ENO1 (Enolase 1, (alpha))
2009-08-01 Bogusz Trojanowicz , Cuong Hoang-Vu , Carsten Sekulla AffiliationIdentity
HGNC
LOCATION
1p36.23
LOCUSID
ALIAS
ENO1L1,HEL-S-17,MPB1,NNE,PPH
FUSION GENES
DNA/RNA
Note
Alpha-Enolase (ENO1, alpha enolase, non-neuronal enolase) is one of the three enolase enzymes, expressed in a wide variety of tissues. The other two enolase genes, ENO2 and ENO3, encode gamma (neuron-specific) and beta (muscle-specific) enolase, respectively. The active enolase enzymes exist as homodimers of non-covalently bound subunits. Each alpha, beta or gamma subunit is encoded by separate genes. The genomic organisation of ENO1 gene is identical with that of human gamma-enolase gene. All the coding exons have exactly the same length and introns occur at analogous positions.
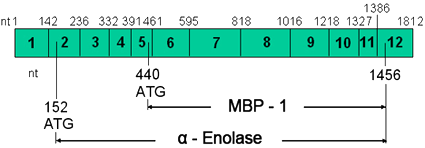
Structure of ENO1 mRNA ; note that nucleotides number (nt), exon positions (1-12) and two translation initiation sites (ATG) are labelled.
Description
The ENO1 gene consists of 12 exons distributed over 17718 bp of genomic DNA. Single alpha-enolase transcript contains two translation initiation positions and encodes two structurally and functionally distinct proteins, alpha-enolase enzyme and MYC promotor-binding protein (MBP-1).
Transcription
Transcription start sites of ENO1 gene are heterogeneous and spread over 38-bp region located at 116 bp upstream from the initiation codon ATG. These multiple start sites of transcription in ENO1 gene are consistent with lack of a canonical TATA box, usually found at the position 19-27 bp upstream of the cap sites. It is worth to notice that promoter of ENO1 gene contains two perfect Myc-Max binding motifs CACGTC. Other regulatory sites found in the 5-flanking region of ENO1 gene include AP1 (T[T/G]AGTCA), AP2 (CCCCAGGC), AP3 (GGGTGTGGAAAG), AP4 (CAGCTGTGG), AP5 (CTGTGGAATG), ATF/CREB ([T/G][A/T]CGTCA), C2 (CATGTG), CTF/NF1 (TGGCTNNNAGCCAA), E2AE-C beta (TGGGAATT), E2F (TTTCGCGC), E4TF1 (GGAAGTG), EF-C (GTTGCNNGGCAAC), MLTF/USF (GGTCACGTGGCC), Ig octamer (ATTTGCAT), PEA2 (GACCGCA), SP1 (GGGCGG), CACCC (may function as CAAT boxes) and viral core (GTGG[A/T][A/T][A/T]G).
Pseudogene
A pseudogene has been identified that is located on the other arm of the same chromosome (provided by RefSeq).
Proteins
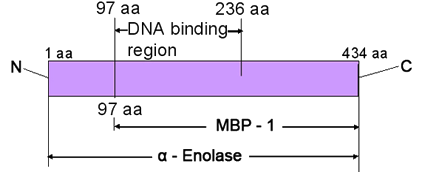
Structure of ENO1/MBP-1 protein; N and C termini, and amino acid (aa) positions are labelled.
Description
Alpha-Enolase (ENO1), like two other isoenzymes (gamma-ENO2 and beta-ENO3), is made up of two identical (homodimer), non-covalently bound alpha alpha subunits; alpha-Enolase is resolved during 1D-PAGE as a protein with molecular weight of about 48kDa (434 amino acids). It was demonstrated that in brain and neurons, specific enolases may exist as heterodimers, such as alpha alpha, alpha beta, beta beta, alpha gamma and gamma gamma. The proportions of isoenzymes alpha alpha, alpha beta and beta beta change in heart and muscle during embryonic development. In both mentioned tissues, isoform alpha was found predominantly in fetus. In adult heart this subunit is replaced by types alpha beta and beta beta, and in muscle by type beta beta. In human adult brain tissues, apha-type and gamma-type enolase subunits are present at similar concentrations.
