Identity
HGNC
LOCATION
6p21.33
LOCUSID
ALIAS
CL1C1,CLCNL1,G6,NCC27
FUSION GENES
DNA/RNA

Genomic organization of the CLIC1 gene on chromosome 6. There are 7 alternatively spliced and 2 unspliced variants of mRNA.
Description
The CLIC1 gene was mapped on chromosome 6 (6p21.33). It covers 9.183 kb, from position 31707540 to 31698358 bp in the minus strand orientation (NCBI 37, August 2010) and contains 6 exons. The gene is also known as CLIC1, DADB-110M10.1, G6, NCC27 or LOC1192, skersla. It has been described as chloride intracellular channel protein 1, hRNCC, p64CLCP, RNCC protein, OTTHUMP00000029133, OTTHUMP00000029137, OTTHUMP00000174486, chloride channel ABP, nuclear chloride ion channel 27, nuclear chloride ion channel protein, regulatory nuclear chloride ion channel protein.
Transcription
Ten separate gt-ag or gc-ag introns can be found within CLIC1. The mRNAs produced by its transcription: 7 alternatively spliced and 2 unspliced variants (see diagram above). These mRNAs appear to differ by truncation of the 5 end, truncation of the 3 end, overlapping exons with different boundaries, and splicing rather than retention of 3 introns. Research shows there are 2 non-overlapping alternative last exons, 4 probable alternative promoters, and 5 validated alternative polyadenylation sites. A translated product, an upstream open reading frame (uORF), reduces the efficacy of translation by initiating at an AUG upstream of the main open reading frame.
Pseudogene
According to the NCBI and HGNC database, there is a pseudogene of CLIC1 within the human known as CLIC1P1 (chloride intracellular channel 1 pseudogene 1). CLIC1P1 was mapped to chromosome 12q24.31 (121352207-121352926). Two other pseudogenes of CLIC1, LOC401864 and LOC100420638, were also recorded in the NCBI database. LOC401864 and LOC100420638 were located on chromosome 16q24.1 (86468450-86469104) and 17 (63639278-63640303), respectively.
Proteins
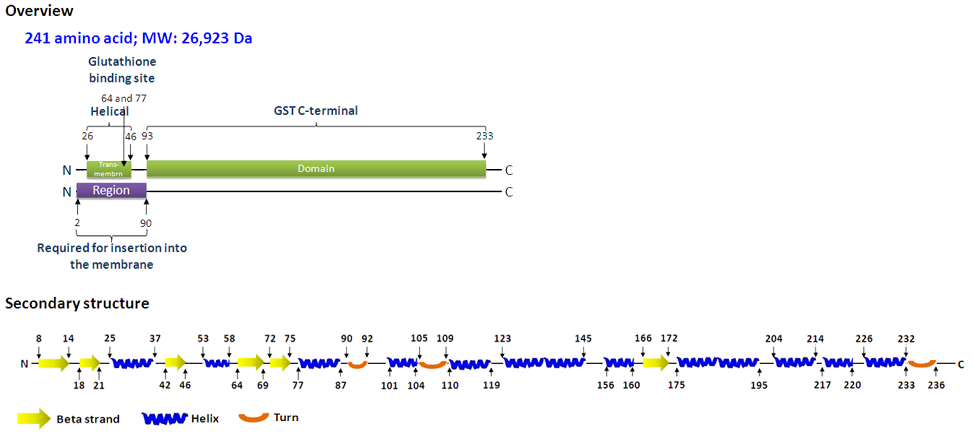
Description
CLIC1, also known as NCC27, is a member of the CLIC family. The family is defined by a C-terminal core segment of 230 amino acids, which has significant structural homology with glutathione-S-transferase (Harrop et al., 2001), and contains seven members, including CLIC1, CLIC2, CLIC3, CLIC4, CLIC5, p64, and parchorin. CLIC1 functions as a chloride channel, much like other CLIC family members, and possesses the biological activities needed to regulate the cell volume and acidity of intracellular organelles. CLIC1 exists in cells as an integral membrane protein as well as a soluble cytoplasm protein. These phenomena indicate that CLIC1 might cycle between membrane-inserted and soluble forms (Tulk et al., 2002).
