MYBBP1A (MYB binding protein (P160) 1a)
2011-01-01 Claudia Perrera , Riccardo Colombo AffiliationDepartment of Cell Biology-Oncology, Nerviano Medical Sciences, Viale Pasteur 10, Nerviano 20014, Italy
Identity
HGNC
LOCATION
17p13.2
LOCUSID
ALIAS
P160,PAP2,Pol5
FUSION GENES
DNA/RNA
Note
Total gene length 16491 bp, mRNA length 4122 bp, telomeric to SPNS2 and centromeric to GGT6. Two variants described. MYBBP1A gene is composed by 26 exones and 28 introns.
Description
The human MYBBP1A gene is located on chromosome 17p13.3 (Keough et al., 1999).
Transcription
The complete MYBBP1A cDNA is 4518 bp, including 25 bp of 5 UTR and 506 bp of 3 UTR up to the polyA tail.
Proteins
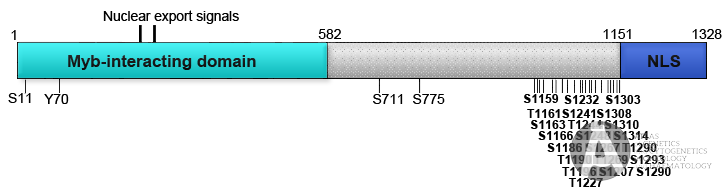
Schematic representation of Mybbp1A protein. aa 1-582 is the domain reported to interact in vitro with Myb. NLS: Nuclear and nucleolar localisation signal. The indicated S, T and Y are phosphorylated residues identified in several phospho-proteomic studies.
Description
Human MYBBP1A is a 1328 aa long protein. Murine MYBBP1A was originally identified as a protein interacting with the leucine zipper of c-Myb (Favier et al., 1994). Subsequently, in 1998, the human gene homologue of MYBBP1A was cloned and its chromosomal location mapped to 17p13.3 (Keough et al., 1999).
Expression
MYBBP1A is ubiquitously expressed (Tavner et al., 1998).
Localisation
MYBBP1A is a nuclear protein, predominantly localized in the nucleolus (Keough et al., 2003). MYBBP1A has been confirmed as a resident protein of the nucleolus by three large-scale proteomic studies that have established a protein inventory of this sub-nuclear compartment (Andersen et al., 2002; Scherl et al., 2002; Andersen et al., 2005). The nuclear/nucleolar localization signals are present in the C-terminal tail of MYBBP1A.
Function
MYBBP1A functions have not yet been completely clarified. It was originally identified as a protein able to interact with the negative regulatory domain (NRD) of c-Myb; however, it was later shown to lack any significant effect in a Myb-dependent transcription reporter assay (Favier et al., 1994). MYBBP1A has been found to interact with and regulate several transcription factors: it binds and represses both Prep1-Pbx1, involved in development and organogenesis, and also PGC-1a, a key regulator of metabolic processes such as mitochondrial biogenesis and respiration and gluconeogenesis in liver (Fan et al., 2004; Diaz et al., 2007). MYBBP1A acts as a co-repressor for RelA/p65, a member of the NFkB family, by competing with the co-activator p300 histone acetyltrasferase for interaction with the transcription activation domain (TAD) of RelA/p65. It is also a co-repressor on the Period2 promoter, repressing the expression of Per2, an essential gene in the regulation of the circadian clock (Owen et al., 2007; Hara et al., 2009). Conversely, MYBBP1A is a positive regulator of the aromatic hydrocarbon receptor (AhR) which mediates transcriptional responses to certain hydrophobic ligands, such as dioxin, by enhancing the ability of AhR to activate transcription (Jones et al., 2002).
MYBBP1A has also been reported to be a component of macromolecular complexes such as the B-WICH complex, a 3 MDa assembly made of proteins and RNAs, formed during active transcription (Cavellan et al., 2006) or part of large interactomes such as the SMN interactome (Fuller et al., 2010).
MYBBP1A can be post-translationally processed in some type of cells to generate an amino-terminal fragments of 67 kDa (p67). Ribosomal stress induced by Actinomycin D (an inhibitor of ribosome biogenesis) treatment causes MYBBP1A processing and translocation from the nucleolus to the nucloplasm (Diaz et al., 2007; Yamauchi et al., 2008), indicating a possible MYBBP1A role in ribosome biogenesis.
Several post-translational modifications have been described for MYBBP1A, even if their biological significance is not yet clarified. MYBBP1A is reported to be a heavily phosphorylated protein in cells, according to several large-scale mass spectrometry-based phosphoproteomic studies (Beausoleil et al., 2004; Beausoleil et al., 2006; Nousiainen et al., 2006; Olsen et al., 2006; Cantin et al., 2008; Daub et al., 2008; Dephoure et al., 2008; Imami et al., 2008). The majority of the phosphosites mapped in MYBBP1A in these studies (18 out of a total of 21) reside within the ~200 amino-acid long C-terminal portion of the protein, which has been shown to be relevant for its nuclear and nucleolar localization (Keough et al., 2003). Notably, MYBBP1A was also found to be also a component of the proteome as well as the phospho-proteome of the human mitotic spindle. MYBBP1A contains several consensus motifs for several kinases, but until now, only Ser1303 has been proven in vitro and in HeLa cells to be indeed phosphorylated by Aurora B kinase (Perrera et al., 2010). In this work, it has been shown that MYBBP1A depletion by RNAi causes a delay in progression through mitosis and defects in mitotic spindle assembly and stability, indicating that, like other nucleolar proteins, MYBBP1A may have a role in insuring correct mitotic progression (Perrera et al., 2010).
