HUS1 (HUS1 checkpoint homolog (S. pombe))
2010-09-01 Amrita Madabushi , Randall C Gunther , A-Lien Lu AffiliationDepartment of Biochemistry, Molecular Biology, School of Medicine, University of Maryland, 108 North Greene Street, Baltimore, Maryland 21201, USA
DNA/RNA
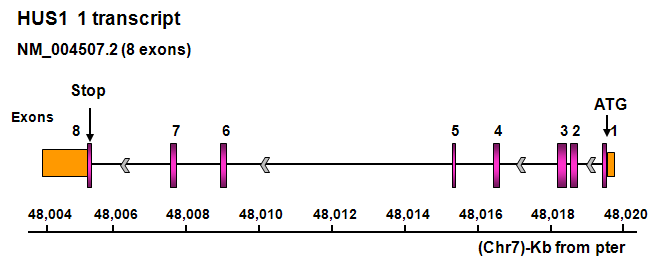
Description
Transcription
Pseudogene
Proteins
Description
Expression
Localisation
Function
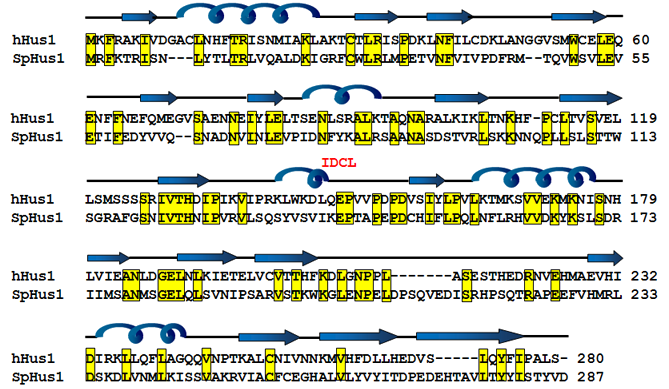
The structure of the 9-1-1 complex (Doré et al., 2009; Sohn and Cho, 2009; Xu et al., 2009) is similar to the sliding clamp proliferating cell nuclear antigen protein (PCNA) (Gulbis et al., 1996; Krishna et al., 1994). Hus1 interacts with Rad9 and Rad1 through its N terminal domain and C terminal domain, respectively. The structure and surface charge distribution of the interdomain connecting loop (IDCL) of Hus1 differs from those of other two subunits (Doré et al., 2009; Sohn and Cho, 2009; Xu et al., 2009). The IDCL of Hus1 contains an N-terminal alpha helix and positive charge cluster. These differences among Hus1, Rad9, and Rad1 may contribute to different binding affinities to their partner proteins. For example, MutY homolog (MYH) has a strong preference to bind Hus1 (Chang and Lu, 2005; Shi et al., 2006).
Recent structural and functional analyses indicate that the Hus1 binding region of MYH adopts a stabilized conformation projecting away from the catalytic domain to form a docking scaffold for Hus1 and binds to Hus1 through electrostatic interaction (Luncsford et al., 2010).
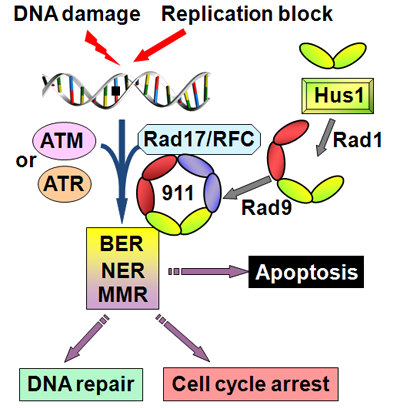
The 9-1-1 complex is required to activate two checkpoint sensors-ATM (ataxia telangiectasia [AT] mutated protein) and ATR (ATM- and Rad3-related protein), which are phosphoinositol phosphate 3 kinase-related kinases (PIKKs) (Zhou and Elledge, 2000). The DNA-bound 9-1-1 complex facilitates ATM- or ATR-mediated phosphorylation of more than 700 proteins including Chk1, Chk2, p53, and BRCA1 (Zhou and Elledge, 2000). Hus1 facilitated phosphorylation of Chk1 kinase is required for the ATR-dependent checkpoint; and regulates S-phase progression, G2/M arrest, and replication fork stabilization (Sancar et al., 2004; Sancar et al., 2004). However, Hus1 is not required for Chk2 phosphorylation in response to certain genotoxins (Weiss et al., 2003).
Besides acting as a DNA damage sensor, the 9-1-1 complex plays an integral role in several DNA repair pathways including base excision repair (BER), mismatch repair (MMR), and nucleotide excision repair (NER) (see figure 2) (Helt et al., 2005).
