CRTC1 (CREB regulated transcription coactivator 1)
2006-05-01 Afrouz Behboudi , Gâran Stenman AffiliationDepartment of Clinical Genetics, Institute of Biomedicine, Gâteborg University, Box 440, 405 30 Gâteborg, Sweden
Identity
HGNC
LOCATION
19p13.11
LOCUSID
ALIAS
MAML2,MECT1,Mam-2,TORC-1,TORC1,WAMTP1
FUSION GENES
DNA/RNA
Description
Spans about 94 kb and includes 14 to 16 exons
Transcription
two RNA variants of 2505 and 2342 bp, respectively.
Proteins
Note
634 amino acids; 67300 Da
Description
Transcriptional coactivator for CREB1.
Expression
Expressed in a restricted number of tissues including fetal brain and liver and adult heart, skeletal muscle, liver and salivary gland.
Localisation
Nucleus.
Function
Interacts through its N-terminal CREB-binding domain with the bZIP domain of CREB1; Binds to CREB1 as a homotetramer. Transcriptional coactivator for CREB1, which activates transcription through both consensus and variant cAMP response element (CRE) sites, leading to activation of CREB1 target genes. Does not appear to modulate CREB1 DNA-binding activity but enhances the interaction of CREB1 with TAF4/TAFII-130.
Implicated in
Entity name
Mucoepidermoid carcinoma (most common type of malignant salivary gland tumor; second most frequent lung tumor of bronchial gland origin); benign Warthins tumor; clear cell hidradenoma of the skin (CCH; a.k.a. nodular hidradenomas or eccrine acrospiromas). all sharing a t(11; 19) (q21; p13)
Cytogenetics
t(11;19)(q21;p13).
Hybrid gene
CRTC1-MAML2. Exon 1 of MECT1 fused to exons 2-5 of MAML2.
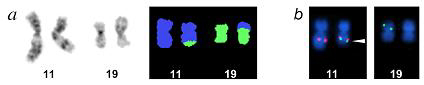
Fig1a (a) Partial G-banded and SKY karyotypes showing the t(11;19) in a mucoepidermoid carcinoma (MEC) case. (b) Dual-color FISH experiment of the MEC using BAC 697H10 (MAML2, red signal) and cosmid LLNLR 255A4 (CRTC1, green signal) as probes. Note the presence of the CRTC1-MAML2 fusion gene on the der(11) (fused red green signals marked by an arrowhead).
Reprinted partially from Publication Exp Cell Res 292, Enlund F., Behboudi A., Andren Y., Oberg C., Lendahl U., Mark J., Stenman G., Altered Notch signaling resulting from expression of a WAMTP1-MAML2 gene fusion in mucoepidermoid carcinomas and benign Warthins tumors, 21-28, Copyright (2004), with permission from Elsevier.
Fusion protein
MECT1-MAML2; in the fusion protein the first 171 aa including the basic domain of MAML2 are replaced by 42 aa of MECT1; there are no sequence similarities in the N-terminal domains of MAML2 and MECT1; the fusion protein activates transcription of the Notch target gene HES1 independently of both Notch ligand and CSL.
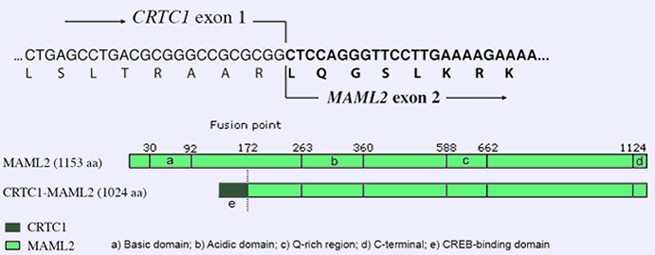
Top: Nucleotide and deduced amino acid sequences of the CRTC1-MAML2 breakpoint junction.
Bottom: Schematic representation of the MAML2 protein and the CRTC1-MAML2 fusion protein. In the fusion protein the N-terminal Notch-binding domain of MAML2 is replaced by the CREB-binding domain of CRTC1.