Two identical subunits of alpha-Enolase facilitate each other in an antiparallel fashion. Each subunit is made up of two distinct domains: N-terminal domain, consisting of three beta-sheets and four alpha-helices (beta3 alpha4 topology), and larger C-terminal domain with eightfold alpha beta barrel structure with beta beta alpha alpha(beta alpha)6 topology. This domain contains two beta-sheets at the beginning, followed by two alpha-helices and ends with a barrel made up of alternating beta-sheets and alpha-helices (beta - sheets are surrounded by the alpha - helices). The N-terminal of one subunit contacts the C-terminal of the second in such way, that glutamic acid at position 20 (Glu20) forms an ionic pair with arginine at position 414 (Arg414).
Alternatively translated product of ENO1 gene, called MBP-1 (MYC promotor-binding protein), is expressed as a 37kDa (338 amino acids) protein and does not posses the enolase enzyme activity.
Two identical subunits of alpha-Enolase facilitate each other in an antiparallel fashion. Each subunit is made up of two distinct domains: N-terminal domain, consisting of three beta-sheets and four alpha-helices (beta3 alpha4 topology), and larger C-terminal domain with eightfold alpha beta barrel structure with beta beta alpha alpha(beta alpha)6 topology. This domain contains two beta-sheets at the beginning, followed by two alpha-helices and ends with a barrel made up of alternating beta-sheets and alpha-helices (beta - sheets are surrounded by the alpha - helices). The N-terminal of one subunit contacts the C-terminal of the second in such way, that glutamic acid at position 20 (Glu20) forms an ionic pair with arginine at position 414 (Arg414).
Alternatively translated product of ENO1 gene, called MBP-1 (MYC promotor-binding protein), is expressed as a 37kDa (338 amino acids) protein and does not posses the enolase enzyme activity.
Expression
Apha-Enolase is widely expressed in variety of tissues including liver, brain, kidney, spleen, adipose as well as thyroid. In comparison with gamma-type subunit found only in neurons, type alpha subunit was also detected in astrocytes, ependymal cells, capillary endothelial cells, Schwann cells and arachnoidal endothelial cells.
Localisation
Alpha-Enolase is most abundantly found in cytoplasm and also on the cell surface. MBP-1 is localised in the nucleus.
Function
Enolase enzymes (2-phospho-D-glycerate hydrolases) catalyse the dehydration of 2-phospho-D-glycerate (PGA) to phosphoenolopyruvate (PEP) in Emden Mayerhoff-Parnas glycolytic pathway (catabolic direction). In anabolic pathway (reverse reaction) during gluconeogenesis, the same enzyme catalyses hydration of PEP to PGA (hence it is called phosphopyruvate hydratase). Metal ions are cofactors impairing the increase of enolase activity; hence it is also called metal-activated metalloenzyme. Magnesium is a natural cofactor causing the highest activity. The relative activation strength profile of metal ions involved in the enzyme activity appears in the following rank of order Mg2+ > Zn2+ > Mn2+ > Fe(II)2+ > Cd2+ > Co2+, Ni2+, Sm3+, Tb3+ and most other divalent metal ions. In reaction catalyzed by enolases, the alpha-proton from a carbon adjacent to a carboxylate group of PGA, is abstracted, and PGA is conversed to enolate anion intermediate. This intermediate is further processed in a variety of chemical reactions, including racemization, cycloisomerization and beta-elimination of either water or ammonia.
