NR3C2 (nuclear receptor subfamily 3, group C, member 2)
2008-12-01 Francesco Di Fabio , Bruce Gottlieb , Lenore K Beitel , Mark Trifiro AffiliationIdentity
HGNC
LOCATION
4q31.23
LOCUSID
ALIAS
MCR,MLR,MR,NR3C2VIT
FUSION GENES
DNA/RNA
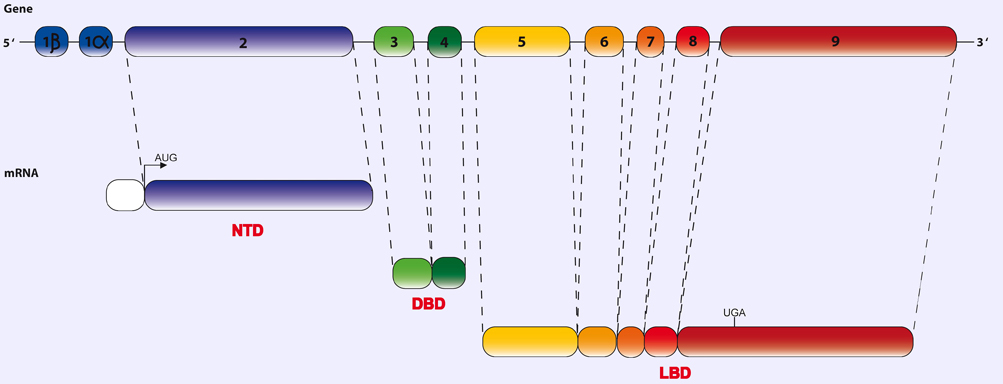
Schematic representation of human mineralocorticoid receptor (MR/NR3C2) structure. The human MR/NR3C2 gene is composed of 10 exons. The first 2 exons (1alpha and 1beta) are untranslated. However, alternative transcription of these exons can give rise to 2 RNA isoforms. The subsequent eight exons encode for the entire MR protein. MR/NR3C2 has three major functional domains: a modulating N-terminal domain (NTD), a central DNA-binding domain (DBD) and a C-terminal steroid ligand-binding domain (LBD). Exon 2 encodes for most of the NTD, exons 3 and 4 encode for each of the two zinc fingers of the DBD and the remaining five exons encode the LBD. AUG: start codon; UGA: stop codon.
Description
10 exons; 363,729 bases DNA.
Transcription
Multiple (at least four) mRNA isoforms are generated by alternative transcription or slicing events. Multiple mRNA isoforms are translated into various protein variants.
Proteins
Description
984 amino acids. Like all member of the nuclear receptor superfamily, the MR has 3 major domains: an N-terminal domain (NTD), a DNA-binding domain (DBD) and a ligand-binding domain (LBD). The NTD (602 amino acids) is composed of several functional domains (AF-1a, a central domain and AF-1b) that recruit various coregulators responsible for selectively modulating the transcriptional activity of MR. The DBD (66 amino acids) recognizes specific target DNA sequences or hormone response elements. The LBD (251 amino acids) is a multifunctional domain allowing selective hormone binding. The LBD includes the ligand-dependent AF-2 functional domain, which undergoes a rearrangement upon ligand binding.
Expression
Ubiquitous. MR expression has been reported in the kidney, colon, breast, salivary glands, sweat glands, liver, cardiomyocytes, endothelial cells, lung, central nervous system (hippocampus, hypothalamus), ocular tissues (retina, iris-ciliary body), adipose tissues (white and brown), inner ear, skin, placenta, uterus, ovaries and testis.
Localisation
Cytoplasm, nucleus, endoplasmic reticulum membrane, peripheral membrane: cytoplasmic and nuclear in the absence of ligand; nuclear after ligand-binding. When bound to 11 beta-hydroxysteroid dehydrogenase type 2 (11 beta-HSD2), it is found associated with the endoplasmic reticulum membrane.
Function
Transcription factor activity, receptor activity, protein binding, sequence-specific DNA binding, steroid binding, steroid hormone receptor activity, metal ion binding, zinc ion binding.
Homology
Member of the nuclear hormone receptor family, NR3 subfamily.
Mutations
Note
Loss-of-function mutations. About 50 distinct mutations in the human MR are known to be responsible for pseudohypoaldosteronism type 1 (PHA1). This is an autosomal genetic disorder caused by loss-of-function mutations. The mutations reported are missense, nonsense, frameshift, splice site mutations and deletions. PHA1 is characterized by salt wasting due to loss-of-function of the MR in the target organs. There are 2 forms of PHA1: the autosomal dominant form, which may be severe at birth, but symptoms remit with age, and the recessive form, which manifests with more severe symptoms persisting into adulthood.