Expression
CLIC1 can be expressed in various cell types. Expression is prominent in the heart, placenta, liver, kidney and pancreas (Berryman and Bretscher, 2000). To find the protein expression of various cell types and normal/cancer tissues, please refer to the database, The Human Protein Atlas.
Localisation
The protein localizes in the nucleus, nucleus membrane, cytoplasm, and cell membrane. Protein generally exists in the nucleus including the nuclear membrane and smaller amounts exist in the cytoplasm as well as the plasma membrane (Valenzuela et al., 1997; Berryman and Bretscher, 2000; Harrop et al., 2001). The Human Protein Atlas database reveals that CLIC1 has weak to strong immunofluorescence staining in various cell types in cytoplasm.
Function
1. Ion channels
The CLIC family of proteins exhibits chloride channel activity when reconstituted in phospholipid vesicles. Due to its ability to spontaneously insert into preformed membranes, CLIC1 appears to cycle between membrane protein and soluble cytoplasmic protein forms, and sometimes functions as an anion-selective channel (chloride ion channels) (Tulk et al., 2000; Tulk et al., 2002; Berryman and Bretscher, 2000). Chloride channels are a diverse group of proteins that regulate fundamental cellular processes including cell volume, stabilization of cell membrane potential, transepithelial transport, maintenance of intracellular pH. In previous studies, CLIC1 ion channels were shown to be strongly and reversibly inhibited by cytosolic F-actin in the absence of other proteins. This effect can be reversed using cytochalasin, which disrupts F-actin. This represents a new possibility for which CLIC1 and other actin-regulated membrane CLICs could be used to modify solute transport at key stages during cellular events such as apoptosis, cell movement, cell-volume regulation, as well as cell and organelle division and fusion (Singh et al., 2007; Fanucchi et al., 2008; Stoychev et al., 2009). In an oxidized state, the crystal structure of CLIC1 drastically changes as a large hydrophobic surface is exposed, and forms a dimer interface. The oxidized CLIC1 dimer maintains its ability to form chloride ion channels in artificial bilayers and vesicles, whereas a reducing environment would inhibit the formation of ion channels by CLIC1 (Littler et al., 2004). Research suggest that oxidation of monomeric CLIC1, in the presence of membranes, promotes its insertion into the bilayer more effectively than the oxidized CLIC1 dimer (Goodchild et al., 2009). The crystal structure of CLIC1 classifies it as a member of the glutathione S-transferase superfamily. This detail helps explain why CLICs can exist in a water-soluble state, and also insert into membranes to form ion channels (Dulhunty et al., 2000; Cromer et al., 2002). As an ion channel, CLIC1 is likely to consist of a tetrameric assembly of subunits, and despite its size and unusual properties, there are indications of its ability to form an ion channel in the absence of any other ancillary proteins (Warton et al., 2002). The structure of CLIC1 with glutathione reveals that glutathione occupies the redox-active site, which is adjacent to an open, elongated slot lined with basic residues. Integration of CLIC1 into the membrane would require major structural changes, most likely within the N-domain (residues 1-90), with its transmembrane helix arising from residues near the redox-active site. The structure indicates that CLIC1 is likely to be controlled by redox-dependent processes (Harrop et al., 2001). In addition, CLIC1 translocates from the cytosol to the plasma membrane after microglial activation where it promotes chloride conductance. The charge generated by the active NADPH oxidase is balanced by the resulting anionic current. Removing the excess charge supports superoxide generation by the enzyme. CLIC1 exhibits an ability to act as both a second messenger and an executer (Averaimo et al., 2010).
