NKX3-1 (NK3 homeobox 1)
2009-03-01 Liang-Nian Song , Edward P Gelmann AffiliationHerbert Irving Comprehensive Cancer Center, Columbia University, New York, NY 10032, USA
Identity
HGNC
LOCATION
8p21.2
LOCUSID
ALIAS
BAPX2,NKX3,NKX3.1,NKX3A
FUSION GENES
DNA/RNA

The gene for NKX3-1 comprises two exons of 334 and 2947 bp, respectively. The length of the intron is 964 bp. Positions of start and stop codons are indicated.
Description
The gene has two exons and one intron.
Transcription
Transcription takes place in a centromere --> telomere orientation. The length of the processed mRNA is about 3200 bp.
Pseudogene
Not known.
Proteins
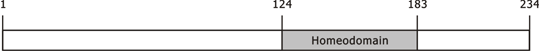
NKX3-1 contains two exons encoding a 234-amino acid protein including a homeodomain (grey).
Description
234 amino acids; 35-38 kDa, contains one N-terminal domain (residues 1-123), one homeodomain (residues 124-183), and one C-terminal domain (residues 184-234).
Expression
Expression is restricted to the adult murine prostate and bulbourethral gland. During early murine embryogenesis NKX3-1 expression has also been detected in developing somites and testes. In the adult human expression is seen in prostate epithelium, testis, ureter, and pulmonary bronchial mucous glands.
Localisation
Nuclear.
Function
Binds to DNA to suppress transcription. Interacts with transcription factors, e.g. serum response factor, to enhance transcriptional activation. Binds to and potentiates topoisomerase I DNA resolving activity. Acts as prostate tumor suppressor.
Homology
Homeodomain protein with membership of the NKX family.
Mutations
Germinal
Twenty-one germ-line variants have been identified in 159 probands of hereditary prostate cancer families. These variants were linked to prostate cancer risk in hereditary prostate cancer families. For example, the C154T (11% of the population) polymorphism mutation is associated with prostatic enlargement and prostate cancer risk. A T164A mutations in one family cosegregates with prostate cancer in three affected brothers. For a more complete list of identified mutations, please visit http://cancerres.aacrjournals.org/cgi/content/full/66/1/69.
Somatic
None.
Implicated in
Entity name
Prostate Cancer
Disease
Prostate cancer is the most commonly diagnosed cancer in American men and the second leading cause of cancer-related deaths. Prostate cancer predominantly occurs in the peripheral zone of the human prostate, with roughly 5 to 10% of cases found in the central zone. Disease development involves the temporal and spatial loss of the basal epithelial compartment accompanied by increased proliferation and dedifferentiation of the luminal (secretory) epithelial cells. Prostate cancer is a slow developing disease that is typically found in men greater than 60 years of age and incidence increases with increasing age.
Prognosis
PSA test combined with digital-rectal exams are used to screen for the presence of disease. If the digital-rectal exams are positive, additional tests including needle core biopsies are taken to assess disease stage and grade. Patients with localized, prostate-restricted disease are theoretically curable with complete removal of the prostate (radical prostatectomy). Patients with extra-prostatic disease are treated with hormone (androgen ablation) therapy, radiation, and/or antiandrogens; however, no curative treatments are available for nonorgan confined, metastatic disease.
Cytogenetics
Various forms of aneuploidy.
