BABAM2 (brain and reproductive organ-expressed (TNFRSF1A modulator))
2010-06-01 Yiu-Loon Chui , Kenneth Ka-Ho Lee , John Yeuk-Hon Chan AffiliationIdentity
HGNC
LOCATION
2p23.2
LOCUSID
ALIAS
BRCC4,BRCC45,BRE
FUSION GENES
DNA/RNA
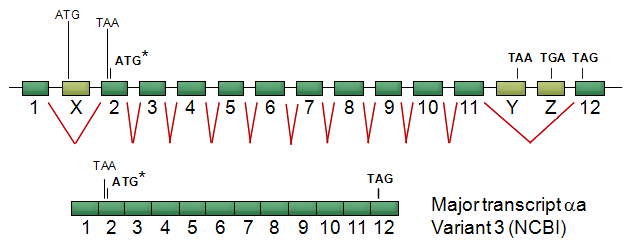
Generation of the major transcript variant of human BRE. Human BRE gene (not drawn in scale) is consisted of 15 exons, three of which are alternatively spliced. The light green boxes, X - Z, are alternative exons which are not present in the major transcript. The asterisked ATG is the translation start. This transcript encodes the major 383-amino-acid BRE protein isoform 2, that has been studied.
Description
The gene spans 448284 bases, telomere to centromere orientation. The first exon is non-coding. In humans, six transcript variants are produced by alternative splicing predominantly at either end of the gene. All human cells examined co-express all of the splice variants, but at different ratios to one another. The major transcript is αa, also known as variant 3 by NCBI nomenclature. This transcript encodes the ubiquitous 383-amino-acid protein, designated by NCBI as protein isoform 2 (NP_954661.1) (Ching et al., 2001). Functions of all minor transcript variants are undetermined. In mice, alternative splicing occurs only at the 5 region of the gene. The major transcript is variant 5 (NCBI nomenclature), which encodes the ubiquitous 383-amino acid protein that is 99% identical to human BRE. The minor transcript variants, unlike the human counterparts, are expressed differentially among tissues. Their functions are undetermined (Ching et al., 2003).
Transcription
Exon 1 is non-coding; its flanking sequences are embedded in a CpG island of 1216 bases long. Transcription start varies over the region between 35 to 112 bases upstream of the last base of exon 1, with the most common site at 40 bases upstream. No TATA or CAAT box is located within 150 bases upstream of any of the transcription start sites. BRE mRNA is expressed ubiquitously, and was initially found to be highly expressed in brain, and reproductive organs; hence the name "BRE" (Li et al., 1995). Subsequent screens using human multiple-tissue RNA dot blot and Northern blot revealed highest transcript expression in adrenal and heart (Miao et al., 2001).
Pseudogene
No pseudogene found.
Proteins
Note
BRE is a 383-amino-acid protein of no identifiable functional domain by sequence homology. No crystal structure of BRE is available. This protein has no paralog. The N-terminal region of 333 residues of human BRE, which is conserved among vertebrate orthologs, has been classified as a single unique domain, pfam06113. It has been recently proprosed that BRE contains 2 ubiquitin E2 variant (UEV) domains (Wang et al., 2009).
Description
BRE is an evolutionarily highly conserved protein with no homolog within the same species. The major protein isoform is 383 amino-acids long. Based on bioinformatic analysis, BRE was proposed to have two ubiquitin-binding UEV (Ubiquitin E2 variant) domains. One was located in the N-terminal region between residues 30 and 147. The other one, however, could only be located in the isoform encoded by a rare transcript variant 1, as the C-terminal one quarter of the putative domain is encoded by the alternative exon Y (Wang et al., 2009). Thus, it is not clear whether the remaining putative UEV domain sequence from residues 275 to 363 of the major BRE isoform is functional.
Expression
BRE is ubiquitously expressed. All mammalian cell lines examined express high levels of BRE. These cell lines include Jurkat, KRC/Y, HeLa, HepG2, HL60, MCF7, NIH3T3, NS0, THP-1, and lymphoblastoid CB14022 cells. Among mouse tissues, the expression levels of BRE detected by Western blot analysis showed the following pattern: lungs = spleen = thymus > adrenal > testis = kidney > brain > heart = liver. Human hepatocytes express little BRE as detected by immumnohistochemistry and Western blot analysis (Chan et al., 2008).