Gain-of-function mutations. Only one mutation, to date, is known to result in a gain-of-function of MR. The single mutation S810L in the LBD leads to a constant MR activation. The inheritance is autosomal dominant. The disease is characterized by the onset of severe hypertension before the age of 20, with severe exacerbation in pregnancy.
Gain-of-function mutations. Only one mutation, to date, is known to result in a gain-of-function of MR. The single mutation S810L in the LBD leads to a constant MR activation. The inheritance is autosomal dominant. The disease is characterized by the onset of severe hypertension before the age of 20, with severe exacerbation in pregnancy.
Implicated in
Entity name
Leukaemia
Note
Some of the leukaemic cell lines express both the MR and the amiloride-sensitive sodium channel.
Oncogenesis
Some leukemic cell lines respond to mineralocorticoid steroids by an altered growth response that may possibly be linked to apoptosis. The altered sodium flux due to the induction of the amiloride-sensitive sodium channel by mineralocorticoids may cause uncontrolled cell proliferation in cancer.
Entity name
Colorectal carcinoma
Note
Reduction of MR mRNA expression is an early event in human sporadic carcinoma progression.
Oncogenesis
A significant inverse association between MR and vascular endothelial growth factor receptor-2 (VEGFR-2) expression at the mRNA level has been detected in human colorectal carcinoma, suggesting a potential tumor-suppressive function for MR. The degree of MR underexpression may have a role in the pro-angiogenic switch of colorectal carcinoma.
Entity name
Cervical carcinoma
Note
In a study of gene expression profiles in squamous cell cervical carcinoma, NR3C2 was observed to be significantly downregulated, which would suggest that it should be considered as a new putative cervical cancer-related gene.
Entity name
Renal cell neoplasms
Note
Using immunohistochemistry, expression of both MR and its related enzyme 11 beta-HSD2 was detected in chromophobe renal cell carcinoma and in oncocytomas. No staining was detected in clear cell renal cell carcinomas. MR and 11 beta-HSD2 may be considered specific immunohistochemical markers of the distal nephron and its related neoplasms (chromophobe renal cell carcinoma and oncocytoma).
Entity name
Lung carcinoma
Note
It has been reported that MR and 11 beta-HSD2 seem to be expressed only in adenocarcinomas or in the adenocarcinomatous component of adenosquamous carcinomas. No MR or 11 beta-HSD2 expression was detected in squamous cell carcinomas. MR and 11 beta-HSD2 immunoreactivity have been significantly correlated with the grade of differentiation of adenocarcinomas. Patterns of MR and 11 beta-HSD2 expression as detected with immunohistochemistry may reflect the cellular origin and differentiation status of primary human lung carcinoma, and may serve as a marker of differentiation.
Entity name
Breast carcinoma
Note
Expression of MR, as detected with immunohistochemistry, appears to be related to ductal differentiation of breast carcinomas.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 3037703 | 1987 | Cloning of human mineralocorticoid receptor complementary DNA: structural and functional kinship with the glucocorticoid receptor. | Arriza JL et al |
| 15063830 | 2004 | Aldosterone signaling modifies capillary formation by human bone marrow endothelial cells. | Chen W et al |
| 17504382 | 2007 | Gene expression profiles in squamous cell cervical carcinoma using array-based comparative genomic hybridization analysis. | Choi YW et al |
| 17703341 | 2007 | Underexpression of mineralocorticoid receptor in colorectal carcinomas and association with VEGFR-2 overexpression. | Di Fabio F et al |
| 17293689 | 2007 | Aldosterone, mineralocorticoid receptors, and vascular inflammation. | Fiebeler A et al |
| 18249428 | 2008 | Nongenotropic aldosterone effects and the EGFR: interaction and biological relevance. | Grossmann C et al |
| 12939263 | 2003 | Aldosterone stimulates epidermal growth factor receptor expression. | Krug AW et al |
| 16847146 | 2006 | Aldosterone impairs bone marrow-derived progenitor cell formation. | Marumo T et al |
| 10865975 | 2000 | Demonstration of the mineralocorticoid hormone receptor and action in human leukemic cell lines. | Mirshahi M et al |