MYBBP1A has been reported to be also sumoylated upon MG132 treatment (Matafora et al., 2009).
MYBBP1A has also been reported to be a component of macromolecular complexes such as the B-WICH complex, a 3 MDa assembly made of proteins and RNAs, formed during active transcription (Cavellan et al., 2006) or part of large interactomes such as the SMN interactome (Fuller et al., 2010).
MYBBP1A can be post-translationally processed in some type of cells to generate an amino-terminal fragments of 67 kDa (p67). Ribosomal stress induced by Actinomycin D (an inhibitor of ribosome biogenesis) treatment causes MYBBP1A processing and translocation from the nucleolus to the nucloplasm (Diaz et al., 2007; Yamauchi et al., 2008), indicating a possible MYBBP1A role in ribosome biogenesis.
Several post-translational modifications have been described for MYBBP1A, even if their biological significance is not yet clarified. MYBBP1A is reported to be a heavily phosphorylated protein in cells, according to several large-scale mass spectrometry-based phosphoproteomic studies (Beausoleil et al., 2004; Beausoleil et al., 2006; Nousiainen et al., 2006; Olsen et al., 2006; Cantin et al., 2008; Daub et al., 2008; Dephoure et al., 2008; Imami et al., 2008). The majority of the phosphosites mapped in MYBBP1A in these studies (18 out of a total of 21) reside within the ~200 amino-acid long C-terminal portion of the protein, which has been shown to be relevant for its nuclear and nucleolar localization (Keough et al., 2003). Notably, MYBBP1A was also found to be also a component of the proteome as well as the phospho-proteome of the human mitotic spindle. MYBBP1A contains several consensus motifs for several kinases, but until now, only Ser1303 has been proven in vitro and in HeLa cells to be indeed phosphorylated by Aurora B kinase (Perrera et al., 2010). In this work, it has been shown that MYBBP1A depletion by RNAi causes a delay in progression through mitosis and defects in mitotic spindle assembly and stability, indicating that, like other nucleolar proteins, MYBBP1A may have a role in insuring correct mitotic progression (Perrera et al., 2010).
MYBBP1A has been reported to be also sumoylated upon MG132 treatment (Matafora et al., 2009).
Homology
Orthologous genes for MYBBP1A sharing a high degree of similarity are present in rat and mice. Protein homologues have also been recognized in dog, bovine, and chicken and a MYBBP1A-like protein spanning 1269 residues and showing a 60% similarity to the human protein has been identified in zebrafish, suggesting that MYBBP1A is significantly conserved across vertebrate species (Amsterdam et al., 2004). MYBBP1A shares some homology to a yeast protein called POL5, reported to be an essential DNA polymerase in Saccharomyces cerevisiae (Yang et al., 2003).
Implicated in
Entity name
Various cancers
Disease
MYBBP1A maps at 17p13.3, a region frequently lost in many solid and haematological tumors, such as breast and ovarian cancer, medulloblastoma, astrocytoma, leukemias, etc. This indicates that this chromosomal band contains one or more tumor suppressor genes. However, MYBBP1A is unlikely a candidate for being a tumor suppressor gene, as it lies centromeric to the regions of LOH described (Keough et al., 1999).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15256591 | 2004 | Identification of 315 genes essential for early zebrafish development. | Amsterdam A et al |
| 15635413 | 2005 | Nucleolar proteome dynamics. | Andersen JS et al |
| 16964243 | 2006 | A probability-based approach for high-throughput protein phosphorylation analysis and site localization. | Beausoleil SA et al |
| 18220336 | 2008 | Combining protein-based IMAC, peptide-based IMAC, and MudPIT for efficient phosphoproteomic analysis. | Cantin GT et al |
| 16603771 | 2006 | The WSTF-SNF2h chromatin remodeling complex interacts with several nuclear proteins in transcription. | Cavellán E et al |
| 18691976 | 2008 | Kinase-selective enrichment enables quantitative phosphoproteomics of the kinome across the cell cycle. | Daub H et al |
| 18669648 | 2008 | A quantitative atlas of mitotic phosphorylation. | Dephoure N et al |
| 17875935 | 2007 | p160 Myb-binding protein interacts with Prep1 and inhibits its transcriptional activity. | Díaz VM et al |
| 14744933 | 2004 | Suppression of mitochondrial respiration through recruitment of p160 myb binding protein to PGC-1alpha: modulation by p38 MAPK. | Fan M et al |
| 8302594 | 1994 | Detection of proteins that bind to the leucine zipper motif of c-Myb. | Favier D et al |