2. Inflammation
At the cellular level, Alzheimers disease is characterized as the accumulation of Aβ in neuritic plaques which have been infiltrated by astrocytes and reactive microglia. A decrease in the expression of CLIC1 could reverse this inflammation if the decrease was used to prevent pro-inflammatory TNF-a and neurotoxic products caused by Aβ-stimulated microglial cells (Novarino et al., 2004).
3. Apoptosis
A specific blocker may be used to reduce CLIC1 chloride conductance, and thereby prevent neural apoptosis in neurons co-cultured with Aβ-treated microglia. In doing so, the cellular process of apoptosis could be controlled, giving hope to possibly control diseases caused by the apoptosis of particular cells (Novarino et al., 2004).
4. Motility
CLIC1 overexpression can promote cell motility and invasion of gallbladder carcinoma cells (GBC-SD18L), whereas interference of CLIC1s RNA can significantly decrease the cell motility and invasive potency of GBC-SD18L in vitro (Wang et al., 2009). Additionally, by simply reducing the CLIC1 expression, the migration ability of endothelial cells can be reduced accordingly (Tung and Kitajewski, 2010).
5. Cell cycle regulation
Cl- ion channel blockers, known to block CLIC1, were shown to inhibit Chinese hamster ovary (CHO-K1) cells in the G2/M stage of the cell cycle. This is the stage in which the ion channel is selectively expressed on the plasma membrane. The prevention of CLIC1-mediated changes in cell volume may prevent cells from completing mitosis, thereby preventing the cells from physically dividing and/or the dissolution of the nuclear envelope. To the same effect, disruption of the CLIC1 function in ionic Cl- regulation may prevent other downstream events, in which case cell cycle checkpoint mechanisms prevent the cell from completing mitosis (Valenzuela et al., 2000).
The CLIC family of proteins exhibits chloride channel activity when reconstituted in phospholipid vesicles. Due to its ability to spontaneously insert into preformed membranes, CLIC1 appears to cycle between membrane protein and soluble cytoplasmic protein forms, and sometimes functions as an anion-selective channel (chloride ion channels) (Tulk et al., 2000; Tulk et al., 2002; Berryman and Bretscher, 2000). Chloride channels are a diverse group of proteins that regulate fundamental cellular processes including cell volume, stabilization of cell membrane potential, transepithelial transport, maintenance of intracellular pH. In previous studies, CLIC1 ion channels were shown to be strongly and reversibly inhibited by cytosolic F-actin in the absence of other proteins. This effect can be reversed using cytochalasin, which disrupts F-actin. This represents a new possibility for which CLIC1 and other actin-regulated membrane CLICs could be used to modify solute transport at key stages during cellular events such as apoptosis, cell movement, cell-volume regulation, as well as cell and organelle division and fusion (Singh et al., 2007; Fanucchi et al., 2008; Stoychev et al., 2009). In an oxidized state, the crystal structure of CLIC1 drastically changes as a large hydrophobic surface is exposed, and forms a dimer interface. The oxidized CLIC1 dimer maintains its ability to form chloride ion channels in artificial bilayers and vesicles, whereas a reducing environment would inhibit the formation of ion channels by CLIC1 (Littler et al., 2004). Research suggest that oxidation of monomeric CLIC1, in the presence of membranes, promotes its insertion into the bilayer more effectively than the oxidized CLIC1 dimer (Goodchild et al., 2009). The crystal structure of CLIC1 classifies it as a member of the glutathione S-transferase superfamily. This detail helps explain why CLICs can exist in a water-soluble state, and also insert into membranes to form ion channels (Dulhunty et al., 2000; Cromer et al., 2002). As an ion channel, CLIC1 is likely to consist of a tetrameric assembly of subunits, and despite its size and unusual properties, there are indications of its ability to form an ion channel in the absence of any other ancillary proteins (Warton et al., 2002). The structure of CLIC1 with glutathione reveals that glutathione occupies the redox-active site, which is adjacent to an open, elongated slot lined with basic residues. Integration of CLIC1 into the membrane would require major structural changes, most likely within the N-domain (residues 1-90), with its transmembrane helix arising from residues near the redox-active site. The structure indicates that CLIC1 is likely to be controlled by redox-dependent processes (Harrop et al., 2001). In addition, CLIC1 translocates from the cytosol to the plasma membrane after microglial activation where it promotes chloride conductance. The charge generated by the active NADPH oxidase is balanced by the resulting anionic current. Removing the excess charge supports superoxide generation by the enzyme. CLIC1 exhibits an ability to act as both a second messenger and an executer (Averaimo et al., 2010).
