MIR30A (microRNA 30a)
2012-07-01 Xiao-Bin Lv AffiliationDept of Breast Surgery, Sun-Yat-Sen Memorial Hospital, Sun-Yat-Sen University, Guangzhou, China
DNA/RNA
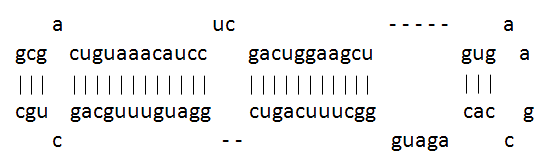
Description
miR-30 microRNA precursor is a small non-coding RNA that regulates gene expression. Animal microRNAs are transcribed as ~70 nucleotide stem-loop precursor and subsequently processed by the Dicer enzyme to give a mature ~22 nucleotide product. In this case the mature sequence comes from both the 3 (miR-30) and 5 (mir-97-6) arms of the precursor. The products are thought to have regulatory roles through complementarity to mRNA. A screen of 17 miRNAs that have been predicted to regulate a number of breast cancer associated genes found variations in the microRNAs miR-17 and miR-30c-1, these patients were noncarriers of BRCA1 or BRCA2 mutations, lending the possibility that familial breast cancer may be caused by variation in these miRNAs. Members of the miR-30 family have been found to be highly expressed in heart cells.
Transcription
miRNAs are transcribed by RNA polymerase II as part of capped and polyadenylated primary transcripts (pri-miRNAs) that can be either protein-coding or non-coding. The primary transcript is cleaved by the Drosha ribonuclease III enzyme to produce an approximately 70-nt stem-loop precursor miRNA (pre-miRNA), which is further cleaved by the cytoplasmic Dicer ribonuclease to generate the mature miRNA.
Pre-miR Length: 71 bases.
gcgactgtaa acatcctcga ctggaagctg tgaagccaca gatgggcttt cagtcggatg tttgcagctg c
Pre-miR Length: 71 bases.
gcgactgtaa acatcctcga ctggaagctg tgaagccaca gatgggcttt cagtcggatg tttgcagctg c
Pseudogene
No reported pseudogenes.
Proteins
Note
miRNAs are not translated into amino acids.
Implicated in
Entity name
Breast cancer
Disease
Overexpression of miR-30a suppressed the migration and invasiveness phenotypes of breast cancer cell lines. Moreover, reduced tumor expression of miR-30a in breast cancer patients was associated with an unfavorable outcome, including late tumor stage, lymph node metastasis, and worse progression (Cheng et al., 2012).
Prognosis
Higher expression levels of hsa-miR-30a-3p, hsa-miR-30c, and hsa-miR-182 were significantly associated with benefit of tamoxifen treatment and with longer PFS (Rodríguez-González et al., 2011).
Entity name
Colon carcinoma
Disease
miR-30a-5p was shown to be down-regulated in human colorectal cancer compared with normal colon mucosa. Overexpression of miR-30-5p suppresses proliferation colon cancer cell lines by targeting denticleless protein homolog (DTL) (Baraniskin et al., 2012).
Entity name
Non-small cell lung cancer
Disease
microRNA-30a expression was found inversely proportional to the invasive potential of various NSCLC cell lines, correlating positively with E-cadherin (epithelial marker) and negatively with N-cadherin (mesenchymal marker) expression. Lucerfise reporter assay indicates snail was a potential target of miR-30a (Kumarswamy et al., 2012).
Entity name
Squamous cell carcinoma
Note
Diagnosis: a 5-microRNA classifier (hsa-miR-210, hsa-miR-182, hsa-miR-486-5p, hsa-miR-30a, and hsa-miR-140-3p) that could distinguish SCC from normal lung tissues (Tan et al., 2011).
Entity name
Gastric cancer
Prognosis
Entity name
Thyroid carcinomas
Disease
miR-30a-5p was down-regulated in Thyroid carcinomas in comparison to normal thyroid tissue (Visone, 2007).
Entity name
Chemotherapy resistance
Note
miR-30a in regulating beclin 1 expression and autophagy reveals a novel function for miRNA in a critical cellular event with significant impacts in cancer development, progression and treatment (Zhu et al., 2009).
miR-30a can sensitize tumor cells to cis-DDP via reducing beclin 1-mediated autophagy and that increasing miR-30a level in tumor cells represents a novel approach to enhance the efficacy of chemotherapy during cancer treatment (Zou et al., 2012).
Imatinib markedly inhibits expression of miR-30a in human CML cells. miR-30a is a potent inhibitor of autophagy by downregulating Beclin 1 and ATG5 expression. miR-30a mimic or knockdown of autophagy genes (ATGs) such as Beclin 1 and ATG5 by short hairpin RNA enhances imatinib-induced cytotoxicity and promotes mitochondria-dependent intrinsic apoptosis. In contrast, knockdown of miR-30a by antagomir-30a increases the expression of Beclin 1 and ATG5, and inhibits imatinib-induced cytotoxicity (Yu et al., 2012)
miR-30a can sensitize tumor cells to cis-DDP via reducing beclin 1-mediated autophagy and that increasing miR-30a level in tumor cells represents a novel approach to enhance the efficacy of chemotherapy during cancer treatment (Zou et al., 2012).
