LGI1 (Leucine-rich, Glioma Inactivated protein 1 precursor)
2007-07-01 Nadia Gabellini AffiliationUniversity of Padua, Department of Biological Chemistry, Viale G. Colombo, 3; 35121, Padua, Italy.
Identity
HGNC
LOCATION
10q23.33
LOCUSID
ALIAS
ADLTE,ADPAEF,ADPEAF,EPITEMPIN,EPT,ETL1,IB1099
DNA/RNA
Note
LGI1 gene spans a 40,274 bp region of chromosome 10 (95,507,632 - 95,547,906)
NCBI assembly annotation: NC_000010.9; NT_030059.12.
Alternate Celera assembly: AC_000053.1; NW_924884.1.
LGI1 is considered a metastasis suppressor gene; it is also implicated in Autosomal Dominant Lateral Temporal Lobe Epilepsy (ADLTE).
NCBI assembly annotation: NC_000010.9; NT_030059.12.
Alternate Celera assembly: AC_000053.1; NW_924884.1.
LGI1 is considered a metastasis suppressor gene; it is also implicated in Autosomal Dominant Lateral Temporal Lobe Epilepsy (ADLTE).
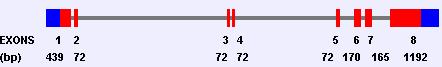
Organization of LGI1 gene, depicting isoform 1 exons (1-8);
exon size: (bp);
red: translated region;
blue: 5 UTR and 3UTR.
Description
The LGI1 gene was isolated by positional cloning from a glioblastoma cell line (T98G) bearing a balanced translocation t(10;19)(q24;q13).
LGI1 gene comprises 8 exons. Exon 1 contains the 5 UTR (224 bp) and encodes the start methione. The size of exon 8 and the position of the stop codon are different in isoform 1 and 2. The 3UTR consists of 356 bp in isoform 1 and of 386 bp in isoform 2.
A minimal promoter region is located immediately upstream of the TSS. Two Poly (A) sites are predicted by SVM from UCSC Genome Browser at the following positions: chr10:95547796-95547828 and chr10:95547881-95547919.
STS markers: IB1099; EST307318; RH51322; SHGC-155057.
LGI1 gene comprises 8 exons. Exon 1 contains the 5 UTR (224 bp) and encodes the start methione. The size of exon 8 and the position of the stop codon are different in isoform 1 and 2. The 3UTR consists of 356 bp in isoform 1 and of 386 bp in isoform 2.
A minimal promoter region is located immediately upstream of the TSS. Two Poly (A) sites are predicted by SVM from UCSC Genome Browser at the following positions: chr10:95547796-95547828 and chr10:95547881-95547919.
STS markers: IB1099; EST307318; RH51322; SHGC-155057.
Transcription
Isoform 1 mRNA is composed of 2290 bases.
Alternative splicing produces isoform 2 consisting of 1456 bases with a shorter exon 8 (425 bases):
Alternative splicing produces isoform 2 consisting of 1456 bases with a shorter exon 8 (425 bases):
Pseudogene
None.
Proteins
Note
The unprocessed precursor of LGI1 comprises 557 Amino Acids with a Molecular weight of 63818 Da (isoform 1, UniProtKB/Swiss-Prot ID: 095970). Three potential N-linked glycosilation sites have been identified at AA positions: 192, 277 and 422.
Isoform 2 includes a sequence variation (AA: 280-291) and lacks the C-terminal AA stretch (292-557) yielding a protein length of 291 AA (Isoform ID: O95970-2).
Isoform 1 is potentially secreted.
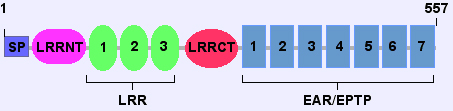
Predicted domains of LGI1 protein (isoform 1):
- signal peptide (SP, AA: 1-34);
- N-terminal LRRNT (AA: 41-71);
- LRRs domains 1-3 (AA: 90-113, 114-137 and 138-161);
- C-terminal LRRCT (AA: 173-222).
The C-terminal half includes the EAR/EPTP repeats 1-7 (AA: 224-267, 270-313, 316-364, 365-415, 418-462, 463-506, and 509-552).
Description
The N-terminal sequence of LGI1 precursor consists of a cleavable N-terminal signal peptide and of three leucine-rich repeats (LRRs) flanked by N-terminal and C-terminal cysteine-rich domains (LRRNT and LRRCT). The LRR domains are structurally similar to arcs and are generally involved in protein-protein interaction.
