KCMF1 (potassium channel modulatory factor 1)
2009-07-01 Roshan Mandrawalia , Ranjan Tamuli AffiliationDepartment of Biotechnology, Indian Institute of Technology Guwahati, Guwahati-781 039, Assam, India
Identity
HGNC
LOCATION
2p11.2
LOCUSID
ALIAS
DEBT91,FIGC,PCMF,ZZZ1
FUSION GENES
DNA/RNA
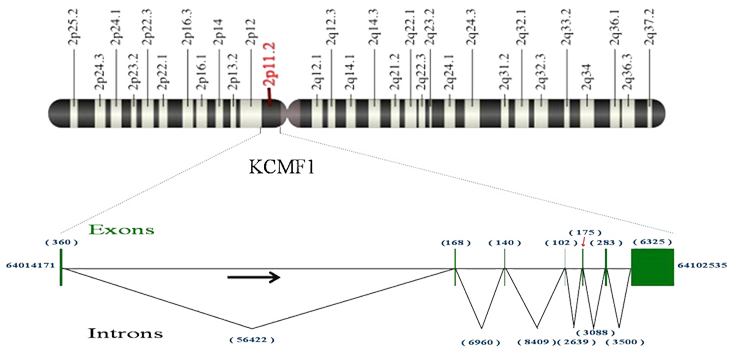
Description
DNA size 87.29 kb, mRNA size 7555 bp, 7 exons.
Proteins
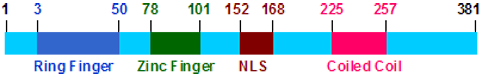
Description
381 amino acids; 41.945 kDa protein.
KCMF1 protein contains ring finger (Zinc finger, ZZ-type) 3-50 (48), zinc finger (C2H2-type) 78-101 (23), nuclear localization signal (NLS) 152-168 (17), and a coiled coil domain 225-257 (33).Isoform bApr07: This partial mRNA is 625 bp long. It is reconstructed from a myeloma cDNA clone. The pre-messenger RNA has 5 exons and covers 74.08 kb. The predicted partial protein has 208 aa (22.9 kDa, pI 5.8) and a very good coding score (7). It contains one Zinc finger, ZZ-type domain, one zinc finger, C2H2-type domain.
Isoform dApr07: The dApr07 mRNA variant is 431 bp long. It is reconstructed from a testis cDNA clone. The pre-mRNA has 4 exons and covers 12.72 kb. The predicted partial protein has 143 aa (15.7 kDa, pI 7.2) and a very good coding score (5). It contains one zinc finger, C2H2-type domain.
KCMF1 protein contains ring finger (Zinc finger, ZZ-type) 3-50 (48), zinc finger (C2H2-type) 78-101 (23), nuclear localization signal (NLS) 152-168 (17), and a coiled coil domain 225-257 (33).
Isoforms:
Two isoforms that predicted to encode proteins containing the zinc finger domain have been identified; other isoforms are relatively shorter and not well defined.
Expression
Ubiquitously expressed. High level of expression is in pharynx, thyroid, respiratory tract and larynx; less expressed in female system, uterus and cervix.
Localisation
Nuclear.
Function
KCMF1 is a transcription factor. Basic functions of the KCMF1 gene are (i) early gene up-regulation during growth factor-induced branching tubulogenesis, (ii) ubiquitination through intrinsic E3 ubiquitin ligase activity, and (iii) a possible role in ion channel activity.
Homology
The percent identity below represents identity of KCMF1 over an aligned region in UniGene.
Pan troglodytes: 97 (Percentage Identity)
Canis lupas familiaris: 91
Bos Taurus: 90
Mus musculus: 96
Gallus gallus: 93
Danio rerio: 85.
Pan troglodytes: 97 (Percentage Identity)
Canis lupas familiaris: 91
Bos Taurus: 90
Mus musculus: 96
Gallus gallus: 93
Danio rerio: 85.
Implicated in
Entity name
Ewings sarcoma family of tumors (ESFT)
Note
KCMF1 is down regulated by high constitutive CD99 (a cell surface glycoprotein) expression in ESFT. KCMF1 expression is inversely correlated with CD99 expression, as seen in a series of 22 primary ESFT. High CD99 expression levels contribute to the malignant properties of ESFT by promoting growth and migration of tumor cells.