The smaller product of ENO1 gene, MBP-1, is known as c-myc binding protein and negative regulator of its expression. C-myc is a DNA-binding phosphoprotein and a key regulator of cell behaviour. Many of c-myc targeting pathways are deregulated in cancer cells and contribute to its enhanced expression. There are four c-myc promoters, designated as P0, P1, P2 and P3, although in normal and cancer cells most mRNAs initiate at the P2 promoter. MBP-1 binds in a region +123 to +153 relative to the c-myc P2 promoter and probably by preventing the formation of a transcription initiation complex, decrease c-myc promoter activity. Hence MBP-1 is considered as tumor suppressor.
ENO1 protein was also found as a structural component of the eye lenses and was designated as tau-crystalin. ENO1 enzyme and tau-crystalin are the products of the same gene. Tau-crystalins are the major components of vertebrate lens. These proteins are mainly found in monomeric form with a low enzymatic activity, while the active ENO1 enzyme exists as a dimer. Irrespective of ENO1 enzyme activity, its significant presence in eye lens (23% of the total protein of the lens) clearly indicate ENO1 structural role in lens and cataracts.
In hypoxic conditions elevated ENO1 levels may provide protection to the cells by increasing anaerobic metabolism.
The smaller product of ENO1 gene, MBP-1, is known as c-myc binding protein and negative regulator of its expression. C-myc is a DNA-binding phosphoprotein and a key regulator of cell behaviour. Many of c-myc targeting pathways are deregulated in cancer cells and contribute to its enhanced expression. There are four c-myc promoters, designated as P0, P1, P2 and P3, although in normal and cancer cells most mRNAs initiate at the P2 promoter. MBP-1 binds in a region +123 to +153 relative to the c-myc P2 promoter and probably by preventing the formation of a transcription initiation complex, decrease c-myc promoter activity. Hence MBP-1 is considered as tumor suppressor.
ENO1 protein was also found as a structural component of the eye lenses and was designated as tau-crystalin. ENO1 enzyme and tau-crystalin are the products of the same gene. Tau-crystalins are the major components of vertebrate lens. These proteins are mainly found in monomeric form with a low enzymatic activity, while the active ENO1 enzyme exists as a dimer. Irrespective of ENO1 enzyme activity, its significant presence in eye lens (23% of the total protein of the lens) clearly indicate ENO1 structural role in lens and cataracts.
In hypoxic conditions elevated ENO1 levels may provide protection to the cells by increasing anaerobic metabolism.

Reaction catalyzed by Enolase.
Homology
Currently, amino acid sequences of more than 50 enolase enzymes are known. The five residues that participate in catalytic activity of this enzyme are highly conserved throughout evolution. Studies in vitro revealed that mutant enolase enzymes that differs at either positions Glu168, Glu211, Lys345, Lys396 or His159, demonstrated dramatically decreased activity level. An integral and conserved part of enolases are two Mg2+ ions that participate in conformational changes of the active site of enolase and enable binding of a substrate or its analogues.
Mutations
Note
The ENO1 gene maps to a region of chromosome 1 (1p35-p36) reported to be often deleted in several human malignancies, including neuroblastoma, melanoma, pheochromocytoma, breast, liver and colon cancer. However screening of neuroblastomas at different stages, failed to detect any mutations in ENO1 gene.
Amplification of ENO1 gene, as well as PAX7 (region 1pter-p33) was found to be a common phenomenon in squamous cell lung carcinoma.
Amplification of ENO1 gene, as well as PAX7 (region 1pter-p33) was found to be a common phenomenon in squamous cell lung carcinoma.
Implicated in
Entity name
Non-Small Cell Lung Cancer (NSCLC)
Note
Higher expression of ENO1 was demonstrated in NSCLC tissues as compared with normal lung tissues. Detection and expression level of ENO1 in primary tumors were the key factors contributing to overall patients survival rates. Relatively higher ENO1 levels in tumors correlated with poorer survival outcomes and tumor recurrence.
Other report suggests that ENO1 down-regulation in patients with NSCLC, predicts more aggressive biological behaviour. The patients whose tumors showed decreased ENO1 production had significantly poorer overall survival when compared with those without ENO1 reduction.