In the BER pathway, the 9-1-1 complex facilitates and interacts with several DNA glycosylases including MYH (Chang and Lu, 2005; Shi et al., 2006; Chang and Lu, 2005; Shi et al., 2006), 8-oxoG glycosylase (OGG1) (Park et al., 2009), NEIL1 (Guan et al., 2007a), and thymine DNA glycosylase (TDG) (Guan et al., 2007b). The 9-1-1 complex also interacts with and stimulates other BER enzymes including APE1 (Gembka et al., 2007), POLbeta (Toueille et al., 2004), FEN1 (Friedrich-Heineken et al., 2005; Wang et al., 2004a), RPA (Wu et al., 2005), and DNA ligase 1 (Smirnova et al., 2005; Wang et al., 2006a). Thus, the 9-1-1 complex may provide a platform for the assembly and function of the BER machinery (Balakrishnan et al., 2009; Lu et al., 2006). The 9-1-1 complex enhances mismatch repair via direct interaction with mismatch recognition proteins (MSH2/MSH3, MSH2/MSH6, and MLH1/PMS2) (Bai et al., 2010; He et al., 2008). Hus1 interacts with MSH2/MSH3 and MSH2/MSH6, but not with MLH1/PMS2 (Bai et al., 2010; He et al., 2008).
In the NER pathway, interactions between Saccharomyces cerevisiae Rad14 (hXPA homolog) and the checkpoint proteins ScDdc1 (hRad9 homolog) and ScMec3 (hHus1 homolog) have been demonstrated (Giannattasio et al., 2004). Inactivation of NER by knock down of XPA and XPC resulted in a decrease of G1 phase cells that displayed Rad9 foci in response to UV light (Warmerdam et al., 2009). UV light-induced Rad9 foci also colocalized with TopBP1 and gamma-H2AX (Warmerdam et al., 2009).
Hus1 interacts with histone deacetylase HDAC1 (Cai et al., 2000). A novel pathway has been proposed that HDAC1 is involved in G(2)/M checkpoint control through the interaction with the 9-1-1 complex.
Jab1 physically associates with the 9-1-1 complex, causes the translocation of the 9-1-1 complex from the nucleus to the cytoplasm, and mediates the rapid degradation of the 9-1-1 complex (Huang et al., 2007).
Homology
Mutations
Note
Implicated in
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 20188637 | 2010 | Interaction between human mismatch repair recognition proteins and checkpoint sensor Rad9-Rad1-Hus1. | Bai H et al |
| 19329425 | 2009 | Long patch base excision repair proceeds via coordinated stimulation of the multienzyme DNA repair complex. | Balakrishnan L et al |
| 12578958 | 2003 | Loading of the human 9-1-1 checkpoint complex onto DNA by the checkpoint clamp loader hRad17-replication factor C complex in vitro. | Bermudez VP et al |
| 10846170 | 2000 | HDAC1, a histone deacetylase, forms a complex with Hus1 and Rad9, two G2/M checkpoint Rad proteins. | Cai RL et al |
| 15533944 | 2005 | Interaction of checkpoint proteins Hus1/Rad1/Rad9 with DNA base excision repair enzyme MutY homolog in fission yeast, Schizosaccharomyces pombe. | Chang DY et al |
| 19446481 | 2009 | Crystal structure of the rad9-rad1-hus1 DNA damage checkpoint complex--implications for clamp loading and regulation. | Doré AS et al |
| 14624239 | 2003 | Biochemical characterization of DNA damage checkpoint complexes: clamp loader and clamp complexes with specificity for 5' recessed DNA. | Ellison V et al |
| 16890531 | 2006 | Telomere and telomerase modulation by the mammalian Rad9/Rad1/Hus1 DNA-damage-checkpoint complex. | Francia S et al |
| 16216273 | 2005 | The two DNA clamps Rad9/Rad1/Hus1 complex and proliferating cell nuclear antigen differentially regulate flap endonuclease 1 activity. | Friedrich-Heineken E et al |
| 17426133 | 2007 | The checkpoint clamp, Rad9-Rad1-Hus1 complex, preferentially stimulates the activity of apurinic/apyrimidinic endonuclease 1 and DNA polymerase beta in long patch base excision repair. | Gembka A et al |
| 14726955 | 2004 | Physical and functional interactions between nucleotide excision repair and DNA damage checkpoint. | Giannattasio M et al |
| 17395641 | 2007 | The human checkpoint sensor Rad9-Rad1-Hus1 interacts with and stimulates NEIL1 glycosylase. | Guan X et al |