2. Inflammation
At the cellular level, Alzheimers disease is characterized as the accumulation of Aβ in neuritic plaques which have been infiltrated by astrocytes and reactive microglia. A decrease in the expression of CLIC1 could reverse this inflammation if the decrease was used to prevent pro-inflammatory TNF-a and neurotoxic products caused by Aβ-stimulated microglial cells (Novarino et al., 2004).
3. Apoptosis
A specific blocker may be used to reduce CLIC1 chloride conductance, and thereby prevent neural apoptosis in neurons co-cultured with Aβ-treated microglia. In doing so, the cellular process of apoptosis could be controlled, giving hope to possibly control diseases caused by the apoptosis of particular cells (Novarino et al., 2004).
4. Motility
CLIC1 overexpression can promote cell motility and invasion of gallbladder carcinoma cells (GBC-SD18L), whereas interference of CLIC1s RNA can significantly decrease the cell motility and invasive potency of GBC-SD18L in vitro (Wang et al., 2009). Additionally, by simply reducing the CLIC1 expression, the migration ability of endothelial cells can be reduced accordingly (Tung and Kitajewski, 2010).
5. Cell cycle regulation
Cl- ion channel blockers, known to block CLIC1, were shown to inhibit Chinese hamster ovary (CHO-K1) cells in the G2/M stage of the cell cycle. This is the stage in which the ion channel is selectively expressed on the plasma membrane. The prevention of CLIC1-mediated changes in cell volume may prevent cells from completing mitosis, thereby preventing the cells from physically dividing and/or the dissolution of the nuclear envelope. To the same effect, disruption of the CLIC1 function in ionic Cl- regulation may prevent other downstream events, in which case cell cycle checkpoint mechanisms prevent the cell from completing mitosis (Valenzuela et al., 2000).
Homology
CLIC1, CLIC2, CLIC3, CLIC5 and CLIC6.
Implicated in
Entity name
Gastric carcinoma
Note
The CLIC1 gene expression in tumor tissues was 1.95 times that of adjacent noncancerous mucosa. These elevated levels are attributed to lymph node metastasis, lymphatic invasion, perineural invasion, and pathological staging. Also, the 5-year survival rate of the low CLIC1 expression group was 1.72 times that of the high expression group. Results represent CLIC1s potential as an effective prognostic marker for gastric cancer (Chen et al., 2006).
Entity name
Hepatocellular carcinoma
Note
An observed overexpression of CLIC1 (60% or 27/45) in high proportions of 45 patient HCC tumors indicates that CLIC1 is also a potential marker for HCC (Huang et al., 2003).
Entity name
Nasopharyngeal carcinoma
Note
The plasma levels of CLIC1 among NPC patients were significantly higher than those in healthy controls, as presented by sandwich ELISA. 75% of NPC tissue specimens showed positive CLIC1 staining by IHC. NPC was successfully discriminated from the benign healthy control group with a sensitivity of 63% and a specificity of 77%. Results indicate that CLIC1 is a potential plasma tumor marker for NPC (Chang et al., 2009).
Entity name
Laryngeal cancer
Note
Suppressing of the CLIC1 gene allows for the acquisition of a radio-resistant phenotype of laryngeal cancer cells via inhibition of ROS production. This indicates that CLIC1 is an important candidate molecule for radiotherapy in radio-resistant laryngeal cells (Kim et al., 2010).
