MUC16 (mucin 16, cell surface associated)
2007-10-01 Shantibhusan Senapati , Moorthy P Ponnusamy , Ajay P Singh , Maneesh Jain , Surinder K Batra AffiliationDepartment of Biochemistry, Molecular Biology, University of Nebraska Medical Center, 985870 Nebraska Medical Center, Durham Research center 7005, Omaha, NE 68198-5870, USA (SKB)
DNA/RNA
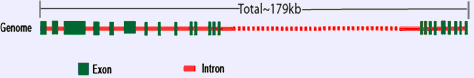
Shows the genomic organization of MUC16 gene.
Description
In the genome, MUC16 is localized in 19p13.2 chromosome and is coded by sequences present within approximatively 179kb of genomic DNA.
Transcription
As per the present available information, there is a discrepancy regarding the total number of exons present in MUC16 genomic DNA. This discrepancy is due to the absence/presence of some of the genomic sequences (particularly for the repeat regions) in the available genomic databases. The terminal nine exons on both 5 and 3 ends code for the amino- and carboxy-terminal domains of MUC16, respectively. At the same time, it has been proposed that five consecutive exons code for a single repeat unit (SRU) of the central tandem repeat domain.
Proteins
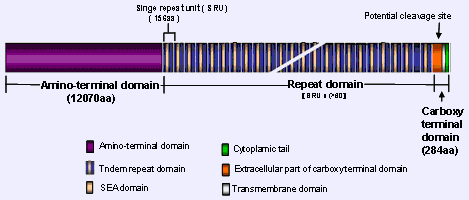
Shows the structural organization of CA125/MUC16 protein.
Description
MUC16 protein harbors a central tandem repeat region, N-terminal domain and carboxy terminal domain. The N-terminal domain has 12070 numbers of amino acids rich in serine/threonine residues and accounts for the major O-glycosylation known to be present in CA125. The MUC16 protein back bone is dominated by tandem repeat region, which has more than 60 repeat domains, each composed of 156 amino acids. Though all the individual repeat units are not similar, most of them occur more than once in the sequence.
The repeat units are rich in serine, threonine and proline residues, which are typical for any mucins. Each repeat unit has some homology to the SEA (Sea-urchin sperm protein, Enterokinase and Agrin) module, whose exact biological function is not known.
The epitopes for known anti-CA125 antibodies (OC125 and M11) are thought to be present on a small cysteine ring region present in the tandem-repeat region of MUC16.
The carboxy-terminal domain has 284 aminoacids and can be divided into three different regions: extra cellular, transmembrane and cytoplasmic tail. The extracellular part of the carboxy-terminal domain has many N-glycosylation sites and some O-glycosyaltion sites. Several in silico analyses suggest a putative cleavage site in the extracellular part of carboxy-terminal domain. The MUC16 cytoplasmic tail is 31 amino acids long and has many possible phosphorylation sites.
The phosphorylation of CA125 in WISH cells has been reported by labeling with 32PO43- and immunoprecipitaion analysis but the exact site of phosphorylation is yet to be mapped. Interestingly, CA125 contains a putative tyrosine phosphorylation site (RRKKEGY), which was first recognized in Src family protein. This sequence is conserved in the translated mouse EST (AK003577) that has homology with CA125/MUC16 at the C-terminal end. Recently, it has been shown that MUC16 cytoplasmic tail, which contains a polybasic aminoacid sequence, interacts with cytoskeleton through ERM (ezrin/radixin/moesin) actin-binding proteins.
The repeat units are rich in serine, threonine and proline residues, which are typical for any mucins. Each repeat unit has some homology to the SEA (Sea-urchin sperm protein, Enterokinase and Agrin) module, whose exact biological function is not known.
The epitopes for known anti-CA125 antibodies (OC125 and M11) are thought to be present on a small cysteine ring region present in the tandem-repeat region of MUC16.
The carboxy-terminal domain has 284 aminoacids and can be divided into three different regions: extra cellular, transmembrane and cytoplasmic tail. The extracellular part of the carboxy-terminal domain has many N-glycosylation sites and some O-glycosyaltion sites. Several in silico analyses suggest a putative cleavage site in the extracellular part of carboxy-terminal domain. The MUC16 cytoplasmic tail is 31 amino acids long and has many possible phosphorylation sites.
The phosphorylation of CA125 in WISH cells has been reported by labeling with 32PO43- and immunoprecipitaion analysis but the exact site of phosphorylation is yet to be mapped. Interestingly, CA125 contains a putative tyrosine phosphorylation site (RRKKEGY), which was first recognized in Src family protein. This sequence is conserved in the translated mouse EST (AK003577) that has homology with CA125/MUC16 at the C-terminal end. Recently, it has been shown that MUC16 cytoplasmic tail, which contains a polybasic aminoacid sequence, interacts with cytoskeleton through ERM (ezrin/radixin/moesin) actin-binding proteins.