Also in proteomic studies, ENO1 was one of the secreted proteins demonstrated to be overexpressed by NSCLC cell line A549 as compared to controls.
Studies in vitro performed on NSCLC cell line H1299, revealed that MBP-1 overexpression correlated with decreased cell proliferation as compared with corresponding controls. Investigations in vivo demonstrated tumor suppressive properties of MBP-1. In mice with induced tumors (injection of H1299) administration of adenovirus MBP-1 construct significantly reduced tumor growth and prolonged animal survival rates.
Other report suggests that ENO1 down-regulation in patients with NSCLC, predicts more aggressive biological behaviour. The patients whose tumors showed decreased ENO1 production had significantly poorer overall survival when compared with those without ENO1 reduction.
Also in proteomic studies, ENO1 was one of the secreted proteins demonstrated to be overexpressed by NSCLC cell line A549 as compared to controls.
Studies in vitro performed on NSCLC cell line H1299, revealed that MBP-1 overexpression correlated with decreased cell proliferation as compared with corresponding controls. Investigations in vivo demonstrated tumor suppressive properties of MBP-1. In mice with induced tumors (injection of H1299) administration of adenovirus MBP-1 construct significantly reduced tumor growth and prolonged animal survival rates.
Entity name
Small Cell Lung Cancer
Note
There is some evidence concerning the role of anti-alpha-enolase antibodies in cancer associated retinopathy with SCLC. In serum obtained from patient with a sudden loss of vision, the only detectable antibodies were those against a 35-kDa anti-retinal protein. Surgical treatment performed after 1 week and 1 month, led to changes in the antibody response from antibodies against p35kDa to alpha-enolase after tumor resection. SCLC may express high levels of alpha-enolase and anti-alpha-enolase antibodies are typically detected after diagnosis of cancer.
Entity name
Thyroid carcinoma
Note
In thyroid oncocytomas, which represent a subgroup of follicular thyroid carcinoma (FTC), the up-regulation of ENO1, GPI (glucose phosphate isomerase) and GAPDH (glyceraldehydes-3-phosphate dehydrogenase) was identified as metabolic signature of thyroid carcinoma.
Important role of ENO1 in progression of thyroid carcinoma was also demonstrated for cell lines established from FTC. Pre-treatment of these cell lines with retinoic acid (RA) used in therapy and chemoprevention of solid cancers, led to decrease in ENO1 and MBP-1 expression, accompanied by reduced invasiveness of the thyroid carcinoma cells. Similar effects were also observed after silencing of common the MYC promoter-binding domain found in ENO1 and MBP-1. Both, RA-mediated and siRNA induced reduction of ENO1/MBP-1 resulted in down-regulation of c-Myc oncoprotein. It seems that in FTC the bi-functional role of ENO1 gene products is diminished and ENO1 posses the enzymatic activity only. It is worth to notice that ENO1 promoter contains two MYC binding sites (CACGTG). C-Myc over-expression and interaction with these sites may result in increased ENO1 expression and/or energy production.
In well differentiated medullary thyroid carcinomas MTC the relatively high amount of alpha beta and gamma gamma enolase isoenzymes was observed, indicating presumed neuroectodermal origin of these tumors. In highly undifferentiated and anaplastic MTCs, the majority of enzyme was represented as alpha alpha-enolase while alpha gamma-enolase was only weakly detectable.
Important role of ENO1 in progression of thyroid carcinoma was also demonstrated for cell lines established from FTC. Pre-treatment of these cell lines with retinoic acid (RA) used in therapy and chemoprevention of solid cancers, led to decrease in ENO1 and MBP-1 expression, accompanied by reduced invasiveness of the thyroid carcinoma cells. Similar effects were also observed after silencing of common the MYC promoter-binding domain found in ENO1 and MBP-1. Both, RA-mediated and siRNA induced reduction of ENO1/MBP-1 resulted in down-regulation of c-Myc oncoprotein. It seems that in FTC the bi-functional role of ENO1 gene products is diminished and ENO1 posses the enzymatic activity only. It is worth to notice that ENO1 promoter contains two MYC binding sites (CACGTG). C-Myc over-expression and interaction with these sites may result in increased ENO1 expression and/or energy production.