Entity name
Lung adenocarcinoma
Note
Among 103 paraffin sections of lung adenocarcinoma tissue samples, the CLIC1 expression was strongly positive in 49 cases (47.6%). This gene expression significantly correlates with the T staging of tumors (p = 0.029). Univariate analysis indicated that the patients ECOG score, T staging, N staging, TNM staging, and CLIC1 expression correlated with prognosis (p = 0.031, 0.001, 0.011, 0.013, and
Entity name
Psoriasis
Note
CLIC1 is one of several genes that can act as genomic classifiers in response to treatment of psoriasis with Alefacept. (Suárez-Fariñas et al., 2010).
Entity name
Alzheimers disease
Note
Research suggests that the blockade of CLIC1 stimulates Aβ phagocytosis in mononuclear phagocytes while inhibiting the induction of iNOS. Results point to CLIC1 as a possible therapeutic target in Alzheimers disease (Paradisi et al., 2008).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 20385134 | 2010 | Chloride intracellular channel 1 (CLIC1): Sensor and effector during oxidative stress. | Averaimo S et al |
| 10793131 | 2000 | Identification of a novel member of the chloride intracellular channel gene family (CLIC5) that associates with the actin cytoskeleton of placental microvilli. | Berryman M et al |
| 19845400 | 2009 | Cell secretome analysis using hollow fiber culture system leads to the discovery of CLIC1 protein as a novel plasma marker for nasopharyngeal carcinoma. | Chang YH et al |
| 17154271 | 2007 | Overexpression of CLIC1 in human gastric carcinoma and its clinicopathological significance. | Chen CD et al |
| 12202911 | 2002 | From glutathione transferase to pore in a CLIC. | Cromer BA et al |
| 18850721 | 2008 | Formation of an unfolding intermediate state of soluble chloride intracellular channel protein CLIC1 at acidic pH. | Fanucchi S et al |
| 19387633 | 2009 | Oxidation promotes insertion of the CLIC1 chloride intracellular channel into the membrane. | Goodchild SC et al |
| 11551966 | 2001 | Crystal structure of a soluble form of the intracellular chloride ion channel CLIC1 (NCC27) at 1.4-A resolution. | Harrop SJ et al |
| 14985104 | 2004 | Diverse cellular transformation capability of overexpressed genes in human hepatocellular carcinoma. | Huang JS et al |
| 20461716 | 2010 | Chloride intracellular channel 1 identified using proteomic analysis plays an important role in the radiosensitivity of HEp-2 cells via reactive oxygen species production. | Kim JS et al |
| 14613939 | 2004 | The intracellular chloride ion channel protein CLIC1 undergoes a redox-controlled structural transition. | Littler DR et al |
| 15190104 | 2004 | Involvement of the intracellular ion channel CLIC1 in microglia-mediated beta-amyloid-induced neurotoxicity. | Novarino G et al |
| 18028448 | 2007 | Functional reconstitution of mammalian 'chloride intracellular channels' CLIC1, CLIC4 and CLIC5 reveals differential regulation by cytoskeletal actin. | Singh H et al |
| 19650640 | 2009 | Structural dynamics of soluble chloride intracellular channel protein CLIC1 examined by amide hydrogen-deuterium exchange mass spectrometry. | Stoychev SH et al |
| 20152045 | 2010 | Personalized medicine in psoriasis: developing a genomic classifier to predict histological response to Alefacept. | Suárez-Fariñas M et al |
| 10834939 | 2000 | Functional characterization of the NCC27 nuclear protein in stable transfected CHO-K1 cells. | Tonini R et al |