Expression
The expression of MUC16 has been reported in human epithelia of conjunctiva, cornea, middle ear and trachea under normal physiological conditions. MUC16 is also expressed in ovarian carcinoma.
Localisation
It is a type I membrane-bound protein and due to cleavage gets secreted into the extracellular space. On the ocular surface, MUC16 is expressed on the tips of the microplicae of the ocular surface.
Function
MUC16 provides a disadhesive protective barrier to the ocular epithelial surface. Overexpression of CA125/MUC16 in ovarian cancer indicates its possible role in cancer pathogenesis. Studies have shown that CA125/MUC16 binds to mesothelin and galectin-1, which are overexpressed in ovarian cancer. It has also been shown that mesothelin-MUC16 interaction has significance in adhesion of ovarian cancer cells to mesothelial cells present on the inner wall of the peritoneum and on the surface of other abdominal organs. This cell to cell adhesion may help in ovarian cancer metastasis. It has been proposed that galectin-1 bound to MUC16 may cause apoptosis of T cells, and thus help in the suppression of the host immunity.
Homology
Similar to mucin 16 of Pan troglodytes, Canis lupus familiaris, Mus musculus, Rattus norvegicus and Gallus gallus.
Implicated in
Entity name
Ovarian cancer
Disease
Epithelial ovarian cancer is the most lethal gynaecologic malignancy in the United States and other parts of the world. In the United States, ovarian cancer accounts for approximately 22,000 new cases and 16,000 deaths occurring every year. The epithelial ovarian carcinomas represent approximately 90% of all types of ovarian malignant neoplasms. Due to lack of specific signs and symptoms of this disease, coupled with lack of reliable screening strategies most patients are diagnosed in the advanced stage of the disease, resulting in low overall cure rates. Ovarian cancer patients are generally treated with surgical resection and subsequent platinum-based chemotherapy. Although, many patients initially respond well to chemotherapy, long term survival remains poor due to eventual tumor recurrence and emergence of drug-resistant disease. Overall, the five year survival rate is 45%.
Prognosis
Since the last 20 years, CA125/MUC16 has been used as a well-established marker for diagnosis of ovarian cancer. It is mostly overexpressed in serous type of ovarian cancers and less likely to be expressed in mucinous tumors. More than 80% of ovarian cancer patients have elevated CA125 level during their treatment period. It has been shown that the disease progression is associated with an increase in serum CA125 level, while a decline in serum CA125 level is associated with response to therapy. In another finding, it has been shown that the trend of serum CA125 level during the first three courses of chemotherapy is a strong forecaster of re-examination findings in patients with ovarian carcinoma at the end of treatment. Interestingly, it has been shown that a normal CA125 level by the end of second or third chemotherapy is strongly linked to the survival of patients in stage 3 or stage 4 conditions. Also, variations in the CA125 value even within the normal range carry useful information regarding prediction of time to treatment failure. Additionally, in patients in stage 1 cancers it has been suggested that CA125 elevations are not related to the tumor mass volume. Recently, the potential of CA125/MUC16 as a therapeutic target has been harnessed by using an armed human antibody (3A5) against MUC16 protein.
Oncogenesis
There is no experimental evidence in the scientific literature for a role of MUC16 in oncogenesis. However, MUC16 possesses many structural similarities with other membrane bound mucins, like MUC1 and MUC4, which are already shown to be functionally involved in different cancers. Transmembrane mucins are hypothesized to serve as sensors of the external environment and can transduce signals via the post-translational modifications of their cytoplasmic tail. Phosphorylation of MUC16 protein has already been reported. Though the exact interacting partner and the site of phosphorylation are unknown, the presence of potential phosphorylation sites in MUC16 cytoplasmic tail indicates the possible role of MUC16 in downstream signal transduction. Further, it has been shown that MUC16 interacts with galectin-1 and mesothelin and these interactions may have a role in cancer progression.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 17898272 | 2007 | Functions of MUC16 in corneal epithelial cells. | Blalock TD et al |
| 16880776 | 2006 | Aberrant expression of MUC4 in ovarian carcinoma: diagnostic significance alone and in combination with MUC1 and MUC16 (CA125). | Chauhan SC et al |
