glycan synthase suppresses tumor formation and metastasis of prostate carcinoma PC3 and LNCaP cells through down-regulation of alpha2beta1 integrin complex. | 34 |
| 15755813 | 2005 | Core 3 synthase is down-regulated in colon carcinoma and profoundly suppresses the metastatic potential of carcinoma cells. | 60 |
| 15755813 | 2005 | Core 3 synthase is down-regulated in colon carcinoma and profoundly suppresses the metastatic potential of carcinoma cells. | 60 |
Citation
Neeru M Sharma ; Prakash Radhakrishnan ; Shuhua Tan ; Pi-Wan Cheng
B3GNT6 (UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 6 (core 3 synthase))
Atlas Genet Cytogenet Oncol Haematol. 2009-05-01
Online version: http://atlasgeneticsoncology.org/gene/44427/b3gnt6
