HOXC13 (homeobox C13)
2009-06-01 Toshiyuki Yamada , Shigeki Tsuchida AffiliationDepartment of Biochemistry, Genome Biology, Hirosaki University Graduate School of Medicine, 5 Zaifu-Cho, Hirosaki 036-8562, Japan
Identity
HGNC
LOCATION
12q13.13
LOCUSID
ALIAS
ECTD9,HOX3,HOX3G
FUSION GENES
DNA/RNA
Description
Spans around 7.7 kb genomic region, contains 2 exons, homeodomain is located in exon 2.
Transcription
Transcript; 2.2 kb, open reading frame; 990 bp.
Proteins
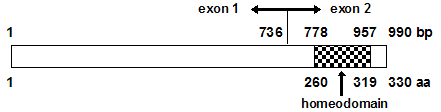
Description
330 amino acids, DNA binding protein with a 60 aa of homeodomain (helix-turn-helix domain) in the C-terminal.
Expression
In nail, hair follicle, tongue (filiform papilla), AML cells, erythroleukemia cells.
Localisation
Nucleus.
Function
Important for patterning of embryo, role in hair follicular formation and hair cycle control, role in leukemogenesis of myeloid cells and differentiation of erythroleukemia cells.
Homology
With homeodomain proteins, ABD-B homeodomain proteins.
Mutations
Germinal
Not known.
Somatic
Not known.
Implicated in
Entity name
t(11;12)(p15;q13)
Disease
AML-M2
Cytogenetics
Chromosome translocation formimg a fusion transcript of 5 NUP98 gene and 3 HOXC13 gene.
Hybrid gene
5 NUP98-3 HOXC13.
Fusion protein
Fusion between FG motifs of NUP98 and the homeodomain of HOXC13.

Entity name
Cytogenetics
Not examined.
Hybrid gene
Not examined.
Oncogenesis
Expressed in human and mouse erythroleukemia cells, overexpression of HOXC13 inhibits differentiation of murine erythroleukemia (MEL) cells.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 9420327 | 1998 | Hoxc13 mutant mice lack external hair. | Godwin AR et al |
| 12619167 | 2003 | Human homeobox gene HOXC13 is the partner of NUP98 in adult acute myeloid leukemia with t(11;12)(p15;q13). | La Starza R et al |
| 1712489 | 1991 | Coordinate regulation of HOX genes in human hematopoietic cells. | Magli MC et al |
| 12461755 | 2003 | Fusion of the NUP98 gene and the homeobox gene HOXC13 in acute myeloid leukemia with t(11;12)(p15;q13). | Panagopoulos I et al |
| 11290294 | 2001 | Overexpression of Hoxc13 in differentiating keratinocytes results in downregulation of a novel hair keratin gene cluster and alopecia. | Tkatchenko AV et al |
| 18076876 | 2008 | Interaction between the homeodomain protein HOXC13 and ETS family transcription factor PU.1 and its implication in the differentiation of murine erythroleukemia cells. | Yamada T et al |
Other Information
Locus ID:
NCBI: 3229
MIM: 142976
HGNC: 5125
Ensembl: ENSG00000123364
Variants:
dbSNP: 3229
ClinVar: 3229
TCGA: ENSG00000123364
COSMIC: HOXC13
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000123364 | ENST00000243056 | P31276 |
Expression (GTEx)
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36833285 | 2023 | Variants Identified in the HOXC13 and HOXD13 Genes Suggest Association with Cervical Cancer in a Cohort of Mexican Women. | 0 |
| 38057852 | 2023 | Silencing HOXC13 exerts anti-prostate cancer effects by inducing DNA damage and activating cGAS/STING/IRF3 pathway. | 0 |
| 36833285 | 2023 | Variants Identified in the HOXC13 and HOXD13 Genes Suggest Association with Cervical Cancer in a Cohort of Mexican Women. | 0 |
| 38057852 | 2023 | Silencing HOXC13 exerts anti-prostate cancer effects by inducing DNA damage and activating cGAS/STING/IRF3 pathway. | 0 |
| 36029503 | 2022 | LncRNA HOXC-AS5 Affects the Proliferation, Invasion and Cell Cycle of Ameloblastoma Cells by Acting on the Target Gene HOXC13. | 1 |
| 36029503 | 2022 | LncRNA HOXC-AS5 Affects the Proliferation, Invasion and Cell Cycle of Ameloblastoma Cells by Acting on the Target Gene HOXC13. | 1 |
| 33427025 | 2021 | HOXC-AS2 mediates the proliferation, apoptosis, and migration of non-small cell lung cancer by combining with HOXC13 gene. | 7 |
| 34309767 | 2021 | HOXC13 promotes cervical cancer proliferation, invasion and Warburg effect through β-catenin/c-Myc signaling pathway. | 8 |
| 34763232 | 2021 | HOXC6/8/10/13 predict poor prognosis and associate with immune infiltrations in glioblastoma. | 5 |
| 33427025 | 2021 | HOXC-AS2 mediates the proliferation, apoptosis, and migration of non-small cell lung cancer by combining with HOXC13 gene. | 7 |
| 34309767 | 2021 | HOXC13 promotes cervical cancer proliferation, invasion and Warburg effect through β-catenin/c-Myc signaling pathway. | 8 |
| 34763232 | 2021 | HOXC6/8/10/13 predict poor prognosis and associate with immune infiltrations in glioblastoma. | 5 |
| 31642514 | 2020 | The homeobox transcription factor HOXC13 upregulates human papillomavirus E1 gene expression and contributes to viral genome maintenance. | 4 |
| 32763778 | 2020 | HOXC13-AS accelerates cell proliferation and migration in oral squamous cell carcinoma via miR-378g/HOXC13 axis. | 13 |
| 31642514 | 2020 | The homeobox transcription factor HOXC13 upregulates human papillomavirus E1 gene expression and contributes to viral genome maintenance. | 4 |
Citation
Toshiyuki Yamada ; Shigeki Tsuchida
HOXC13 (homeobox C13)
Atlas Genet Cytogenet Oncol Haematol. 2009-06-01
Online version: http://atlasgeneticsoncology.org/gene/473/hoxc13