| 11940526 | 2002 | CLIC1 inserts from the aqueous phase into phospholipid membranes, where it functions as an anion channel. | Tulk BM et al |
| 10874038 | 2000 | CLIC-1 functions as a chloride channel when expressed and purified from bacteria. | Tulk BM et al |
| 21040583 | 2010 | Chloride intracellular channel 1 functions in endothelial cell growth and migration. | Tung JJ et al |
| 11195932 | 2000 | The nuclear chloride ion channel NCC27 is involved in regulation of the cell cycle. | Valenzuela SM et al |
| 19299076 | 2009 | Identification of metastasis-associated proteins involved in gallbladder carcinoma metastasis by proteomic analysis and functional exploration of chloride intracellular channel 1. | Wang JW et al |
| 21858536 | 2011 | The expression and clinical significance of CLIC1 and HSP27 in lung adenocarcinoma. | Wang W et al |
| 11978800 | 2002 | Recombinant CLIC1 (NCC27) assembles in lipid bilayers via a pH-dependent two-state process to form chloride ion channels with identical characteristics to those observed in Chinese hamster ovary cells expressing CLIC1. | Warton K et al |
Other Information
Locus ID:
NCBI: 1192
MIM: 602872
HGNC: 2062
Ensembl: ENSG00000213719
Variants:
dbSNP: 1192
ClinVar: 1192
TCGA: ENSG00000213719
COSMIC: CLIC1
RNA/Proteins
Expression (GTEx)
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37104099 | 2024 | CLIC1-mediated autophagy confers resistance to DDP in gastric cancer. | 3 |
| 38257829 | 2024 | Chloride Intracellular Channel Protein 1 (CLIC1) Is a Critical Host Cellular Factor for Influenza A Virus Replication. | 0 |
| 37104099 | 2024 | CLIC1-mediated autophagy confers resistance to DDP in gastric cancer. | 3 |
| 38257829 | 2024 | Chloride Intracellular Channel Protein 1 (CLIC1) Is a Critical Host Cellular Factor for Influenza A Virus Replication. | 0 |
| 34240838 | 2022 | CLIC1 promotes the progression of cervical cancer through the PTEN/PI3K/AKT pathway. | 1 |
| 34969734 | 2022 | Chloride Intracellular Channel 1 Expression Is Associated With Poor Prognosis of Lung Adenocarcinoma. | 5 |
| 35722791 | 2022 | CLIC1 Drives Angiogenesis in Hepatocellular Carcinoma by Modulating VEGFA. | 3 |
| 36181109 | 2022 | Comprehensive analysis of clinical prognosis and CLIC1 immune invasion in lung adenocarcinoma. | 1 |
| 34240838 | 2022 | CLIC1 promotes the progression of cervical cancer through the PTEN/PI3K/AKT pathway. | 1 |
| 34969734 | 2022 | Chloride Intracellular Channel 1 Expression Is Associated With Poor Prognosis of Lung Adenocarcinoma. | 5 |
| 35722791 | 2022 | CLIC1 Drives Angiogenesis in Hepatocellular Carcinoma by Modulating VEGFA. | 3 |
| 36181109 | 2022 | Comprehensive analysis of clinical prognosis and CLIC1 immune invasion in lung adenocarcinoma. | 1 |
| 32586242 | 2021 | A Knock-Down Cell-Based Study for the Functional Analysis of Chloride Intracellular Channel 1 (CLIC1): Integrated Proteomics and Microarray Study. | 0 |
| 32656583 | 2021 | CLIC1 knockout inhibits invasion and migration of gastric cancer by upregulating AMOT-p130 expression. | 8 |
| 32841520 | 2021 | Exosomal CLIC1 released by CLL promotes HUVECs angiogenesis by regulating ITGβ1-MAPK/ERK axis. | 6 |
Citation
Pao-Chi Liao ; Ying-Hwa Chang
CLIC1 (chloride intracellular channel 1)
Atlas Genet Cytogenet Oncol Haematol. 2011-11-01
Online version: http://atlasgeneticsoncology.org/gene/50543/clic1