Oncogenesis
Nkx3.1 plays an essential role in normal murine prostate development. Loss of function of Nkx3.1 leads to defects in prostatic protein secretions and in ductal morphogenesis. Loss-of-function of Nkx3.1 also contributes to prostate carcinogenesis. For example, Nkx3.1 mutant mice develop prostatic dysplasia. Nkx3.1 loss potentiates prostate carcinogenesis in a Pten+/- background. Furthermore, by a variety of mechanisms NKX3.1 expression is reduced in noninvasive and early stage human prostate cancer, suggesting that its decreased expression is one of the earliest steps in the majority of human prostate cancers.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 18557759 | 2008 | Integrating differentiation and cancer: the Nkx3.1 homeobox gene in prostate organogenesis and carcinogenesis. | Abate-Shen C et al |
| 11839815 | 2002 | Conditional loss of Nkx3.1 in adult mice induces prostatic intraepithelial neoplasia. | Abdulkadir SA et al |
| 15734999 | 2005 | Deletion, methylation, and expression of the NKX3.1 suppressor gene in primary human prostate cancer. | Asatiani E et al |
| 17108105 | 2006 | Decreased NKX3.1 protein expression in focal prostatic atrophy, prostatic intraepithelial neoplasia, and adenocarcinoma: association with gleason score and chromosome 8p deletion. | Bethel CR et al |
| 10215624 | 1999 | Roles for Nkx3.1 in prostate development and cancer. | Bhatia-Gaur R et al |
| 11085535 | 2000 | Loss of NKX3.1 expression in human prostate cancers correlates with tumor progression. | Bowen C et al |
| 17234752 | 2007 | NKX3.1 homeodomain protein binds to topoisomerase I and enhances its activity. | Bowen C et al |
| 12661036 | 2003 | Expression of NKX3.1 in normal and malignant tissues. | Gelmann EP et al |
| 9226374 | 1997 | A novel human prostate-specific, androgen-regulated homeobox gene (NKX3.1) that maps to 8p21, a region frequently deleted in prostate cancer. | He WW et al |
| 18794125 | 2008 | Nkx3-1 and LEF-1 function as transcriptional inhibitors of estrogen receptor activity. | Holmes KA et al |
| 16814806 | 2006 | Physical and functional interactions between the prostate suppressor homeoprotein NKX3.1 and serum response factor. | Ju JH et al |
| 11854455 | 2002 | Cooperativity of Nkx3.1 and Pten loss of function in a mouse model of prostate carcinogenesis. | Kim MJ et al |
| 15311057 | 2004 | Analysis of androgen regulated homeobox gene NKX3.1 during prostate carcinogenesis. | Korkmaz CG et al |
| 11137288 | 2000 | Full-length cDNA sequence and genomic organization of human NKX3A - alternative forms and regulation by both androgens and estrogens. | Korkmaz KS et al |
| 16581776 | 2006 | NKX3.1 is regulated by protein kinase CK2 in prostate tumor cells. | Li X et al |
| 12676585 | 2003 | Haploinsufficiency at the Nkx3.1 locus. A paradigm for stochastic, dosage-sensitive gene regulation during tumor initiation. | Magee JA et al |
| 18757402 | 2008 | Inflammatory cytokines induce phosphorylation and ubiquitination of prostate suppressor protein NKX3.1. | Markowski MC et al |
| 17602165 | 2007 | Haploinsufficient prostate tumor suppression by Nkx3.1: a role for chromatin accessibility in dosage-sensitive gene regulation. | Mogal AP et al |
| 11257711 | 2001 | Expression studies and mutational analysis of the androgen regulated homeobox gene NKX3.1 in benign and malignant prostate epithelium. | Ornstein DK et al |
| 9537602 | 1998 | Isolation and androgen regulation of the human homeobox cDNA, NKX3.1. | Prescott JL et al |
| 16442598 | 2006 | Effect of homeodomain protein NKX3.1 R52C polymorphism on prostate gland size. | Rodriguez Ortner E et al |
| 10906459 | 2000 | Targeted disruption of the Nkx3.1 gene in mice results in morphogenetic defects of minor salivary glands: parallels to glandular duct morphogenesis in prostate. | Schneider A et al |
| 9142502 | 1997 | Tissue-specific expression of murine Nkx3.1 in the male urogenital system. | Sciavolino PJ et al |
| 14648854 | 2003 | Roles of the Nkx3.1 homeobox gene in prostate organogenesis and carcinogenesis. | Shen MM et al |