| 19928837 | 2010 | The SMN interactome includes Myb-binding protein 1a. | Fuller HR et al |
| 19129230 | 2009 | Molecular characterization of Mybbp1a as a co-repressor on the Period2 promoter. | Hara Y et al |
| 18187866 | 2008 | Automated phosphoproteome analysis for cultured cancer cells by two-dimensional nanoLC-MS using a calcined titania/C18 biphasic column. | Imami K et al |
| 11956195 | 2002 | Myb-binding protein 1a augments AhR-dependent gene expression. | Jones LC et al |
| 10644447 | 1999 | Molecular cloning and chromosomal mapping of the human homologue of MYB binding protein (P160) 1A (MYBBP1A) to 17p13.3. | Keough R et al |
| 12941609 | 2003 | Myb-binding protein 1a is a nucleocytoplasmic shuttling protein that utilizes CRM1-dependent and independent nuclear export pathways. | Keough RA et al |
| 19596686 | 2009 | Proteomics analysis of nucleolar SUMO-1 target proteins upon proteasome inhibition. | Matafora V et al |
| 16565220 | 2006 | Phosphoproteome analysis of the human mitotic spindle. | Nousiainen M et al |
| 17081983 | 2006 | Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. | Olsen JV et al |
| 17196614 | 2007 | MYBBP1a is a novel repressor of NF-kappaB. | Owen HR et al |
| 20177074 | 2010 | Identification of Myb-binding protein 1A (MYBBP1A) as a novel substrate for aurora B kinase. | Perrera C et al |
| 12429849 | 2002 | Functional proteomic analysis of human nucleolus. | Scherl A et al |
| 9447996 | 1998 | Molecular cloning reveals that the p160 Myb-binding protein is a novel, predominantly nucleolar protein which may play a role in transactivation by Myb. | Tavner FJ et al |
| 18173745 | 2008 | Ribosomal stress induces processing of Mybbp1a and its translocation from the nucleolus to the nucleoplasm. | Yamauchi T et al |
| 12695662 | 2003 | Yeast POL5 is an evolutionarily conserved regulator of rDNA transcription unrelated to any known DNA polymerases. | Yang W et al |
Other Information
Locus ID:
NCBI: 10514
MIM: 604885
HGNC: 7546
Ensembl: ENSG00000132382
Variants:
dbSNP: 10514
ClinVar: 10514
TCGA: ENSG00000132382
COSMIC: MYBBP1A
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37298588 | 2023 | Intratumoral Restoration of miR-137 Plus Cholesterol Favors Homeostasis of the miR-137/Coactivator p160/AR Axis and Negatively Modulates Tumor Progression in Advanced Prostate Cancer. | 1 |
| 37298588 | 2023 | Intratumoral Restoration of miR-137 Plus Cholesterol Favors Homeostasis of the miR-137/Coactivator p160/AR Axis and Negatively Modulates Tumor Progression in Advanced Prostate Cancer. | 1 |
| 35485735 | 2022 | Characterization of the impact of the MYBBP1A gene and rs3809849 on asparaginase sensitivity and cellular functions. | 0 |
| 35485735 | 2022 | Characterization of the impact of the MYBBP1A gene and rs3809849 on asparaginase sensitivity and cellular functions. | 0 |
| 31066170 | 2019 | c-MYB- and PGC1a-dependent metabolic switch induced by MYBBP1A loss in renal cancer. | 13 |
| 31066170 | 2019 | c-MYB- and PGC1a-dependent metabolic switch induced by MYBBP1A loss in renal cancer. | 13 |
| 25612917 | 2016 | The effects of SP110's associated genes on fresh cavitary pulmonary tuberculosis in Han Chinese population. | 6 |
| 25612917 | 2016 | The effects of SP110's associated genes on fresh cavitary pulmonary tuberculosis in Han Chinese population. | 6 |
| 25543088 | 2015 | Regulation and function of Myb-binding protein 1A (MYBBP1A) in cellular senescence and pathogenesis of head and neck cancer. | 9 |
| 26044764 | 2015 | Gradual reduction in rRNA transcription triggers p53 acetylation and apoptosis via MYBBP1A. | 18 |
| 25543088 | 2015 | Regulation and function of Myb-binding protein 1A (MYBBP1A) in cellular senescence and pathogenesis of head and neck cancer. | 9 |
| 26044764 | 2015 | Gradual reduction in rRNA transcription triggers p53 acetylation and apoptosis via MYBBP1A. | 18 |
| 24375404 | 2014 | The nucleolar protein Myb-binding protein 1A (MYBBP1A) enhances p53 tetramerization and acetylation in response to nucleolar disruption. | 21 |
| 25303791 | 2014 | TPPII, MYBBP1A and CDK2 form a protein-protein interaction network. | 1 |
| 24375404 | 2014 | The nucleolar protein Myb-binding protein 1A (MYBBP1A) enhances p53 tetramerization and acetylation in response to nucleolar disruption. | 21 |
Citation
Claudia Perrera ; Riccardo Colombo
MYBBP1A (MYB binding protein (P160) 1a)
Atlas Genet Cytogenet Oncol Haematol. 2011-01-01
Online version: http://atlasgeneticsoncology.org/gene/41467/mybbp1a