The C-terminal portion of LGI1 contains 7 repeats, termed Epilepsy Associated Repeats (EAR) or Epitempin (EPTP). The repeats potentially fold as beta-sheet and form a seven-bladed beta-propeller structure. Similar domains, identified in a number of proteins, probably represent protein interfaces. Isoform 2 lacks the six C-terminal EAR/EPTP domains.
The C-terminal portion of LGI1 contains 7 repeats, termed Epilepsy Associated Repeats (EAR) or Epitempin (EPTP). The repeats potentially fold as beta-sheet and form a seven-bladed beta-propeller structure. Similar domains, identified in a number of proteins, probably represent protein interfaces. Isoform 2 lacks the six C-terminal EAR/EPTP domains.
Expression
LGI1 is highly expressed in neural tissue, particularly in specific brain regions comprising both neurons and glial cells; strong expression is also reported in some areas of the prostate, kidney, sebaceous glands, islets of Langerhans, endometrium, ovary and testis.
Expression is low or absent in the majority of glioma, glioblastoma, neuroblastoma, melanoma and breast cancer cell lines. The decrease of LGI1 expression correlates with the increasing grade of malignancy in astrocytic gliomas.
DNA microarray data substantiate high expression in brain, spinal cord, DRG, and in pituitary gland.
The expression profiles by SAGE and EST number support high expression in cerebellum and cerebrum, peripheral nerve, and also in B-lymphocytes, eye, lung, muscle, testis, and thymus; low or absent expression in neoplasia and tumors.
Expression is low or absent in the majority of glioma, glioblastoma, neuroblastoma, melanoma and breast cancer cell lines. The decrease of LGI1 expression correlates with the increasing grade of malignancy in astrocytic gliomas.
DNA microarray data substantiate high expression in brain, spinal cord, DRG, and in pituitary gland.
The expression profiles by SAGE and EST number support high expression in cerebellum and cerebrum, peripheral nerve, and also in B-lymphocytes, eye, lung, muscle, testis, and thymus; low or absent expression in neoplasia and tumors.
Localisation
Isoform 1 can be secreted, whereas a shorter isoform (which might correspond to isoform 2) is retained within the cell. Some mutants of LGI1 (isoform 1) linked to ADLTE, fail to be secreted and remain in the endoplasmic reticulum and Golgi.
Function
LGI1 is involved in the control of cell proliferation, cell migration and neurogenesis. Like other neuronal LRR proteins LGI1 may modulate synaptic function.
Re-expression in LGI1-null glioblastoma cells decreases cell proliferation through the inhibition of the ERK1/2 pathway and consequent down-regulation of matrix metalloproteinases. Increased expression in neuroblastoma cells reduces proliferation and triggers intrinsic apoptosis by inhibiting the PI3K/AKT pathway.
LGI1 forms membrane complexes with Kv1.1 potassium channels within the cell antagonizing the N-type inactivation by the Kvbeta1 subunit. It has been recognized as a ligand of the trans-membrane protein receptor ADAM22, which also causes seizure when mutated.
Re-expression in LGI1-null glioblastoma cells decreases cell proliferation through the inhibition of the ERK1/2 pathway and consequent down-regulation of matrix metalloproteinases. Increased expression in neuroblastoma cells reduces proliferation and triggers intrinsic apoptosis by inhibiting the PI3K/AKT pathway.
LGI1 forms membrane complexes with Kv1.1 potassium channels within the cell antagonizing the N-type inactivation by the Kvbeta1 subunit. It has been recognized as a ligand of the trans-membrane protein receptor ADAM22, which also causes seizure when mutated.
Homology
It belongs to a family comprising four highly homologous members denoted LGI1, LGI2, LGI3, and LGI4.
LRR repeats flanked by cysteine rich regions, are also part of adhesive proteins and receptors of the LRR superfamily. With respect to this domain LGI1 is particularly related to the Drosophila protein slit, involved in growth-cone guidance and neuronal migration; and to the portion of the mammalian Trk receptors involved in neurotrophin binding. These proteins are crucial for the development of the nervous system. A comparable role for LGI1 is consistent with its involvement in epilepsy and tumors.
The C-terminal seven-fold repeat shows the largest identity with the other members of the LGI protein family, and with a segment of the G protein coupled receptor MASS1/VLGR1, which carries mutations in a mouse model of audiogenic epilepsy.