Reprinted partially from publication above mentioned
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 16444749 | 2006 | Molecular classification of mucoepidermoid carcinomas-prognostic significance of the MECT1-MAML2 fusion oncogene. | Behboudi A et al |
| 14536081 | 2003 | TORCs: transducers of regulated CREB activity. | Conkright MD et al |
| 14720503 | 2004 | Altered Notch signaling resulting from expression of a WAMTP1-MAML2 gene fusion in mucoepidermoid carcinomas and benign Warthin's tumors. | Enlund F et al |
| 14506290 | 2003 | Identification of a family of cAMP response element-binding protein coactivators by genome-scale functional analysis in mammalian cells. | Iourgenko V et al |
| 15826837 | 2005 | Fusion oncogenes and tumor type specificity--insights from salivary gland tumors. | Stenman G et al |
| 12539049 | 2003 | t(11;19)(q21;p13) translocation in mucoepidermoid carcinoma creates a novel fusion product that disrupts a Notch signaling pathway. | Tonon G et al |
| 15961999 | 2005 | Transforming activity of MECT1-MAML2 fusion oncoprotein is mediated by constitutive CREB activation. | Wu L et al |
Other Information
Locus ID:
NCBI: 23373
MIM: 607536
HGNC: 16062
Ensembl: ENSG00000105662
Variants:
dbSNP: 23373
ClinVar: 23373
TCGA: ENSG00000105662
COSMIC: CRTC1
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000105662 | ENST00000321949 | Q6UUV9 |
| ENSG00000105662 | ENST00000338797 | Q6UUV9 |
| ENSG00000105662 | ENST00000594658 | M0QX46 |
| ENSG00000105662 | ENST00000601916 | M0QXN6 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36675234 | 2023 | Establishment of Mucoepidermoid Carcinoma Cell Lines from Surgical and Recurrence Biopsy Specimens. | 0 |
| 36675234 | 2023 | Establishment of Mucoepidermoid Carcinoma Cell Lines from Surgical and Recurrence Biopsy Specimens. | 0 |
| 34657306 | 2022 | Salivary mucoepidermoid carcinoma: histological variants, grading systems, CRTC1/3-MAML2 fusions, and clinicopathological features. | 8 |
| 35993578 | 2022 | Cutaneous Melanocytic Tumor With CRTC1::TRIM11 Translocation : An Emerging Entity Analyzed in a Series of 41 Cases. | 3 |
| 36115844 | 2022 | Oncogenic RAS commandeers amino acid sensing machinery to aberrantly activate mTORC1 in multiple myeloma. | 7 |
| 34657306 | 2022 | Salivary mucoepidermoid carcinoma: histological variants, grading systems, CRTC1/3-MAML2 fusions, and clinicopathological features. | 8 |
| 35993578 | 2022 | Cutaneous Melanocytic Tumor With CRTC1::TRIM11 Translocation : An Emerging Entity Analyzed in a Series of 41 Cases. | 3 |
| 36115844 | 2022 | Oncogenic RAS commandeers amino acid sensing machinery to aberrantly activate mTORC1 in multiple myeloma. | 7 |
| 33473104 | 2021 | Targeting Notch and EGFR signaling in human mucoepidermoid carcinoma. | 10 |
| 33556406 | 2021 | Structural Insights into the Interaction Between CRTCs and 14-3-3. | 5 |
| 33561653 | 2021 | mTORC1 silencing during intestinal epithelial Caco-2 cell differentiation is mediated by the activation of the AMPK/TSC2 pathway. | 2 |
| 33830080 | 2021 | The CRTC1-MAML2 fusion is the major oncogenic driver in mucoepidermoid carcinoma. | 25 |
| 33473104 | 2021 | Targeting Notch and EGFR signaling in human mucoepidermoid carcinoma. | 10 |
| 33556406 | 2021 | Structural Insights into the Interaction Between CRTCs and 14-3-3. | 5 |
| 33561653 | 2021 | mTORC1 silencing during intestinal epithelial Caco-2 cell differentiation is mediated by the activation of the AMPK/TSC2 pathway. | 2 |
Citation
Afrouz Behboudi ; Gâran Stenman
CRTC1 (CREB regulated transcription coactivator 1)
Atlas Genet Cytogenet Oncol Haematol. 2006-05-01
Online version: http://atlasgeneticsoncology.org/gene/471/crtc1
Historical Card
2003-07-01 CRTC1 (CREB regulated transcription coactivator 1) by Goran Stenman Affiliation
Lundberg Laboratory for Cancer Research, Department of Pathology, Goteborg University, Sahlgrenska University Hospital, SE-413 45 Goteborg, Sweden