| 17285301 | 2007 | Is the vascular endothelium under the control of aldosterone? Facts and hypothesis. | Oberleithner H et al |
| 12943727 | 2003 | Dissecting mineralocorticoid receptor structure and function. | Rogerson FM et al |
| 9216657 | 1997 | Localization of mineralocorticoid receptor and 11 beta-hydroxysteroid dehydrogenase type II in human breast and its disorders. | Sasano H et al |
| 10769675 | 2000 | Expression of 11 beta-hydroxysteroid dehydrogenase type 2 and mineralocorticoid receptor in primary lung carcinomas. | Suzuki S et al |
| 18174920 | 2007 | The mineralocorticoid receptor: insights into its molecular and (patho)physiological biology. | Viengchareun S et al |
| 18408592 | 2008 | Mineralocorticoid receptor and 11beta-hydroxysteroid dehydrogenase type II expression in renal cell neoplasms: a tissue microarray and quantitative RT-PCR study. | Yakirevich E et al |
| 7673127 | 1995 | Human mineralocorticoid receptor genomic structure and identification of expressed isoforms. | Zennaro MC et al |
Other Information
Locus ID:
NCBI: 4306
MIM: 600983
HGNC: 7979
Ensembl: ENSG00000151623
Variants:
dbSNP: 4306
ClinVar: 4306
TCGA: ENSG00000151623
COSMIC: NR3C2
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
PharmGKB
| Entity ID | Name | Type | Evidence | Association | PK | PD | PMIDs |
|---|---|---|---|---|---|---|---|
| PA444552 | Hypertension | Disease | ClinicalAnnotation | associated | PD | 24059494 | |
| PA449456 | enalapril | Chemical | ClinicalAnnotation | associated | PD | 24059494 |
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38127122 | 2024 | Mineralocorticoid receptor overactivation: targeting systemic impact with non-steroidal mineralocorticoid receptor antagonists. | 1 |
| 38255827 | 2024 | First Evidence of Mineralocorticoid Receptor Gene and Protein Expression in Rat and Human Thyroid Tissues and Cell Cultures. | 0 |
| 38127122 | 2024 | Mineralocorticoid receptor overactivation: targeting systemic impact with non-steroidal mineralocorticoid receptor antagonists. | 1 |
| 38255827 | 2024 | First Evidence of Mineralocorticoid Receptor Gene and Protein Expression in Rat and Human Thyroid Tissues and Cell Cultures. | 0 |
| 36440858 | 2023 | Emerging vascular cell-specific roles for mineralocorticoid receptor: implications for understanding sex differences in cardiovascular disease. | 4 |
| 36538242 | 2023 | CircSTK39 suppresses the proliferation and invasion of bladder cancer by regulating the miR-135a-5p/NR3C2-mediated epithelial-mesenchymal transition signaling pathway. | 2 |
| 36734065 | 2023 | Associations of ARMS2 and NR3C2 genes polymorphisms with central serous chorioretinopathy in a Greek population. | 0 |
| 36808340 | 2023 | The mineralocorticoid receptor gene (NR3C2) is linked to and associated with polycystic ovarian syndrome in Italian families. | 0 |
| 36919662 | 2023 | NR3C2 mediates oxidised low-density lipoprotein-induced human coronary endothelial cells dysfunction via modulation of NLRP3 inflammasome activation. | 3 |
| 36950803 | 2023 | NR3C2 inhibits the proliferation of colorectal cancer via regulating glucose metabolism and phosphorylating AMPK. | 5 |
| 37130784 | 2023 | Down-Regulation of the Mineralocorticoid Receptor (MR) and Up-Regulation of Hydroxysteroid 11-Beta Dehydrogenase Type 2 (HSD11B2) Isoenzyme in Critically Ill Patients. | 0 |
| 37175424 | 2023 | Mineralocorticoid Receptor Antagonists for Preventing Chronic Kidney Disease Progression: Current Evidence and Future Challenges. | 2 |
| 38054065 | 2023 | Effects of environmental and genetic interactions on job burnout in coal miners: interactions between occupational stress, coping styles, and NR3C2 gene polymorphisms. | 0 |
| 36440858 | 2023 | Emerging vascular cell-specific roles for mineralocorticoid receptor: implications for understanding sex differences in cardiovascular disease. | 4 |
| 36538242 | 2023 | CircSTK39 suppresses the proliferation and invasion of bladder cancer by regulating the miR-135a-5p/NR3C2-mediated epithelial-mesenchymal transition signaling pathway. | 2 |
Citation
Francesco Di Fabio ; Bruce Gottlieb ; Lenore K Beitel ; Mark Trifiro
NR3C2 (nuclear receptor subfamily 3, group C, member 2)
Atlas Genet Cytogenet Oncol Haematol. 2008-12-01
Online version: http://atlasgeneticsoncology.org/gene/44262/nr3c2