Imatinib markedly inhibits expression of miR-30a in human CML cells. miR-30a is a potent inhibitor of autophagy by downregulating Beclin 1 and ATG5 expression. miR-30a mimic or knockdown of autophagy genes (ATGs) such as Beclin 1 and ATG5 by short hairpin RNA enhances imatinib-induced cytotoxicity and promotes mitochondria-dependent intrinsic apoptosis. In contrast, knockdown of miR-30a by antagomir-30a increases the expression of Beclin 1 and ATG5, and inhibits imatinib-induced cytotoxicity (Yu et al., 2012)
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 22476851 | 2012 | MicroRNA-30a inhibits cell migration and invasion by downregulating vimentin expression and is a potential prognostic marker in breast cancer. | Cheng CW et al |
| 21633953 | 2012 | MicroRNA-30a inhibits epithelial-to-mesenchymal transition by targeting Snai1 and is downregulated in non-small cell lung cancer. | Kumarswamy R et al |
| 19951901 | 2010 | Survival prediction of gastric cancer by a seven-microRNA signature. | Li X et al |
| 20490652 | 2011 | MicroRNA-30c expression level is an independent predictor of clinical benefit of endocrine therapy in advanced estrogen receptor positive breast cancer. | Rodríguez-González FG et al |
| 21890451 | 2011 | A 5-microRNA signature for lung squamous cell carcinoma diagnosis and hsa-miR-31 for prognosis. | Tan X et al |
| 17563749 | 2007 | Specific microRNAs are downregulated in human thyroid anaplastic carcinomas. | Visone R et al |
| 22395361 | 2012 | Targeting microRNA-30a-mediated autophagy enhances imatinib activity against human chronic myeloid leukemia cells. | Yu Y et al |
| 19535919 | 2009 | Regulation of autophagy by a beclin 1-targeted microRNA, miR-30a, in cancer cells. | Zhu H et al |
| 22157765 | 2012 | MicroRNA-30a sensitizes tumor cells to cis-platinum via suppressing beclin 1-mediated autophagy. | Zou Z et al |
Other Information
Locus ID:
NCBI: 407029
MIM: 612329
HGNC: 31624
Ensembl: ENSG00000207827
miRBase:
Variants:
dbSNP: 407029
ClinVar: 407029
TCGA: ENSG00000207827
COSMIC: MIR30A
RNA/Proteins
Pathways
| Pathway | Source | External ID |
|---|---|---|
| MicroRNAs in cancer | KEGG | hsa05206 |
| MicroRNAs in cancer | KEGG | ko05206 |
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37479901 | 2024 | Tumor-suppressive action of miR-30a-5p in lung adenocarcinoma correlates with ABL2 inhibition and PI3K/AKT pathway inactivation. | 0 |
| 38705985 | 2024 | miR-30a-5p attenuates hypoxia/reoxygenation-induced cardiomyocyte apoptosis by regulating PTEN protein expression and activating PI3K/Akt signaling pathway. | 0 |
| 38802035 | 2024 | MiR-30a-5p isoform -1|1 promotes the progression of gastric cancer by inhibiting TMEM66 and reducing intratumoral cytotoxic T cells. | 0 |
| 38821626 | 2024 | Deciphering the Role of miR-30a-5p and DLGAP1 Gene in Non-small Cell Lung Cancer. | 0 |
| 37479901 | 2024 | Tumor-suppressive action of miR-30a-5p in lung adenocarcinoma correlates with ABL2 inhibition and PI3K/AKT pathway inactivation. | 0 |
| 38705985 | 2024 | miR-30a-5p attenuates hypoxia/reoxygenation-induced cardiomyocyte apoptosis by regulating PTEN protein expression and activating PI3K/Akt signaling pathway. | 0 |
| 38802035 | 2024 | MiR-30a-5p isoform -1|1 promotes the progression of gastric cancer by inhibiting TMEM66 and reducing intratumoral cytotoxic T cells. | 0 |
| 38821626 | 2024 | Deciphering the Role of miR-30a-5p and DLGAP1 Gene in Non-small Cell Lung Cancer. | 0 |
| 35404750 | 2023 | LINC00488 Induces Tumorigenicity in Retinoblastoma by Regulating microRNA-30a-5p/EPHB2 Axis. | 1 |
| 37196609 | 2023 | MiR-30a-5p inhibits cell behaviors in esophageal cancer via modulating CBX2. | 1 |
| 37437929 | 2023 | MiR-30a-5p Enhances Cisplatin Sensitivity by Downregulating RIF1 in Ovarian Cancer. | 2 |
| 37543170 | 2023 | MiR-30a inhibits silica dust-induced epithelial-mesenchymal transition by targeting Snail. | 0 |
| 37781039 | 2023 | Exosomal miR-30a-5p promoted intrahepatic cholangiocarcinoma progression by increasing angiogenesis and vascular permeability in PDCD10 dependent manner. | 1 |
| 35404750 | 2023 | LINC00488 Induces Tumorigenicity in Retinoblastoma by Regulating microRNA-30a-5p/EPHB2 Axis. | 1 |
| 37196609 | 2023 | MiR-30a-5p inhibits cell behaviors in esophageal cancer via modulating CBX2. | 1 |
Citation
Xiao-Bin Lv
MIR30A (microRNA 30a)
Atlas Genet Cytogenet Oncol Haematol. 2012-07-01
Online version: http://atlasgeneticsoncology.org/gene/51667/mir30a