In well differentiated medullary thyroid carcinomas MTC the relatively high amount of alpha beta and gamma gamma enolase isoenzymes was observed, indicating presumed neuroectodermal origin of these tumors. In highly undifferentiated and anaplastic MTCs, the majority of enzyme was represented as alpha alpha-enolase while alpha gamma-enolase was only weakly detectable.
Entity name
Hepatocellular carcinoma (HCC)
Note
In proteomic studies performed on HCC cell lines and tissues, ENO1 was identified as a protein that showed stronger expression in tumor tissues when comparing to nontumorous samples. Additionally, expression of ENO1 increased with tumor dedifferentiation status. Significantly higher ENO1 expression was found in poorly differentiated HCC than in well differentiated HCC. Moreover, expression of ENO1 positively correlated with tumor size and venous invasion. Also reduction of ENO1 by specific siRNAs decreased the proliferation rates of HCC cell lines and prolonged the G2/M phase of the cell cycle.
Investigations of MBP-1 revealed its significant reduction in cirrhosis and even more diminished expression in HCC. This reduction was surprisingly accompanied by decrease in c-myc expression.
Investigations of MBP-1 revealed its significant reduction in cirrhosis and even more diminished expression in HCC. This reduction was surprisingly accompanied by decrease in c-myc expression.
Entity name
Breast carcinoma
Note
Increased expression of ENO1 was found in HER-2/neu positive breast tumors and cell lines when compared with corresponding controls. HER-2/neu is the receptor tyrosine kinase found to be overexpressed in up to 30% of breast cancers and is associated with increased metastasis rate and poor prognosis.
Introduction of MBP-1 gene into human breast carcinoma cells MDA-MB-231 and MCF-7 reduced their ability to penetrate basement membrane matrix and suppressed tumor formation in athymic nude mice. It is worth to notice that MCF-7 cell line is estrogen receptor positive and estrogen dependent for tumorigenicity.
It was demonstrated that translation of ENO1 mRNA in MCF-7 cell line is glucose concentration-dependent. Low glucose concentrations increased the level of MBP-1 protein accompanied by reduced proliferation rates. The levels of ENO1 mRNA remained unaffected. This suggests that effects induced by low glucose concentrations are mediated by preferential translation of MBP-1 (using the down-stream ATG codon). In contrast, physiologic or high glucose concentrations correlated with reduced levels of MBP-1 protein and markedly induced growth of the cells. Interestingly the low glucose group exhibited a dramatic increase in c-Myc expression, not observed in physiologic or high glucose conditions. As demonstrated for follicular thyroid carcinoma cells, also in this case c-Myc might directly transactivate ENO1 promoter, resulting in an increase in glucose uptake and elevated proliferation rates.
Introduction of MBP-1 gene into human breast carcinoma cells MDA-MB-231 and MCF-7 reduced their ability to penetrate basement membrane matrix and suppressed tumor formation in athymic nude mice. It is worth to notice that MCF-7 cell line is estrogen receptor positive and estrogen dependent for tumorigenicity.
It was demonstrated that translation of ENO1 mRNA in MCF-7 cell line is glucose concentration-dependent. Low glucose concentrations increased the level of MBP-1 protein accompanied by reduced proliferation rates. The levels of ENO1 mRNA remained unaffected. This suggests that effects induced by low glucose concentrations are mediated by preferential translation of MBP-1 (using the down-stream ATG codon). In contrast, physiologic or high glucose concentrations correlated with reduced levels of MBP-1 protein and markedly induced growth of the cells. Interestingly the low glucose group exhibited a dramatic increase in c-Myc expression, not observed in physiologic or high glucose conditions. As demonstrated for follicular thyroid carcinoma cells, also in this case c-Myc might directly transactivate ENO1 promoter, resulting in an increase in glucose uptake and elevated proliferation rates.