Localisation
BRE is located in cytoplasm and nucleus.
Function
DNA-repair and anti-apoptosis via regulation of ubiquitination. BRE was shown able to bind K48- and K63-linked polyubiquitin chains (Wang et al., 2009). BRE and its mouse ortholog are expressed in cytosolic and nuclear compartments (Li et al., 2004). In the nucleus, BRE is part of the BRCA1-A complex involved in DNA repair and maintaining G2/M arrest in response to DNA damage. BRCA1-A complex consists of BRCA1, BARD1, Abraxas/Abra1/CCDC98, RAP80, BRCC36, BRE, and MERIT40/NBA1 (Dong et al., 2003; Sobhian et al., 2007; Feng et al., 2009; Shao et al., 2009; Wang et al., 2009). BRE interacts strongly with MERIT40 and is responsible for binding the latter to the complex of Abraxas, RAP80 and BRCC36 (Feng et al., 2009). BRE may also regulate the K63 deubiquitinase activity of BRCC36 (Sobhian et al., 2007). In conjunction with BRCC36, BRE was shown to potentiate the E3 activity of BRCA1-BARD1 complex (Dong et al., 2003). Furthermore, depletion of BRE by siRNA sensitized cells to death induced by ionizing irradiation (Dong et al., 2003; Feng et al., 2009). This protein also forms multiprotein BRISC (Brcc36 isopeptidase complex) in the cytoplasm. BRISC, containing at least 3 proteins, FAM175B/ABRO1, BRCC36 and MERIT40/NBA1, in addition to BRE, specifically cleaves K63-linked polyubiquitin chains (Cooper et al., 2009). It is not known whether such cytosolic complex is responsible for attenuating apoptotic response emanating from the activated death receptors, TNF-R1 and Fas. BRE also binds to the cytoplasmic region of TNF-R1 and Fas, as well as the death-inducing signaling complex (DISC) during apoptotic induction (Gu et al., 1998; Li et al., 2004). The anti-apoptotic role of BRE has been shown by the increased apoptotic response to TNF-alpha of HeLa cell line depleted of BRE by siRNA, and the attenuated response of HeLa and Jurkat to TNF-alpha and anti-Fas agonist antibody by over-expression of the protein. As over-expression of BRE also reduced intrinsic apoptotic response induced by stress-related and genotoxic stimuli, it has been proposed that the death receptor-associating BRE inhibits the recruitment of mitochondrial apoptotic machinery, which is necessary for amplifying the death-receptor-initiated apoptosis of CD95 type II cell types, which include HeLa, Jurkat, and hepatocytes (Scaffidi et al., 1998; Engels et al., 2000). Ectopic expression of BRE in mouse Lewis lung carcinoma cells was shown to promote tumor growth in footpad injection model, but have no effect on cell proliferation in culture condition (Chan et al., 2005). Over-expression of BRE was found in 74% of 123 samples of human hepatocellular carcinoma, and the protein expression level correlated with poor prognosis. Immortalized human cell lines also uniformly express high levels of BRE regardless of the tissue origin of these cell lines. Transgenic expression of BRE in mouse liver attenuated acute fulminant hepatitis induced by anti-Fas antibody, and promoted diethylnitrosamine-induced, but not spontaneous, liver tumors (Chan et al., 2008; Chui et al., 2010). Thus, it is likely that BRE over-expression enhances tumor survival through its anti-apoptotic activity, rather than initiates tumor formation.
Homology
No homologous protein of BRE found within the same species.
Mutations
Note
According to HapMap genotyped SNP data, there is no SNP polymorphism in any of the coding exons of BRE.
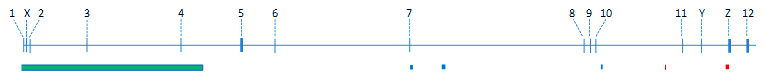

Copy number polymorphism (CNP) of BRE gene. Regions with CNP are shown in colored boxes. Blue and red indicate copy loss and gain, respectively. Green indicates loss and gain at different segments of the contiguous region. The largest CNP region on the far left spans the first non-coding and the next 3 coding exons (exons 1, 2, 3 and 4) and extends further upstream into the neighboring ribokinase gene. The copy gain variant at the far right spans the alternative exon Z. Data obtained from HapMap.