| 17510422 | 2007 | Armed antibodies targeting the mucin repeats of the ovarian cancer antigen, MUC16, are highly efficacious in animal tumor models. | Chen Y et al |
| 16848908 | 2006 | Methylation mediated silencing of TMS1/ASC gene in prostate cancer. | Das PM et al |
| 17604678 | 2007 | MUC16 is produced in tracheal surface epithelium and submucosal glands and is present in secretions from normal human airway and cultured bronchial epithelial cells. | Davies JR et al |
| 16500040 | 2006 | Distinct evolution of the human carcinoma-associated transmembrane mucins, MUC1, MUC4 AND MUC16. | Duraisamy S et al |
| 14764598 | 2004 | Solution structure of the SEA domain from the murine homologue of ovarian cancer antigen CA125 (MUC16). | Maeda T et al |
| 12218296 | 2002 | The CA 125 gene: a newly discovered extension of the glycosylated N-terminal domain doubles the size of this extracellular superstructure. | O'Brien TJ et al |
| 12615972 | 2003 | The cancer antigen CA125 represents a novel counter receptor for galectin-1. | Seelenmeyer C et al |
| 11920644 | 2002 | Ovarian cancer antigen CA125 is encoded by the MUC16 mucin gene. | Yin BW et al |
| 11369781 | 2001 | Molecular cloning of the CA125 ovarian cancer antigen: identification as a new mucin, MUC16. | Yin BW et al |
Other Information
Locus ID:
NCBI: 94025
MIM: 606154
HGNC: 15582
Ensembl: ENSG00000181143
Variants:
dbSNP: 94025
ClinVar: 94025
TCGA: ENSG00000181143
COSMIC: MUC16
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000181143 | ENST00000397910 | Q8WXI7 |
| ENSG00000181143 | ENST00000596768 | M0R2Y5 |
| ENSG00000181143 | ENST00000599436 | M0R2S7 |
| ENSG00000181143 | ENST00000601404 | M0QZZ9 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
PharmGKB
| Entity ID | Name | Type | Evidence | Association | PK | PD | PMIDs |
|---|---|---|---|---|---|---|---|
| PA445048 | Nausea | Disease | ClinicalAnnotation | associated | PD | ||
| PA446051 | Vomiting | Disease | ClinicalAnnotation | associated | PD | ||
| PA452618 | opioids | Chemical | ClinicalAnnotation | associated | PD |
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38285795 | 2024 | Human Epididymis Protein 4 (HE4) and Cancer Antigen 125 (CA125) for Prediction of Optimal Primary Surgery in Non-Mucinous Epithelial Ovarian Cancer. | 0 |
| 38285795 | 2024 | Human Epididymis Protein 4 (HE4) and Cancer Antigen 125 (CA125) for Prediction of Optimal Primary Surgery in Non-Mucinous Epithelial Ovarian Cancer. | 0 |
| 35705629 | 2023 | CA125: a novel cardiac biomarker for infants with congenital diaphragmatic hernia. | 1 |
| 36208679 | 2023 | Prognostic value of carbohydrate antigen 125 in patients undergoing surgical left ventricular reconstruction. | 0 |
| 36436416 | 2023 | MUC16 mutation is associated with tumor grade, clinical features, and prognosis in glioma patients. | 5 |
| 36586479 | 2023 | Cancer Antigen 125 Expression Enhances the Gemcitabine/Cisplatin-Resistant Tumor Microenvironment in Bladder Cancer. | 4 |
| 36596280 | 2023 | The Utility of Preoperative Serum CA125 Combined with HE4 to Predict Lymph Node Metastasis in Endometrial Cancer. | 1 |
| 36918912 | 2023 | MUC16 promotes triple-negative breast cancer lung metastasis by modulating RNA-binding protein ELAVL1/HUR. | 7 |
| 36928256 | 2023 | Diffusion-weighted magnetic resonance sequence and CA125/CEA ratio can be used as add-on tools to ultrasound for the differentiation of ovarian from non-ovarian pelvic masses. | 1 |
| 37129994 | 2023 | Radiologically enlarged cardiophrenic lymph nodes and CA-125 in relation to diaphragmatic carcinomatosis, surgical outcome, and overall survival in advanced ovarian cancer. | 1 |
| 37341998 | 2023 | Development and validation of a MUC16 mutation-associated immune prognostic model for lung adenocarcinoma. | 0 |
| 37549441 | 2023 | Serum CA-125 as a predictor in the early diagnosis of ectopic pregnancy in Vietnam - A case-control study. | 0 |
| 37603503 | 2023 | Diagnostic value of serum human epididymis protein 4, carbohydrate antigen 125 and their combination in endometrial cancer: A meta-analysis. | 2 |
| 37644468 | 2023 | MUC16 stimulates neutrophils to an inflammatory and immunosuppressive phenotype in ovarian cancer. | 6 |
| 37932988 | 2023 | Comprehensive characterization of MUC16 mutations in lung adenocarcinoma for immunotherapies and prognosis: An observational study. | 0 |
Citation
Shantibhusan Senapati ; Moorthy P Ponnusamy ; Ajay P Singh ; Maneesh Jain ; Surinder K Batra
MUC16 (mucin 16, cell surface associated)
Atlas Genet Cytogenet Oncol Haematol. 2007-10-01
Online version: http://atlasgeneticsoncology.org/gene/41455/muc16