LRR repeats flanked by cysteine rich regions, are also part of adhesive proteins and receptors of the LRR superfamily. With respect to this domain LGI1 is particularly related to the Drosophila protein slit, involved in growth-cone guidance and neuronal migration; and to the portion of the mammalian Trk receptors involved in neurotrophin binding. These proteins are crucial for the development of the nervous system. A comparable role for LGI1 is consistent with its involvement in epilepsy and tumors.
The C-terminal seven-fold repeat shows the largest identity with the other members of the LGI protein family, and with a segment of the G protein coupled receptor MASS1/VLGR1, which carries mutations in a mouse model of audiogenic epilepsy.
Mutations
Note
The human LGI1 gene disclosed about 200 Single Nucleotide Polimorphism (SNP), NCBI Assembly Reference Cluster Report: rs1111820 - rs3083468.
Heterozygous point mutations are associated with ADLTE.
Large-scale homozygous mutations are linked to the development of brain malignancy.
Apparently the incidence of brain tumors is not increased in ADLTE.
Heterozygous point mutations are associated with ADLTE.
Large-scale homozygous mutations are linked to the development of brain malignancy.
Apparently the incidence of brain tumors is not increased in ADLTE.
Germinal
Several loss of function mutations (missense/nonsense, splicing, small deletions and insertions) have been reported in ADLTE patients.
Somatic
Complete loss of LGI1 expression is associated with malignant brain tumors. Rearrangement or deletion of the region 10q23-q26, following the complete loss of one copy of chromosome 10, frequently occurs in high-grade gliomas. Genetic abnormalities in this region, comprising tumor suppressor genes such as PTEN and DMBT next to the metastasis suppressor LGI1 gene, enhance the malignant progression. Even if rearrangements or mutations of LGI1 locus are absent in low-grade tumors LGI1 expression is often reduced, possibly due to epigenetic silencing.
Implicated in
Entity name
Malignant brain tumors.
Entity name
Epilepsy with auditory features (ADLTE).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 11404414 | 2001 | Repellent signaling by Slit requires the leucine-rich repeats. | Battye R et al |
| 12942323 | 2003 | Expression of the LGI1 gene product in astrocytic gliomas: downregulation with malignant progression. | Besleaga R et al |
| 14643004 | 2003 | No evidence for a seriously increased malignancy risk in LGI1-caused epilepsy. | Brodtkorb E et al |
| 9879993 | 1998 | A novel gene, LGI1, from 10q24 is rearranged and downregulated in malignant brain tumors. | Chernova OB et al |
| 16990550 | 2006 | Epilepsy-related ligand/receptor complex LGI1 and ADAM22 regulate synaptic transmission. | Fukata Y et al |
| 16518856 | 2006 | Increased expression of LGI1 gene triggers growth inhibition and apoptosis of neuroblastoma cells. | Gabellini N et al |
| 12023020 | 2002 | The LGI1 gene involved in lateral temporal lobe epilepsy belongs to a new subfamily of leucine-rich repeat proteins. | Gu W et al |
| 17565425 | 2007 | Defining the expression pattern of the LGI1 gene in BAC transgenic mice. | Head K et al |
| 11810107 | 2002 | Mutations in LGI1 cause autosomal-dominant partial epilepsy with auditory features. | Kalachikov S et al |
| 17471552 | 2007 | Leucine-rich repeat proteins of synapses. | Ko J et al |
| 7817399 | 1994 | The leucine-rich repeat: a versatile binding motif. | Kobe B et al |
| 11907806 | 2002 | Physical and functional characterization of the human LGI1 gene and its possible role in glioma development. | Krex D et al |
| 15047712 | 2004 | LGI1, a putative tumor metastasis suppressor gene, controls in vitro invasiveness and expression of matrix metalloproteinases in glioma cells through the ERK1/2 pathway. | Kunapuli P et al |
| 11606593 | 2002 | Very large G protein-coupled receptor-1, the largest known cell surface protein, is highly expressed in the developing central nervous system. | McMillan DR et al |
