HBP1 (HMG-box transcription factor 1)
2009-03-01 K Eric Paulson , Amy S Yee AffiliationDepartments of Biochemistry, Radiation Oncology, Tufts Medical School, Boston, MA 02111, USA (KEP); Department of Biochemistry, Tufts Medical School, Boston, MA 02111, USA (ASY)
DNA/RNA
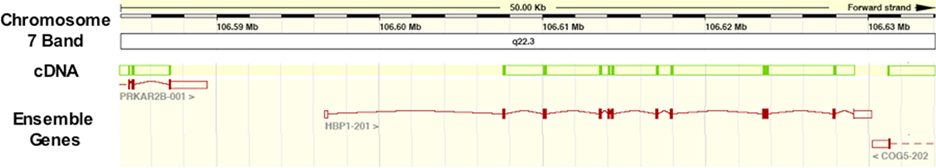
HBP1 Locus on 7q22. Chromosome 7, band q22 is shown at the top of the diagram with the distance on the forward strand marked as indicated. The HBP1 gene is located between the PRKAR2B gene (left) and the COG5 gene (right). Coding exons are shown in bold red, while non-coding are shown with open red boxes. cDNA coding sequence is shown in dark green.
Description
Sequence length 33515; cDNA length 2829; Coding sequence 1545. The gene is comprised of 11 exons; max. exon length 1121, min. exon length 54. Number of SNPs 6.
Transcription
The consensus normal transcript is 2829 nt and the coding sequence is 1545 nt. The consensus normal transcript is produced from 11 exons. The first exon is non-coding. There are 16 different mRNAs produced, including 13 different alternatively spliced variants. Several of these alternatively spliced variants appear to be produced only in tumor cells.
Pseudogene
None known.
Proteins
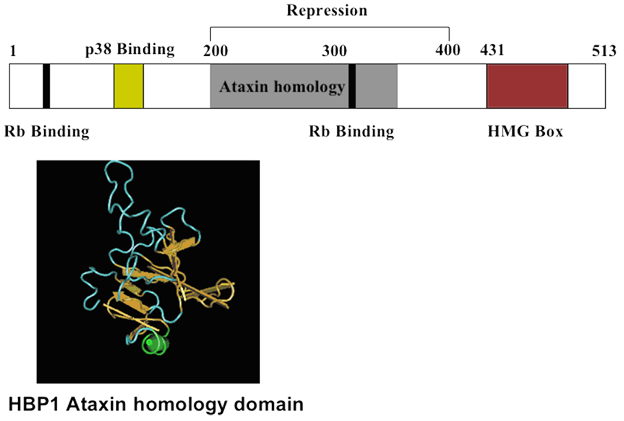
HBP1 Protein Schematic. HBP1 contains two defined domains; an ataxin homology domain (gray) and an HMG box DNA binding domain (red). There are two Rb-interacting motifs (black line) and a p38 MAP kinase-interacting region (yellow). A repression domain (aa200-400) encompasses the Ataxin homology domain.
Description
HBP1 encodes a 515 amino acid, 62-kDa, transcriptional repressor. On western blots, HBP1 runs anomolously at 75 kD. HBP1 represses numerous target genes when overexpressed, including N-Myc, cyclin D1 and c-myc. The protein contains two known domains: an HMG box DNA binding domain and a repression domain that contains an ataxin homology domain.
Expression
HBP1 mRNA is expressed in several human organs and tissues: brain; uterus; testis; mixed; uncharacterized tissue; lung; bladder; kidney; heart; lymph node; blood; prostate; trachea; esophagus; cervix; skin; adrenal gland; eye; intestine; vascular; amniotic fluid; mouth; embryonic tissue; spleen; thymus; placenta; connective tissue; liver; mammary gland; muscle; pancreas; stomach; ovary; ascites; ganglia; bone; pharynx; thyroid; adipose tissue; parathyroid; ear; pineal gland; nerve; bone marrow; umbilical cord.
Localisation
HBP1 is predominantly a nuclear protein.
Function
Cell Cycle. HBP1 was isolated as a cell cycle inhibitor and HMG-box transcriptional repressor. HBP1 binds to Rb and p107 via an LXCXE and IXCXE binding motif as part of HBP1 function in cell cycle regulation. HBP1 expression was uniquely associated with oncogene-mediated senescence in an RB-dependent manner.