| 17855402 | 2007 | The human checkpoint sensor Rad9-Rad1-Hus1 interacts with and stimulates DNA repair enzyme TDG glycosylase. | Guan X et al |
| 8861913 | 1996 | Structure of the C-terminal region of p21(WAF1/CIP1) complexed with human PCNA. | Gulbis JM et al |
| 10777662 | 2000 | Physical interactions among human checkpoint control proteins HUS1p, RAD1p, and RAD9p, and implications for the regulation of cell cycle progression. | Hang H et al |
| 18842633 | 2008 | Rad9 plays an important role in DNA mismatch repair through physical interaction with MLH1. | He W et al |
| 15876866 | 2005 | Evidence that DNA damage detection machinery participates in DNA repair. | Helt CE et al |
| 17583730 | 2007 | Jab1 mediates protein degradation of the Rad9-Rad1-Hus1 checkpoint complex. | Huang J et al |
| 11920544 | 2002 | Downregulation of Hus1 by antisense oligonucleotides enhances the sensitivity of human lung carcinoma cells to cisplatin. | Kinzel B et al |
| 8001157 | 1994 | Crystal structure of the eukaryotic DNA polymerase processivity factor PCNA. | Krishna TS et al |
| 15919177 | 2005 | Conditional inactivation of the mouse Hus1 cell cycle checkpoint gene. | Levitt PS et al |
| 17220276 | 2007 | Genome maintenance defects in cultured cells and mice following partial inactivation of the essential cell cycle checkpoint gene Hus1. | Levitt PS et al |
| 16720376 | 2006 | MutY and MutY homologs (MYH) in genome maintenance. | Lu AL et al |
| 20816984 | 2010 | A structural hinge in eukaryotic MutY homologues mediates catalytic activity and Rad9-Rad1-Hus1 checkpoint complex interactions. | Luncsford PJ et al |
| 18794804 | 2008 | Loss of Hus1 sensitizes cells to etoposide-induced apoptosis by regulating BH3-only proteins. | Meyerkord CL et al |
| 19615952 | 2009 | Repair activities of human 8-oxoguanine DNA glycosylase are stimulated by the interaction with human checkpoint sensor Rad9-Rad1-Hus1 complex. | Park MJ et al |
| 15189136 | 2004 | Molecular mechanisms of mammalian DNA repair and the DNA damage checkpoints. | Sancar A et al |
| 16879101 | 2006 | Physical and functional interactions between MutY glycosylase homologue (MYH) and checkpoint proteins Rad9-Rad1-Hus1. | Shi G et al |
| 15871698 | 2005 | The human checkpoint sensor and alternative DNA clamp Rad9-Rad1-Hus1 modulates the activity of DNA ligase I, a component of the long-patch base excision repair machinery. | Smirnova E et al |
| 19464297 | 2009 | Crystal structure of the human rad9-hus1-rad1 clamp. | Sohn SY et al |
| 10359610 | 1999 | The human G2 checkpoint control protein hRAD9 is a nuclear phosphoprotein that forms complexes with hRAD1 and hHUS1. | St Onge RP et al |
| 15314187 | 2004 | The human Rad9/Rad1/Hus1 damage sensor clamp interacts with DNA polymerase beta and increases its DNA substrate utilisation efficiency: implications for DNA repair. | Toueille M et al |
| 18950845 | 2009 | Evaluating new candidate SNPs as low penetrance risk factors in sporadic breast cancer: a two-stage Spanish case-control study. | Vega A et al |
| 9872989 | 1999 | Human homologs of Schizosaccharomyces pombe rad1, hus1, and rad9 form a DNA damage-responsive protein complex. | Volkmer E et al |
| 15556996 | 2004 | The human Rad9-Rad1-Hus1 checkpoint complex stimulates flap endonuclease 1. | Wang W et al |
| 16731526 | 2006 | Mechanism of stimulation of human DNA ligase I by the Rad9-rad1-Hus1 checkpoint complex. | Wang W et al |
| 14762204 | 2004 | Involvement of Hus1 in the chain elongation step of DNA replication after exposure to camptothecin or ionizing radiation. | Wang X et al |
| 16278671 | 2006 | The effect of Hus1 on ionizing radiation sensitivity is associated with homologous recombination repair but is independent of nonhomologous end-joining. | Wang X et al |
| 19411845 | 2009 | Cell cycle-dependent processing of DNA lesions controls localization of Rad9 to sites of genotoxic stress. | Warmerdam DO et al |
