PEG3 (paternally expressed 3)
2010-08-01 Yinhua Yu , Weiwei Feng , Zhen Lu , Robert C Bast Jr AffiliationThe University of Texas, MD Anderson Cancer Center, 1515 Holcombe Blvd, Box 354, Houston, TX 77030, USA
DNA/RNA
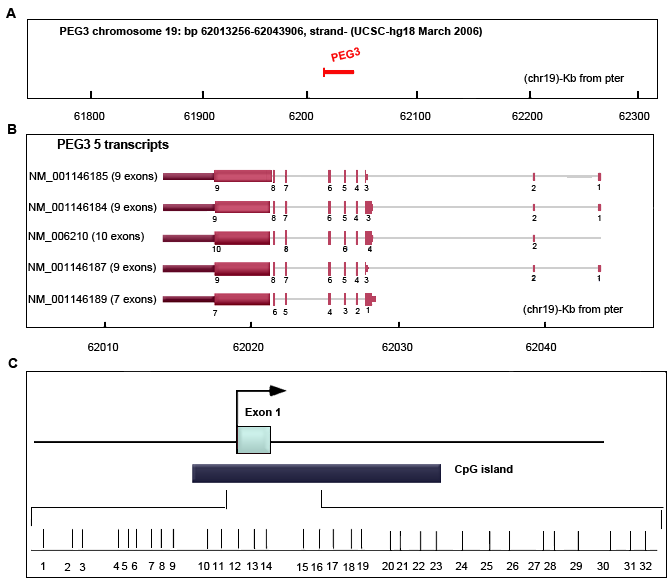
Figure A. PEG3 DNA located on chromosome 19, starts at 62013257 and ends at 62043906 bp.
Figure B. The PEG3 gene contains 7-10 exons with 5 different transcripts.
Figure C. PEG3 promoter and first exon. The transcription start site and direction are indicated with an arrow. The horizontal bars are 32 CpG sites. The dark blue box is a CpG island surrounded the first exon (Figure modified from Feng et al., 2008).
Description
Human PEG3 was originally identified as an ortholog of murine Peg3 that is the first imprinted gene detected in mouse chromosome 7. Human PEG3 is located ~2 Mb proximal of the 19q telomere, which is rich in Krüppel-type ZNF genes (Kim et al., 1997). Bisulfite sequencing across PEG3 revealed that all CpG dinucleotides examined were differentially methylated in human fetal brain, kidney, liver, and pancreas. PEG3 is monoallelically expressed during fetal development, it is a maternal-imprinted gene (Murphy et al., 2001).
Transcription
The PEG3 gene contains 7-10 exons with 5 different transcripts.
Pseudogene
There are no known pseudogenes, but an antisense transcript gene (APEG3), which is transcribed in the opposite direction of PEG3, has been identified from human. It is confirmed the presence of APEG3 in total RNAs from brain thalamus and testis. The functional significance of APEG3 is not known. The paternal allele-specific expression of anti-sense Peg3 (imprinting) was detected in mouse brain, but not in human yet (Choo et al., 2002).
Proteins
Description
PEG3 is maternal-imprinted and paternally expressed. The imprinted status of PEG3 throughout development and adult life. PEG3 may be a susceptibility locus for cancer and for neurobehavioral deficits (Murphy et al., 2001).
Expression
PEG3 is expressed at higher levels in ovary and placenta (Kim et al., 1997; Hiby et al., 2001). It was also strongly expressed in the adult brain (Kohda et al., 2001).
Localisation
High levels of PEG3 have localized to the layer of villous cytotrophoblast cells in the human placenta, PEG3 expression is also detected in the ovary stroma (Hiby et al., 2001).
Function
PEG3 is an imprinted gene expressed exclusively from the paternal allele. The precise function of PEG3 is not clear, but recent evidence suggests that it plays an important role in the p53/c-myc-mediated apoptosis pathway.
PEG3 is a maternally imprinted tumor suppressor gene that is downregulated in gliomas, ovarian cancers, breast cancers and other gynecologic cancers (Kohda et al., 2001; Dowdy et al., 2005; Feng et al., 2008).