Growth and Differentiation. Liver-specific expression of HBP1 was shown to inhibit liver regeneration. In addition, ectopic expression of HBP1 regulates differentiation in muscle cells. Similarly, transgenic mice overexpressing HBP1 exhibited altered thymus cellularity and decreased thymocyte development. Finally, HBP1 expression is maintained in the developing testis beyond the onset of spermatogenesis, and the expression of HBP1 in XY germ cells appears to correlate with the onset of mitotic arrest. The repression domain of HBP1 contains an Ataxin homology domain which interacts with the Sin3 corepressor PAH2 domain, thus recruiting HDAC1 to a repression complex.
Signaling and Transcription. HBP1 inhibits a number of genes through direct binding to its cognate recognition sequence, including N-myc and p47 phox. In addition, HBP1 is an inhibitor of Wnt signaling and blocks beta-catenin/LEF/TCF complexes. Wnt target genes regulated by HBP1 include c-myc and cyclin D1. HBP1 is regulated by p38 MAP kinase signaling and contains both a p38 MAP kinase docking sequence and a p38 MAP kinase phosphorylation site. Phosphorylation results in HBP1 protein stabilization and increased repression function. p38 MAP kinase phosphorylation is also required for HBP1 induction of premature senescence. HBP1 mRNA is stabilized by the green tea compound epigallo-catechin galate (EGCG), resulting in increased HBP1 protein and HBP1-dependent repression of Wnt signaling.
Growth and Differentiation. Liver-specific expression of HBP1 was shown to inhibit liver regeneration. In addition, ectopic expression of HBP1 regulates differentiation in muscle cells. Similarly, transgenic mice overexpressing HBP1 exhibited altered thymus cellularity and decreased thymocyte development. Finally, HBP1 expression is maintained in the developing testis beyond the onset of spermatogenesis, and the expression of HBP1 in XY germ cells appears to correlate with the onset of mitotic arrest. The repression domain of HBP1 contains an Ataxin homology domain which interacts with the Sin3 corepressor PAH2 domain, thus recruiting HDAC1 to a repression complex.
Signaling and Transcription. HBP1 inhibits a number of genes through direct binding to its cognate recognition sequence, including N-myc and p47 phox. In addition, HBP1 is an inhibitor of Wnt signaling and blocks beta-catenin/LEF/TCF complexes. Wnt target genes regulated by HBP1 include c-myc and cyclin D1. HBP1 is regulated by p38 MAP kinase signaling and contains both a p38 MAP kinase docking sequence and a p38 MAP kinase phosphorylation site. Phosphorylation results in HBP1 protein stabilization and increased repression function. p38 MAP kinase phosphorylation is also required for HBP1 induction of premature senescence. HBP1 mRNA is stabilized by the green tea compound epigallo-catechin galate (EGCG), resulting in increased HBP1 protein and HBP1-dependent repression of Wnt signaling.
Homology
HBP1 contains two recognized homology motifs; an Ataxin homology domain and an HMG box DNA binding domain.
Mutations
Somatic
In an analysis of 76 breast tumors, 10 HBP1 mutations/variants were identified that were associated with fully invasive breast cancer. Some of these mutants/variants were shown to be the result of genomic mutations.
Implicated in
Entity name
Breast Cancer
Disease
Aberrations in HBP1 are associated with invasive breast cancer. The HBP1 gene is either mutated or reduced in breast cancer. As cited above in the "Somatic Mutation section", HBP1 mutations/variants were associated with fully invasive breast cancer, some of which arose from genomic mutations. In a new analysis, a subset of invasive breast cancer tumors had markedly reduced expression of the HBP1 mRNA.