| 10921903 | 2000 | Inactivation of mouse Hus1 results in genomic instability and impaired responses to genotoxic stress. | Weiss RS et al |
| 12529385 | 2003 | Critical role for mouse Hus1 in an S-phase DNA damage cell cycle checkpoint. | Weiss RS et al |
| 11790307 | 2002 | Hus1 acts upstream of chk1 in a mammalian DNA damage response pathway. | Weiss RS et al |
| 15897895 | 2005 | Interaction and colocalization of Rad9/Rad1/Hus1 checkpoint complex with replication protein A in human cells. | Wu X et al |
| 19535328 | 2009 | Structure and functional implications of the human rad9-hus1-rad1 cell cycle checkpoint complex. | Xu M et al |
| 19918068 | 2009 | Dual inactivation of Hus1 and p53 in the mouse mammary gland results in accumulation of damaged cells and impaired tissue regeneration. | Yazinski SA et al |
| 11100718 | 2000 | The DNA damage response: putting checkpoints in perspective. | Zhou BB et al |
| 15485898 | 2004 | Human immunodeficiency virus type 1 Vpr-mediated G2 arrest requires Rad17 and Hus1 and induces nuclear BRCA1 and gamma-H2AX focus formation. | Zimmerman ES et al |
| 18156970 | 2008 | Expression of DNA damage checkpoint protein Hus1 in epithelial ovarian tumors correlates with prognostic markers. | de la Torre J et al |
Other Information
Locus ID:
NCBI: 3364
MIM: 603760
HGNC: 5309
Ensembl: ENSG00000136273
Variants:
dbSNP: 3364
ClinVar: 3364
TCGA: ENSG00000136273
COSMIC: HUS1
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 33711383 | 2021 | MiR-340-3p-HUS1 axis suppresses proliferation and migration in lung adenocarcinoma cells. | 3 |
| 33711383 | 2021 | MiR-340-3p-HUS1 axis suppresses proliferation and migration in lung adenocarcinoma cells. | 3 |
| 33137086 | 2020 | In vivo miRNA knockout screening identifies miR-190b as a novel tumor suppressor. | 9 |
| 33137086 | 2020 | In vivo miRNA knockout screening identifies miR-190b as a novel tumor suppressor. | 9 |
| 30182378 | 2019 | HUS1 checkpoint clamp component (HUS1) is a potential tumor suppressor in primary hepatocellular carcinoma. | 6 |
| 30182378 | 2019 | HUS1 checkpoint clamp component (HUS1) is a potential tumor suppressor in primary hepatocellular carcinoma. | 6 |
| 25911100 | 2015 | Genome Protection by the 9-1-1 Complex Subunit HUS1 Requires Clamp Formation, DNA Contacts, and ATR Signaling-independent Effector Functions. | 0 |
| 26088138 | 2015 | Intramolecular Binding of the Rad9 C Terminus in the Checkpoint Clamp Rad9-Hus1-Rad1 Is Closely Linked with Its DNA Binding. | 7 |
| 26377631 | 2015 | Distinct functional consequences of MUTYH variants associated with colorectal cancer: Damaged DNA affinity, glycosylase activity and interaction with PCNA and Hus1. | 16 |
| 26600172 | 2015 | The Roles of p21(Waf1/CIP1) and Hus1 in Generation and Transmission of Damage Signals Stimulated by Low-Dose Alpha-Particle Irradiation. | 1 |
| 25911100 | 2015 | Genome Protection by the 9-1-1 Complex Subunit HUS1 Requires Clamp Formation, DNA Contacts, and ATR Signaling-independent Effector Functions. | 0 |
| 26088138 | 2015 | Intramolecular Binding of the Rad9 C Terminus in the Checkpoint Clamp Rad9-Hus1-Rad1 Is Closely Linked with Its DNA Binding. | 7 |
| 26377631 | 2015 | Distinct functional consequences of MUTYH variants associated with colorectal cancer: Damaged DNA affinity, glycosylase activity and interaction with PCNA and Hus1. | 16 |
| 26600172 | 2015 | The Roles of p21(Waf1/CIP1) and Hus1 in Generation and Transmission of Damage Signals Stimulated by Low-Dose Alpha-Particle Irradiation. | 1 |
| 24062075 | 2013 | Expression of cell cycle regulatory factors hus1, gadd45a, rb1, cdkn2a and mre11a correlates with expression of clock gene per2 in human colorectal carcinoma tissue. | 8 |
Citation
Amrita Madabushi ; Randall C Gunther ; A-Lien Lu
HUS1 (HUS1 checkpoint homolog (S. pombe))
Atlas Genet Cytogenet Oncol Haematol. 2010-09-01
Online version: http://atlasgeneticsoncology.org/gene/40899/hus1