| 11978770 | 2002 | Mutations in the LGI1/Epitempin gene on 10q24 cause autosomal dominant lateral temporal epilepsy. | Morante-Redolat JM et al |
| 16000246 | 2005 | Differential expression of the LGI and SLIT families of genes in human cancer cells. | Rossi MR et al |
| 12095917 | 2002 | A common protein interaction domain links two recently identified epilepsy genes. | Scheel H et al |
| 16504945 | 2006 | The epilepsy-linked Lgi1 protein assembles into presynaptic Kv1 channels and inhibits inactivation by Kvbeta1. | Schulte U et al |
| 15857855 | 2005 | ADPEAF mutations reduce levels of secreted LGI1, a putative tumor suppressor protein linked to epilepsy. | Senechal KR et al |
| 17067999 | 2006 | The epilepsy gene LGI1 encodes a secreted glycoprotein that binds to the cell surface. | Sirerol-Piquer MS et al |
| 10920229 | 2000 | Identification of the promoter, genomic structure, and mouse ortholog of LGI1. | Somerville RP et al |
| 12217514 | 2002 | The novel EPTP repeat defines a superfamily of proteins implicated in epileptic disorders. | Staub E et al |
Other Information
Locus ID:
NCBI: 9211
MIM: 604619
HGNC: 6572
Ensembl: ENSG00000108231
Variants:
dbSNP: 9211
ClinVar: 9211
TCGA: ENSG00000108231
COSMIC: LGI1
RNA/Proteins
Expression (GTEx)
Pathways
| Pathway | Source | External ID |
|---|---|---|
| Developmental Biology | REACTOME | R-HSA-1266738 |
| LGI-ADAM interactions | REACTOME | R-HSA-5682910 |
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38473828 | 2024 | Leucine-Rich Glioma-Inactivated 1 (LGI1) Protein Stimulates Proliferation and IL-10 Production in Peripheral Blood Mononuclear Cells of Patients with LGI1 Antibody-Mediated Autoimmune Encephalitis In Vitro. | 0 |
| 38663634 | 2024 | The downregulation of Kv(1) channels in Lgi1(-/-)mice is accompanied by a profound modification of its interactome and a parallel decrease in Kv(2) channels. | 0 |
| 38473828 | 2024 | Leucine-Rich Glioma-Inactivated 1 (LGI1) Protein Stimulates Proliferation and IL-10 Production in Peripheral Blood Mononuclear Cells of Patients with LGI1 Antibody-Mediated Autoimmune Encephalitis In Vitro. | 0 |
| 38663634 | 2024 | The downregulation of Kv(1) channels in Lgi1(-/-)mice is accompanied by a profound modification of its interactome and a parallel decrease in Kv(2) channels. | 0 |
| 34767694 | 2022 | A novel LGI1 mutation causing autosomal dominant lateral temporal lobe epilepsy confirmed by a precise knock-in mouse model. | 3 |
| 35643238 | 2022 | Autosomal dominant lateral temporal epilepsy in a family exhibiting a rare heterozygous mutation and deletion in the leucine-rich glioma inactivated 1 gene. | 0 |
| 36304459 | 2022 | Characterization of cardiac bradyarrhythmia associated with LGI1-IgG autoimmune encephalitis. | 1 |
| 34767694 | 2022 | A novel LGI1 mutation causing autosomal dominant lateral temporal lobe epilepsy confirmed by a precise knock-in mouse model. | 3 |
| 35643238 | 2022 | Autosomal dominant lateral temporal epilepsy in a family exhibiting a rare heterozygous mutation and deletion in the leucine-rich glioma inactivated 1 gene. | 0 |
| 36304459 | 2022 | Characterization of cardiac bradyarrhythmia associated with LGI1-IgG autoimmune encephalitis. | 1 |
| 32232701 | 2021 | Clinical features of nine cases of leucine-rich glioma inactivated 1 protein antibody-associated encephalitis. | 4 |
| 32232701 | 2021 | Clinical features of nine cases of leucine-rich glioma inactivated 1 protein antibody-associated encephalitis. | 4 |
| 31900946 | 2020 | Human Cerebrospinal Fluid Monoclonal LGI1 Autoantibodies Increase Neuronal Excitability. | 48 |
| 32799011 | 2020 | Serum and CSF cytokine levels mirror different neuroimmunological mechanisms in patients with LGI1 and Caspr2 encephalitis. | 10 |
| 32911250 | 2020 | Clinical characteristics of patients double positive for CASPR2 and LGI1-antibodies. | 3 |
Citation
Nadia Gabellini
LGI1 (Leucine-rich, Glioma Inactivated protein 1 precursor)
Atlas Genet Cytogenet Oncol Haematol. 2007-07-01
Online version: http://atlasgeneticsoncology.org/gene/311/lgi1