Prognosis
Statistical analysis of a breast cancer patient database predicted that reduced HBP1 mRNA levels were associated with a decreased relapse-free survival and recurrence with distant metastases (Paulson et al., 2007).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15024088 | 2004 | HBP1 repression of the p47phox gene: cell cycle regulation via the NADPH oxidase. | Berasi SP et al |
| 10030670 | 1998 | Activation and repression of p21(WAF1/CIP1) transcription by RB binding proteins. | Gartel AL et al |
| 16495219 | 2006 | Suppression of Wnt signaling by the green tea compound (-)-epigallocatechin 3-gallate (EGCG) in invasive breast cancer cells. Requirement of the transcriptional repressor HBP1. | Kim J et al |
| 9178770 | 1997 | The HMG-box transcription factor HBP1 is targeted by the pocket proteins and E1A. | Lavender P et al |
| 10958660 | 2000 | Involvement of retinoblastoma protein and HBP1 in histone H1(0) gene expression. | Lemercier C et al |
| 17616670 | 2007 | Alterations of the HBP1 transcriptional repressor are associated with invasive breast cancer. | Paulson KE et al |
| 11500377 | 2001 | Negative regulation of the Wnt-beta-catenin pathway by the transcriptional repressor HBP1. | Sampson EM et al |
| 16210625 | 2005 | Human high mobility group box transcription factor 1 affects thymocyte development and transgene variegation. | Sekkali B et al |
| 9671483 | 1998 | Regulation of differentiation by HBP1, a target of the retinoblastoma protein. | Shih HH et al |
| 11486012 | 2001 | HMG box transcriptional repressor HBP1 maintains a proliferation barrier in differentiated liver tissue. | Shih HH et al |
| 18258815 | 2008 | Evidence for the involvement of miRNA in redox regulated angiogenic response of human microvascular endothelial cells. | Shilo S et al |
| 15162515 | 2004 | HMG box transcription factor gene Hbp1 is expressed in germ cells of the developing mouse testis. | Smith JM et al |
| 15235594 | 2004 | HBP1 and Mad1 repressors bind the Sin3 corepressor PAH2 domain with opposite helical orientations. | Swanson KA et al |
| 9030690 | 1997 | HBP1: a HMG box transcriptional repressor that is targeted by the retinoblastoma family. | Tevosian SG et al |
| 14612426 | 2003 | The transcriptional repressor HBP1 is a target of the p38 mitogen-activated protein kinase pathway in cell cycle regulation. | Xiu M et al |
| 15225871 | 2004 | The HBP1 transcriptional repressor and the p38 MAP kinase: unlikely partners in G1 regulation and tumor suppression. | Yee AS et al |
| 16966377 | 2006 | The HBP1 transcriptional repressor participates in RAS-induced premature senescence. | Zhang X et al |
| 10562551 | 1999 | Human HMG box transcription factor HBP1: a role in hCD2 LCR function. | Zhuma T et al |
| 15893665 | 2005 | The AXH domain adopts alternative folds the solution structure of HBP1 AXH. | de Chiara C et al |
Other Information
Locus ID:
NCBI: 26959
MIM: 616714
HGNC: 23200
Ensembl: ENSG00000105856
Variants:
dbSNP: 26959
ClinVar: 26959
TCGA: ENSG00000105856
COSMIC: HBP1
RNA/Proteins
Expression (GTEx)
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37414199 | 2023 | Targeting of HBP1/TIMP3 axis as a novel strategy against breast cancer. | 2 |
| 37414199 | 2023 | Targeting of HBP1/TIMP3 axis as a novel strategy against breast cancer. | 2 |
| 34013373 | 2021 | UHRF1 regulates the transcriptional repressor HBP1 through MIF in T acute lymphoblastic leukemia. | 7 |
| 34013373 | 2021 | UHRF1 regulates the transcriptional repressor HBP1 through MIF in T acute lymphoblastic leukemia. | 7 |
| 32549582 | 2020 | Suppression of p38/HBP1 pathway alleviates hyperosmotic stress-induced senescent progression of chondrocyte senescence. | 0 |
| 32677158 | 2020 | Regulatory role of transcription factor HBP1 in anticancer efficacy of EGFR inhibitor erlotinib in HNSCC. | 2 |
| 32964956 | 2020 | HBP1 deficiency protects against stress-induced premature senescence of nucleus pulposus. | 2 |
| 33327874 | 2020 | miR-146b Functions as an Oncogene in Oral Squamous Cell Carcinoma by Targeting HBP1. | 5 |
| 32549582 | 2020 | Suppression of p38/HBP1 pathway alleviates hyperosmotic stress-induced senescent progression of chondrocyte senescence. | 0 |
| 32677158 | 2020 | Regulatory role of transcription factor HBP1 in anticancer efficacy of EGFR inhibitor erlotinib in HNSCC. | 2 |
| 32964956 | 2020 | HBP1 deficiency protects against stress-induced premature senescence of nucleus pulposus. | 2 |
| 33327874 | 2020 | miR-146b Functions as an Oncogene in Oral Squamous Cell Carcinoma by Targeting HBP1. | 5 |
| 30191992 | 2019 | Matrix metalloproteinase-13 is a target gene of high-mobility group box-containing protein 1 in modulating oral cancer cell invasion. | 4 |
| 30538293 | 2019 | ALK positively regulates MYCN activity through repression of HBP1 expression. | 16 |
| 30683982 | 2019 | The HMG box transcription factor HBP1: a cell cycle inhibitor at the crossroads of cancer signaling pathways. | 21 |
Citation
K Eric Paulson ; Amy S Yee
HBP1 (HMG-box transcription factor 1)
Atlas Genet Cytogenet Oncol Haematol. 2009-03-01
Online version: http://atlasgeneticsoncology.org/gene/40791/hbp1