Entity name
Prostate cancer
Note
Investigations performed on human prostate cancer cells PC3, revealed that tumor suppressive function of MBP-1 is diminished. Reduction of endogenous MBP-1 by employing specific siRNAs resulted in decreased proliferation rates accompanied by inhibition of cyclin A1 and cyclin B1 expression. Additionally, the cell size increased after depletion of MBP-1. Introduction of exogenous MBP-1 restored cyclins expression, leading to dose-dependent increase in cyclin A1 and B1 levels.
Entity name
Brain tumors
Note
Generally, the increased levels and activity of alpha alpha-enolase correlate with brain tumorigenicity. In astrocytomas with different degrees of malignancy, oligodendrogliomas, meningiomas and ependymomas, alpha alpha-enolase was more abundant than in normal brain tissues. Among astrocytic tumors, glioblastomas revealed the highest proportion of alpha alpha-enolase as compared with control tissues.
Introduction of full length, exogenous ENO1 sequence into 1p-deleted or other neuroblastoma cell lines, led to reduction of cell growth. This suggests that in this cell lines ENO1 is preferentially translated as MBP-1 and probably does not posses the enolase enzyme activity.
Introduction of full length, exogenous ENO1 sequence into 1p-deleted or other neuroblastoma cell lines, led to reduction of cell growth. This suggests that in this cell lines ENO1 is preferentially translated as MBP-1 and probably does not posses the enolase enzyme activity.
Entity name
Multiple myeloma (plasma cell myeloma, Kahlers disease)
Note
Entity name
Acute lung inflammation (pneumonia)
Note
Increased ENO1 cell-surface expression on peripheral blood monocytes (PBMs) and strong ENO1 production in mononuclear cells in the alveolar space were demonstrated for pneumonia patients when compared with healthy volunteers. Elevated cell-surface expression of ENO1 on PBMs and on human leukemic monocyte lymphoma cell line U937, led to increased plasmin generation, enhanced monocyte migration through epithelial monolayers and promoted matrix degradation.
Entity name
Vasculitis
Note
In sera from patients with clinically proven vaculitis, anti-neutrophil cytoplasmic antibodies (ANCA) reacted with proteins present in the granules of human neutrophils. 37.3 % of these sera contained the antibodies raised against 48kDa protein, identified further as cytoplasmic alpha-enolase. Antibodies directed against enolase protein, recognised only alpha isoform and were detected in sera giving ANCA staining pattern.
Disease
Vasculitis (inflammatory destruction of blood vessels).
Entity name
Nephritis
Note
In two independent studies antibodies raised against alpha-enolase were detected in 10/41 and 9/33 sera of patients with clinically proved SLE, respectively. 80% of patients from the first report and 66.7% from the second one, suffered from active nephritis.
Disease
Nephritis (renal disease) caused by systemic lupus erythematosus (SLE, chronic autoimmune connective tissue disease that can affect any part of the body).
Entity name
Ulcerative colitis
Note
Alpha-enolase antibodies were found in about 10% of ulcerative colitis patients.
Entity name
Crohns disease
Note
Alpha-enolase antibodies were found in about 18% of patients with Crohns disease.
Disease
Crohns disease (autoimmune, inflammatory disease of the intestines that may affect any part of the gastrointestinal tract).