Germinal
According to the current HapMap_rel27 for all the 4 populations (CEU, CHB, JPT and YRI), the number of nucleotide positions in BRE gene with HapMap genotyped SNP is 453. Given the size of BRE gene of 448284 bases long, the number of bases with SNP fits well to the average genome-wide figure of one SNP per 1000 bases (Dutt and Beroukhim, 2007). It is, however, noteworthy that no SNP has been found in any of the coding exons. All of the SNPs, except one located in the 5 UTR, are present in the introns. Two recombination hotspots are located in the introns, one of which is from position 28271535 to 28276573, located between coding exons 7 and 8. The other one is from position 28338948 to 28341210, located between coding exons 10 and 11. Copy number polymorphisms (CNP) involving a large contiguous region of 163295 bases encompassing the first 3 coding exons and the upstream sequence of the neighbouring ribokinase gene and smaller downstream regions have also been identified (see diagram above) (The International HapMap Consortium, 2003).
Somatic
One R9L mutation was identified in a lung carcinoma cell line, NIH-H2126, and a synonymous mutation S182S in a clear cell renal cell carcinoma sample, PD2198a.
Implicated in
Entity name
Hepatocellular carcinoma (HCC)
Note
Immunohistochemical analysis, supplemented by immunoblotting, has revealed overexpression of BRE in the tumoral regions of 72% of the 123 human HCC samples examined. Non-tumoral liver regions, cirrhotic or otherwise, expressed little BRE (Chan et al., 2008).
Prognosis
The over-expression levels of BRE correlated with poor differentiation of HCC cells and therefore poor prognosis.
Cytogenetics
Not determined.
Hybrid gene
Not determined.
Fusion protein
No fusion protein reported.
Oncogenesis
The transgenic mouse model with liver-specific over-expression of human BRE showed no enhanced spontaneous tumor development, indicating that BRE over-expression alone is not tumorigenic. These mice, however, showed significant attenuation of liver apoptosis induced by injection anti-Fas agonist antibody. These findings indicate that the over-expression of BRE in HCC is related to the anti-apoptotic activity of the protein which promotes growth of the carcinoma (Chan et al., 2008). Recent work on inducing liver carcinoma to the above transgenic mice by neonatal injection of diethylnitrosamine (DEN) confirmed that BRE over-expression in the liver could only promote growth of the already initiated tumor, rather than on initiating tumor formation. Interestingly, the DEN-induced liver tumors of the non-transgenic controls also showed up-regulation of endogenous BRE, suggesting that the BRE is important in liver carcinogenesis through its anti-apoptotic activity (Chui et al., 2010).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 17704801 | 2008 | BRE is an antiapoptotic protein in vivo and overexpressed in human hepatocellular carcinoma. | Chan BC et al |
| 15582573 | 2005 | BRE enhances in vivo growth of tumor cells. | Chan BC et al |
| 14565866 | 2003 | Expression of a conserved mouse stress-modulating gene, Bre: comparison with the human ortholog. | Ching AK et al |
| 20035718 | 2010 | BRE over-expression promotes growth of hepatocellular carcinoma. | Chui YL et al |
| 19214193 | 2009 | K63-specific deubiquitination by two JAMM/MPN+ complexes: BRISC-associated Brcc36 and proteasomal Poh1. | Cooper EM et al |
| 14636569 | 2003 | Regulation of BRCC, a holoenzyme complex containing BRCA1 and BRCA2, by a signalosome-like subunit and its role in DNA repair. | Dong Y et al |
| 17133111 | 2007 | Single nucleotide polymorphism array analysis of cancer. | Dutt A et al |
| 11030145 | 2000 | Caspase-8/FLICE functions as an executioner caspase in anticancer drug-induced apoptosis. | Engels IH et al |