There are several studies revealed murine Peg3 acts as an intermediary between p53 and Bax in a cell death pathway activated by DNA damage in primary mouse cortical neurons, inhibiting Peg3 activity blocks p53-induced apoptosis (Johnson et al., 2002). Pw1/Peg3 interacts with a p53-inducible gene product Siah1a. Coexpression of Pw1/Peg3 with Siah1a induces apoptosis independently of p53. Inhibiting Pw1/Peg3 activity blocks p53-induced apoptosis (Relaix et al., 2000). Since human PEG3 is highly conserved with murine Peg3, PEG3 may have same function, Jiang et al. (2010) demonstrated that enforced overexpression of PEG3 mRNA during zebrafish embryogenesis decreased beta-catenin protein expression and inhibited Wnt-dependent tail development. Peg3/Pw1 also inhibited Wnt signaling in human cells by binding to beta-catenin and promoting its degradation via a p53/Siah1-dependent, GSK3beta-independent proteasomal pathway. Hypermethylation of the PEG3 promoter in primary human gliomas led to a loss of imprinting and decreased PEG3 mRNA expression that correlated with increasing tumor grade (Jiang et al., 2010).
The transcription factor YY1 can bind to the first intron of human PEG3, and specifically to the paternal allele of the gene. YY1-binding sites are methylated only on the maternal chromosome. YY1-binding region may function as a methylation-sensitive insulator that could influence the imprinted expression of PEG3 (Kim et al., 2003).
PEG3 is a maternally imprinted tumor suppressor gene that is downregulated in gliomas, ovarian cancers, breast cancers and other gynecologic cancers (Kohda et al., 2001; Dowdy et al., 2005; Feng et al., 2008).
There are several studies revealed murine Peg3 acts as an intermediary between p53 and Bax in a cell death pathway activated by DNA damage in primary mouse cortical neurons, inhibiting Peg3 activity blocks p53-induced apoptosis (Johnson et al., 2002). Pw1/Peg3 interacts with a p53-inducible gene product Siah1a. Coexpression of Pw1/Peg3 with Siah1a induces apoptosis independently of p53. Inhibiting Pw1/Peg3 activity blocks p53-induced apoptosis (Relaix et al., 2000). Since human PEG3 is highly conserved with murine Peg3, PEG3 may have same function, Jiang et al. (2010) demonstrated that enforced overexpression of PEG3 mRNA during zebrafish embryogenesis decreased beta-catenin protein expression and inhibited Wnt-dependent tail development. Peg3/Pw1 also inhibited Wnt signaling in human cells by binding to beta-catenin and promoting its degradation via a p53/Siah1-dependent, GSK3beta-independent proteasomal pathway. Hypermethylation of the PEG3 promoter in primary human gliomas led to a loss of imprinting and decreased PEG3 mRNA expression that correlated with increasing tumor grade (Jiang et al., 2010).
The transcription factor YY1 can bind to the first intron of human PEG3, and specifically to the paternal allele of the gene. YY1-binding sites are methylated only on the maternal chromosome. YY1-binding region may function as a methylation-sensitive insulator that could influence the imprinted expression of PEG3 (Kim et al., 2003).
Homology
Human PEG3 is known to have orthologs in mice and cow. Human PEG3 gene sequences revealed a high level of conservation with Murine Peg3 (83% similarity), but one of the two proline-rich repeats is absent from the human PEG3 (Kim et al., 1997).
Mutations
Note
Mutations have not been detected.
Implicated in
Entity name
Glioma, glioblastoma
Note
A significant decrease in PEG3 expression was more commonly observed in glioma cell lines as compared with that in primary cultures of astrocytes. Transfection of PEG3 cDNA in a glioma cell line resulted in a loss of tumorigenicity in nude mice (Kohda et al., 2001). The epigenetic silencing of PEG3 expression in glioma cell lines depends on aberrant DNA methylation of an exonic CpG island. Treatment of glioma cell lines with the DNA demethylating agent 5-aza-2-deoxycytidine reversed the silencing of PEG3 biallelically (Maegawa et al., 2001). Re-expression of PEG3 in glioma cells suppresses their proliferation. Hypermethylation of the PEG3 promoter in primary human gliomas led to a loss of imprinting and decreased PEG3 mRNA expression that correlated with tumor grade (Jiang et al., 2010). The lower gene expression was confirmed statistically in glioblastoma (Otsuka et al., 2009).