Entity name
Primary biliary cirrhosis and autoimmune hepatitis
Note
Alpha-enolase antibodies were present in 28.6% of patients with primary biliary cirrhosis and in 31.6% with autoimmune hepatitis. Normal subjects revealed significantly lower levels of alpha-enolase antibodies when compared with both diseases. Note that antibodies against beta and gamma enolases were not found in any serum sample analysed.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 7499243 | 1995 | Non-neuronal enolase is an endothelial hypoxic stress protein. | Aaronson RM et al |
| 9126798 | 1997 | Identification of an autoantibody against alpha-enolase in primary biliary cirrhosis. | Akisawa N et al |
| 14764826 | 2004 | Transcriptional profiling reveals coordinated up-regulation of oxidative metabolism genes in thyroid oncocytic tumors. | Baris O et al |
| 17020980 | 2006 | Identification of alpha-enolase as an autoantigen in lung cancer: its overexpression is associated with clinical outcomes. | Chang GC et al |
| 14506152 | 2003 | Enolase-alpha is frequently down-regulated in non-small cell lung cancer and predicts aggressive biological behavior. | Chang YS et al |
| 15808190 | 2005 | Anti-alpha-enolase antibodies in cancer-associated retinopathy with small cell carcinoma of the lung. | Dot C et al |
| 16359544 | 2005 | Introduction of in vitro transcribed ENO1 mRNA into neuroblastoma cells induces cell death. | Ejeskär K et al |
| 11318533 | 2001 | Expression of c-myc promoter binding protein (MBP-1), a novel eukaryotic repressor gene, in cirrhosis and human hepatocellular carcinoma. | Fan X et al |
| 10802057 | 2000 | ENO1 gene product binds to the c-myc promoter and acts as a transcriptional repressor: relationship with Myc promoter-binding protein 1 (MBP-1). | Feo S et al |
| 17178888 | 2006 | Tumor-suppressive effects of MBP-1 in non-small cell lung cancer cells. | Ghosh AK et al |
| 2373081 | 1990 | Structure of the human gene for alpha-enolase. | Giallongo A et al |
| 18813785 | 2008 | Glycolysis module activated by hypoxia-inducible factor 1alpha is related to the aggressive phenotype of hepatocellular carcinoma. | Hamaguchi T et al |
| 17094902 | 2006 | Proteomic analysis of secreted proteins of non-small cell lung cancer. | Huang LJ et al |
| 8632173 | 1996 | Enolase activity and isoenzyme distribution in human brain regions and tumors. | Joseph J et al |
| 9169614 | 1997 | Biochemical characterization of the mouse muscle-specific enolase: developmental changes in electrophoretic variants and selective binding to other proteins. | Merkulova T et al |
| 8455367 | 1993 | Alpha-enolase: a novel cytosolic autoantigen in ANCA positive vasculitis. | Moodie FD et al |
| 3965094 | 1985 | Enolase isozymes in differentiated and undifferentiated medullary thyroid carcinomas. | Oskam R et al |
| 11497239 | 2001 | Multifunctional alpha-enolase: its role in diseases. | Pancholi V et al |
| 10648026 | 2000 | Autoantibodies specific for alpha-enolase in systemic autoimmune disorders. | Pratesi F et al |
| 10853020 | 2000 | Gene amplification at chromosome 1pter-p33 including the genes PAX7 and ENO1 in squamous cell lung carcinoma. | Rácz A et al |
| 7641187 | 1995 | Human breast carcinoma cells transfected with the gene encoding a c-myc promoter-binding protein (MBP-1) inhibits tumors in nude mice. | Ray RB et al |
| 17595061 | 2007 | c-myc promoter binding protein regulates the cellular response to an altered glucose concentration. | Sedoris KC et al |
| 10681589 | 2000 | Structural analysis of alpha-enolase. Mapping the functional domains involved in down-regulation of the c-myc protooncogene. | Subramanian A et al |
| 15800975 | 2005 | Overexpression of alpha enolase in hepatitis C virus-related hepatocellular carcinoma: association with tumor progression as determined by proteomic analysis. | Takashima M et al |
