AGAP2 (Centaurin , gamma1)
2007-05-01 Chan Chi Bun , Ye Keqiang AffiliationDepartment of Pathology, Laboratory Medicine, Emory University School of Medicine, Atlanta, USA
Identity
DNA/RNA
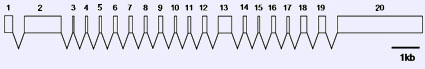
Description
Transcription
Pseudogene
Proteins
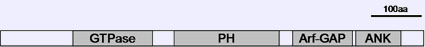
Description
Expression
Localisation
Function
PIKE-A is also a physiological interacting partner of protein kinase B (Akt). It was reported that PIKE-A specifically interacted with the regulatory domain and partial catalytic domain of activated Akt thorough its GTPase domain. Moreover, this interaction was guanine nucleotide dependent as the presence of GTPgammaS strongly stimulated their binding. Through interacting with PIKE-A, both basal and growth factor (e.g. EGF) stimulated Akt activity is greatly enhanced. This enhanced Akt activity is not triggered by uplifting PI3-kinase activity as PIKE-A neither interacts with PI3-kinase nor affects its activity. Instead, PIKE-A maintains and initiates Akt activation directly in both U87MG and LN-Z308 cells.
Overexpression of PIKE-A in U87MG glioblastoma cells promotes cell proliferation and enhances its invasion activity by stimulating Akt activity. In contrast, depletion of PIKE-A decreases U87MG viability upon staurosporine treatment via enhancing apoptosis.
Homology
Mutations
Germinal
Somatic
Implicated in
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15118108 | 2004 | PIKE-A is amplified in human cancers and prevents apoptosis by up-regulating Akt. | Ahn JY et al |
| 14761976 | 2004 | PIKE (phosphatidylinositol 3-kinase enhancer)-A GTPase stimulates Akt activity and mediates cellular invasion. | Ahn JY et al |
| 9192850 | 1997 | Transcript mapping in a 46-kb sequenced region at the core of 12q13.3 amplification in human cancers. | Elkahloun AG et al |
| 16263930 | 2005 | Phosphoinositol lipids bind to phosphatidylinositol 3 (PI3)-kinase enhancer GTPase and mediate its stimulatory effect on PI3-kinase and Akt signalings. | Hu Y et al |
| 16150119 | 2005 | Genetic alteration and expression of the phosphoinositol-3-kinase/Akt pathway genes PIK3CA and PIKE in human glioblastomas. | Knobbe CB et al |
| 17297440 | 2007 | PIKE-A is a proto-oncogene promoting cell growth, transformation and invasion. | Liu X et al |
| 8724849 | 1996 | Prediction of the coding sequences of unidentified human genes. V. The coding sequences of 40 new genes (KIAA0161-KIAA0200) deduced by analysis of cDNA clones from human cell line KG-1. | Nagase T et al |
| 16841086 | 2007 | Src-family tyrosine kinase fyn phosphorylates phosphatidylinositol 3-kinase enhancer-activating Akt, preventing its apoptotic cleavage and promoting cell survival. | Tang X et al |
| 16854451 | 2006 | Pike tyrosine phosphorylation regulates its apoptotic cleavage during programmed cell death. | Tang X et al |
| 12640130 | 2003 | GGAPs, a new family of bifunctional GTP-binding and GTPase-activating proteins. | Xia C et al |
Other Information
Locus ID:
NCBI: 116986
MIM: 605476
HGNC: 16921
Ensembl: ENSG00000135439
Variants:
dbSNP: 116986
ClinVar: 116986
TCGA: ENSG00000135439
COSMIC: AGAP2
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 34411567 | 2021 | PIKE-A promotes glioblastoma growth by driving PPP flux through increasing G6PD expression mediated by phosphorylation of STAT3. | 9 |
| 34411567 | 2021 | PIKE-A promotes glioblastoma growth by driving PPP flux through increasing G6PD expression mediated by phosphorylation of STAT3. | 9 |
| 30660615 | 2019 | Role of AGAP2 in the profibrogenic effects induced by TGFβ in LX-2 hepatic stellate cells. | 10 |
| 30674964 | 2019 | SP1 and RARα regulate AGAP2 expression in cancer. | 8 |
| 31386624 | 2019 | Exosomes derived from microRNA-199a-overexpressing mesenchymal stem cells inhibit glioma progression by down-regulating AGAP2. | 53 |
| 30660615 | 2019 | Role of AGAP2 in the profibrogenic effects induced by TGFβ in LX-2 hepatic stellate cells. | 10 |
| 30674964 | 2019 | SP1 and RARα regulate AGAP2 expression in cancer. | 8 |
| 31386624 | 2019 | Exosomes derived from microRNA-199a-overexpressing mesenchymal stem cells inhibit glioma progression by down-regulating AGAP2. | 53 |
| 28096359 | 2017 | α-Synuclein binds and sequesters PIKE-L into Lewy bodies, triggering dopaminergic cell death via AMPK hyperactivation. | 14 |
| 28368413 | 2017 | Co-amplification of phosphoinositide 3-kinase enhancer A and cyclin-dependent kinase 4 triggers glioblastoma progression. | 13 |
| 28096359 | 2017 | α-Synuclein binds and sequesters PIKE-L into Lewy bodies, triggering dopaminergic cell death via AMPK hyperactivation. | 14 |
| 28368413 | 2017 | Co-amplification of phosphoinositide 3-kinase enhancer A and cyclin-dependent kinase 4 triggers glioblastoma progression. | 13 |
| 26001218 | 2016 | Fyn-phosphorylated PIKE-A binds and inhibits AMPK signaling, blocking its tumor suppressive activity. | 21 |
| 26977005 | 2016 | Phosphoinositide-3-Kinase Enhancers, PIKEs: Their Biological Functions and Roles in Cancer. | 6 |
| 26001218 | 2016 | Fyn-phosphorylated PIKE-A binds and inhibits AMPK signaling, blocking its tumor suppressive activity. | 21 |
Citation
Chan Chi Bun ; Ye Keqiang
AGAP2 (Centaurin , gamma1)
Atlas Genet Cytogenet Oncol Haematol. 2007-05-01
Online version: http://atlasgeneticsoncology.org/gene/44037/agap2