Entity name
Gastric cancer
Note
KCMF1 (also known as FIGC) encode a RING finger protein, has intrinsic E3 ubiquitin ligase activity and promotes ubiquitination. KCMF1 contains a novel C6H2-type RING finger domain at the NH2-terminal region, consensus sequence CX2C(7-11) CX2CXA5CX2CX(5-9) HX (1-3) H (XA: acidic residues). Using differential display approach with basic fibroblast growth factor (b-FGF) inducible genes in gastric cancer cells, it was observed that FIGC upregulation in response to bFGF in gastric cancer. This suggests that FIGC might be implicated in gastric carcinogenesis through dysregulation of growth modulator.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15581609 | 2004 | FIGC, a novel FGF-induced ubiquitin-protein ligase in gastric cancers. | Jang JH et al |
| 16314831 | 2006 | Suppression of KCMF1 by constitutive high CD99 expression is involved in the migratory ability of Ewing's sarcoma cells. | Kreppel M et al |
| 12810064 | 2003 | Debt91, a putative zinc finger protein differentially expressed during epithelial morphogenesis. | Li Z et al |
Other Information
Locus ID:
NCBI: 56888
MIM: 614719
HGNC: 20589
Ensembl: ENSG00000176407
Variants:
dbSNP: 56888
ClinVar: 56888
TCGA: ENSG00000176407
COSMIC: KCMF1
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000176407 | ENST00000409785 | Q9P0J7 |
| ENSG00000176407 | ENST00000428691 | C9JSW5 |
| ENSG00000176407 | ENST00000453448 | C9J3I2 |
| ENSG00000176407 | ENST00000456682 | A0A1D5RMQ3 |
Expression (GTEx)
Pathways
| Pathway | Source | External ID |
|---|---|---|
| Immune System | REACTOME | R-HSA-168256 |
| Innate Immune System | REACTOME | R-HSA-168249 |
| Neutrophil degranulation | REACTOME | R-HSA-6798695 |
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36515749 | 2023 | KCMF1 regulates autophagy and ion channels' function in renal cell carcinoma: a future therapeutic target. | 0 |
| 36515749 | 2023 | KCMF1 regulates autophagy and ion channels' function in renal cell carcinoma: a future therapeutic target. | 0 |
| 35032791 | 2022 | CircHIPK3 contributes to human villous trophoblast growth, migration and invasion via modulating the pathway of miR-346/KCMF1. | 3 |
| 35032791 | 2022 | CircHIPK3 contributes to human villous trophoblast growth, migration and invasion via modulating the pathway of miR-346/KCMF1. | 3 |
| 25582440 | 2015 | KCMF1 (potassium channel modulatory factor 1) Links RAD6 to UBR4 (ubiquitin N-recognin domain-containing E3 ligase 4) and lysosome-mediated degradation. | 19 |
| 25582440 | 2015 | KCMF1 (potassium channel modulatory factor 1) Links RAD6 to UBR4 (ubiquitin N-recognin domain-containing E3 ligase 4) and lysosome-mediated degradation. | 19 |
| 24980667 | 2014 | MicroRNA-210 contributes to preeclampsia by downregulating potassium channel modulatory factor 1. | 42 |
| 24980667 | 2014 | MicroRNA-210 contributes to preeclampsia by downregulating potassium channel modulatory factor 1. | 42 |
| 16314831 | 2006 | Suppression of KCMF1 by constitutive high CD99 expression is involved in the migratory ability of Ewing's sarcoma cells. | 26 |
| 16314831 | 2006 | Suppression of KCMF1 by constitutive high CD99 expression is involved in the migratory ability of Ewing's sarcoma cells. | 26 |
| 15581609 | 2004 | FIGC, a novel FGF-induced ubiquitin-protein ligase in gastric cancers. | 16 |
| 15581609 | 2004 | FIGC, a novel FGF-induced ubiquitin-protein ligase in gastric cancers. | 16 |
Citation
Roshan Mandrawalia ; Ranjan Tamuli
KCMF1 (potassium channel modulatory factor 1)
Atlas Genet Cytogenet Oncol Haematol. 2009-07-01
Online version: http://atlasgeneticsoncology.org/gene/46364/kcmf1