| 19060179 | 2009 | Retinoic acid-mediated down-regulation of ENO1/MBP-1 gene products caused decreased invasiveness of the follicular thyroid carcinoma cell lines. | Trojanowicz B et al |
| 8978760 | 1996 | Report of the second international workshop on human chromosome 1 mapping 1995. | Weith A et al |
| 10375612 | 1999 | Identification of c-myc promoter-binding protein and X-box binding protein 1 as interleukin-6 target genes in human multiple myeloma cells. | Wen XY et al |
| 2462567 | 1988 | Tau-crystallin/alpha-enolase: one gene encodes both an enzyme and a lens structural protein. | Wistow GJ et al |
| 19182206 | 2009 | Enolase-1 promotes plasminogen-mediated recruitment of monocytes to the acutely inflamed lung. | Wygrecka M et al |
| 16820872 | 2006 | Gene expression profiling of human HBV- and/or HCV-associated hepatocellular carcinoma cells using expressed sequence tags. | Yoon SY et al |
| 16048908 | 2005 | Proteomic study reveals that proteins involved in metabolic and detoxification pathways are highly expressed in HER-2/neu-positive breast cancer. | Zhang D et al |
Other Information
Locus ID:
NCBI: 2023
MIM: 172430
HGNC: 3350
Ensembl: ENSG00000074800
Variants:
dbSNP: 2023
ClinVar: 2023
TCGA: ENSG00000074800
COSMIC: ENO1
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38273010 | 2024 | Multi-omics reveals the role of ENO1 in bladder cancer and constructs an epithelial-related prognostic model to predict prognosis and efficacy. | 0 |
| 38517039 | 2024 | ENO1 contributes to the gemcitabine resistance of pancreatic cancer through the YAP1 signaling pathway. | 0 |
| 38661062 | 2024 | ENO1 promotes trophoblast invasion regulated by E2F8 in recurrent miscarriage. | 1 |
| 38823761 | 2024 | N6-Methyladenosine enhances the translation of ENO1 to promote the progression of bladder cancer by inhibiting PCNA ubiquitination. | 0 |
| 38273010 | 2024 | Multi-omics reveals the role of ENO1 in bladder cancer and constructs an epithelial-related prognostic model to predict prognosis and efficacy. | 0 |
| 38517039 | 2024 | ENO1 contributes to the gemcitabine resistance of pancreatic cancer through the YAP1 signaling pathway. | 0 |
| 38661062 | 2024 | ENO1 promotes trophoblast invasion regulated by E2F8 in recurrent miscarriage. | 1 |
| 38823761 | 2024 | N6-Methyladenosine enhances the translation of ENO1 to promote the progression of bladder cancer by inhibiting PCNA ubiquitination. | 0 |
| 36494582 | 2023 | Glioblastoma glycolytic signature predicts unfavorable prognosis, immunological heterogeneity, and ENO1 promotes microglia M2 polarization and cancer cell malignancy. | 5 |
| 36614179 | 2023 | ENO1 Promotes OSCC Migration and Invasion by Orchestrating IL-6 Secretion from Macrophages via a Positive Feedback Loop. | 3 |
| 36674531 | 2023 | Determination of the Predictive Roles and Potentially Pathogenic Antigen Epitopes of α-Enolase Related to the Development of Miscarriage in Females with Autoimmune Thyroiditis. | 0 |
| 37011206 | 2023 | The moonlighting function of glycolytic enzyme enolase-1 promotes choline phospholipid metabolism and tumor cell proliferation. | 6 |
| 37083649 | 2023 | miR-22-3p Suppresses Cell Proliferation and Migration of Gastric Cancer by Targeting ENO1. | 0 |
| 37247894 | 2023 | Overexpression of c-MYC Promoter Binding Protein-1 Enhances Proliferation and Glucose Metabolism of Melanoma Cells Lines. | 0 |
| 37288669 | 2023 | Alpha-enolase 1 knockdown facilitates the proliferation and invasion of villous trophoblasts by upregulating COX-2. | 1 |
Citation
Bogusz Trojanowicz ; Cuong Hoang-Vu ; Carsten Sekulla
ENO1 (Enolase 1, (alpha))
Atlas Genet Cytogenet Oncol Haematol. 2009-08-01
Online version: http://atlasgeneticsoncology.org/gene/40453/eno1