| 19261748 | 2009 | MERIT40 facilitates BRCA1 localization and DNA damage repair. | Feng L et al |
| 9737713 | 1998 | BRE: a modulator of TNF-alpha action. | Gu C et al |
| 14685227 | 2003 | The International HapMap Project. | |
| 7826398 | 1995 | Identification of a brain- and reproductive-organs-specific gene responsive to DNA damage and retinoic acid. | Li L et al |
| 15465831 | 2004 | A death receptor-associated anti-apoptotic protein, BRE, inhibits mitochondrial apoptotic pathway. | Li Q et al |
| 11259452 | 2001 | Differential expression of a stress-modulating gene, BRE, in the adrenal gland, in adrenal neoplasia, and in abnormal adrenal tissues. | Miao J et al |
| 9501089 | 1998 | Two CD95 (APO-1/Fas) signaling pathways. | Scaffidi C et al |
| 19261746 | 2009 | MERIT40 controls BRCA1-Rap80 complex integrity and recruitment to DNA double-strand breaks. | Shao G et al |
| 17525341 | 2007 | RAP80 targets BRCA1 to specific ubiquitin structures at DNA damage sites. | Sobhian B et al |
| 19261749 | 2009 | NBA1, a new player in the Brca1 A complex, is required for DNA damage resistance and checkpoint control. | Wang B et al |
Other Information
Locus ID:
NCBI: 9577
MIM: 610497
HGNC: 1106
Ensembl: ENSG00000158019
Variants:
dbSNP: 9577
ClinVar: 9577
TCGA: ENSG00000158019
COSMIC: BABAM2
RNA/Proteins
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 30287910 | 2019 | Pharmacogenomics in epithelial ovarian cancer first-line treatment outcome: validation of GWAS-associated NRG3 rs1649942 and BRE rs7572644 variants in an independent cohort. | 5 |
| 30287910 | 2019 | Pharmacogenomics in epithelial ovarian cancer first-line treatment outcome: validation of GWAS-associated NRG3 rs1649942 and BRE rs7572644 variants in an independent cohort. | 5 |
| 28871137 | 2018 | C-terminal BRE overexpression in 11q23-rearranged and t(8;16) acute myeloid leukemia is caused by intragenic transcription initiation. | 2 |
| 29416040 | 2018 | BRE/BRCC45 regulates CDC25A stability by recruiting USP7 in response to DNA damage. | 23 |
| 28871137 | 2018 | C-terminal BRE overexpression in 11q23-rearranged and t(8;16) acute myeloid leukemia is caused by intragenic transcription initiation. | 2 |
| 29416040 | 2018 | BRE/BRCC45 regulates CDC25A stability by recruiting USP7 in response to DNA damage. | 23 |
| 27733185 | 2016 | Over-expression of the long non-coding RNA HOTTIP inhibits glioma cell growth by BRE. | 20 |
| 27733185 | 2016 | Over-expression of the long non-coding RNA HOTTIP inhibits glioma cell growth by BRE. | 20 |
| 22555662 | 2012 | Improved classification of MLL-AF9-positive acute myeloid leukemia patients based on BRE and EVI1 expression. | 7 |
| 22555662 | 2012 | Improved classification of MLL-AF9-positive acute myeloid leukemia patients based on BRE and EVI1 expression. | 7 |
| 21282113 | 2011 | NBA1/MERIT40 and BRE interaction is required for the integrity of two distinct deubiquitinating enzyme BRCC36-containing complexes. | 46 |
| 21937695 | 2011 | High BRE expression predicts favorable outcome in adult acute myeloid leukemia, in particular among MLL-AF9-positive patients. | 19 |
| 21282113 | 2011 | NBA1/MERIT40 and BRE interaction is required for the integrity of two distinct deubiquitinating enzyme BRCC36-containing complexes. | 46 |
| 21937695 | 2011 | High BRE expression predicts favorable outcome in adult acute myeloid leukemia, in particular among MLL-AF9-positive patients. | 19 |
| 19757177 | 2010 | Differential expression of a novel gene BRE (TNFRSF1A modulator/BRCC45) in response to stress and biological signals. | 3 |
Citation
Yiu-Loon Chui ; Kenneth Ka-Ho Lee ; John Yeuk-Hon Chan
BABAM2 (brain and reproductive organ-expressed (TNFRSF1A modulator))
Atlas Genet Cytogenet Oncol Haematol. 2010-06-01
Online version: http://atlasgeneticsoncology.org/gene/839/babam2