| 16201967 | 2006 | Nkx3.1 binds and negatively regulates the transcriptional activity of Sp-family members in prostate-derived cells. | Simmons SO et al |
| 10871372 | 2000 | DNA-binding sequence of the human prostate-specific homeodomain protein NKX3.1. | Steadman DJ et al |
| 11002344 | 2000 | Nkx3.1, a murine homolog of Ddrosophila bagpipe, regulates epithelial ductal branching and proliferation of the prostate and palatine glands. | Tanaka M et al |
| 10415359 | 1999 | Expression of the Nkx3.1 homobox gene during pre and postnatal development. | Tanaka M et al |
| 9377551 | 1997 | Coding region of NKX3.1, a prostate-specific homeobox gene on 8p21, is not mutated in human prostate cancers. | Voeller HJ et al |
| 10688034 | 2000 | Expression profile of an androgen regulated prostate specific homeobox gene NKX3.1 in primary prostate cancer. | Xu LL et al |
| 18296735 | 2008 | Structural and functional analysis of domains mediating interaction between the bagpipe homologue, Nkx3.1 and serum response factor. | Zhang Y et al |
| 16397218 | 2006 | Germ-line mutation of NKX3.1 cosegregates with hereditary prostate cancer and alters the homeodomain structure and function. | Zheng SL et al |
Other Information
Locus ID:
NCBI: 4824
MIM: 602041
HGNC: 7838
Ensembl: ENSG00000167034
Variants:
dbSNP: 4824
ClinVar: 4824
TCGA: ENSG00000167034
COSMIC: NKX3-1
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000167034 | ENST00000380871 | Q99801 |
| ENSG00000167034 | ENST00000523261 | Q99801 |
Expression (GTEx)
Pathways
| Pathway | Source | External ID |
|---|---|---|
| Prostate cancer | KEGG | ko05215 |
| Pathways in cancer | KEGG | hsa05200 |
| Prostate cancer | KEGG | hsa05215 |
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38525603 | 2024 | NKX3.1 Expression in Non-Prostatic Tumors and Characterizing its Expression in Esophageal/Gastroesophageal Adenocarcinoma. | 0 |
| 38525603 | 2024 | NKX3.1 Expression in Non-Prostatic Tumors and Characterizing its Expression in Esophageal/Gastroesophageal Adenocarcinoma. | 0 |
| 36746884 | 2023 | NKX3.1 Expression and Molecular Characterization of Secretory Myoepithelial Carcinoma (SMCA): Advancing the Case for a Salivary Mucous Acinar Phenotype. | 1 |
| 37334663 | 2023 | Fibroblast-derived exosomal microRNA regulates NKX3-1 expression in androgen-sensitive, androgen receptor-dependent prostate cancer cells. | 1 |
| 37893500 | 2023 | Comprehensive Analysis of NKX3.2 in Liver Hepatocellular Carcinoma by Bigdata. | 0 |
| 36746884 | 2023 | NKX3.1 Expression and Molecular Characterization of Secretory Myoepithelial Carcinoma (SMCA): Advancing the Case for a Salivary Mucous Acinar Phenotype. | 1 |
| 37334663 | 2023 | Fibroblast-derived exosomal microRNA regulates NKX3-1 expression in androgen-sensitive, androgen receptor-dependent prostate cancer cells. | 1 |
| 37893500 | 2023 | Comprehensive Analysis of NKX3.2 in Liver Hepatocellular Carcinoma by Bigdata. | 0 |
| 34996754 | 2022 | Gene of the month: NKX3.1. | 3 |
| 35968571 | 2022 | Genome-Wide 3'-UTR Single Nucleotide Polymorphism Association Study Identifies Significant Prostate Cancer Risk-Associated Functional Loci at 8p21.2 in Chinese Population. | 5 |
| 34996754 | 2022 | Gene of the month: NKX3.1. | 3 |
| 35968571 | 2022 | Genome-Wide 3'-UTR Single Nucleotide Polymorphism Association Study Identifies Significant Prostate Cancer Risk-Associated Functional Loci at 8p21.2 in Chinese Population. | 5 |
| 33302222 | 2021 | NKX3.1 a useful marker for mesenchymal chondrosarcoma: An immunohistochemical study. | 11 |
| 33640159 | 2021 | NKX3.1 immunohistochemistry is highly specific for the diagnosis of mesenchymal chondrosarcomas: experience in the Australian population. | 2 |
| 34686726 | 2021 | Nkx3-1 and Fech genes might be switch genes involved in pituitary non-functioning adenoma invasiveness. | 4 |
Citation
Liang-Nian Song ; Edward P Gelmann
NKX3-1 (NK3 homeobox 1)
Atlas Genet Cytogenet Oncol Haematol. 2009-03-01
Online version: http://atlasgeneticsoncology.org/gene/41541/nkx3-1