Entity name
Endometrial cancer, cervical cancer, choriocarcinomas
Note
PEG3 is silenced in all endometrial and cervical cancer cell lines studied. In contrast, loss of maternal imprinting and relatively high PEG3 expression levels were detected in all four choriocarcinomas cell lines studied (Dowdy et al., 2005).
Entity name
Breast cancer, ovarian cancer
Note
Five of the eight ovarian cancer cell lines were found to be PEG3 negative, the remaining three express low levels of PEG3 mRNA (Dowdy et al., 2005). PEG3 was down-regulated in 75% of ovarian cancers. PEG3 was hypermethylated in 11 of 42 ovarian cancers (26%), and PEG3 expression was down-regulated in 10 of those 11 cancers. LOH was detected in 5 of 25 informative cases for PEG3 (20%). Re-expression of PEG3 markedly inhibited ovarian cancer growth. PEG3 expression could be restored by treatment with 5-aza-2-deoxycytidine and trichostatin A (Feng et al., 2008).
Entity name
Hydatidiform moles
Note
PEG3 is not mutated in women with familial recurrent hydatidiform moles, there is allele-specific methylation of the CpG island and expression from the paternal allele in two independent informative pedigrees (Van den Veyver et al., 2001).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 18166281 | 2008 | Imprinting of an evolutionarily conserved antisense transcript gene APeg3. | Choo JH et al |
| 16023706 | 2005 | Biallelic methylation and silencing of paternally expressed gene 3 (PEG3) in gynecologic cancer cell lines. | Dowdy SC et al |
| 18286529 | 2008 | Imprinted tumor suppressor genes ARHI and PEG3 are the most frequently down-regulated in human ovarian cancers by loss of heterozygosity and promoter methylation. | Feng W et al |
| 11331620 | 2001 | Paternal monoallelic expression of PEG3 in the human placenta. | Hiby SE et al |
| 20064927 | 2010 | The imprinted gene PEG3 inhibits Wnt signaling and regulates glioma growth. | Jiang X et al |
| 11943780 | 2002 | Peg3/Pw1 is a mediator between p53 and Bax in DNA damage-induced neuronal death. | Johnson MD et al |
| 9149948 | 1997 | The human homolog of a mouse-imprinted gene, Peg3, maps to a zinc finger gene-rich region of human chromosome 19q13.4. | Kim J et al |
| 12554678 | 2003 | Methylation-sensitive binding of transcription factor YY1 to an insulator sequence within the paternally expressed imprinted gene, Peg3. | Kim J et al |
| 11260267 | 2001 | Tumour suppressor activity of human imprinted gene PEG3 in a glioma cell line. | Kohda T et al |
| 11398192 | 2001 | Epigenetic silencing of PEG3 gene expression in human glioma cell lines. | Maegawa S et al |
| 11161803 | 2001 | Imprinting of PEG3, the human homologue of a mouse gene involved in nurturing behavior. | Murphy SK et al |
| 19367087 | 2009 | Aberrant promoter methylation and expression of the imprinted PEG3 gene in glioma. | Otsuka S et al |
| 10681424 | 2000 | Pw1/Peg3 is a potential cell death mediator and cooperates with Siah1a in p53-mediated apoptosis. | Relaix F et al |
| 11677152 | 2001 | The human homologue (PEG3) of the mouse paternally expressed gene 3 (Peg3) is maternally imprinted but not mutated in women with familial recurrent hydatidiform molar pregnancies. | Van den Veyver IB et al |
Other Information
Locus ID:
NCBI: 5178
MIM: 601483
HGNC: 8826
Ensembl: ENSG00000198300
Variants:
dbSNP: 5178
ClinVar: 5178
TCGA: ENSG00000198300
COSMIC: PEG3
RNA/Proteins
Expression (GTEx)
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37001595 | 2023 | Epigenetic reactivation of PEG3 by EZH2 inhibitors suppresses renal clear cell carcinoma progress. | 2 |
| 37001595 | 2023 | Epigenetic reactivation of PEG3 by EZH2 inhibitors suppresses renal clear cell carcinoma progress. | 2 |
| 34192708 | 2021 | miR-155-5p Promotes Cell Proliferation and Migration of Clear Cell Renal Cell Carcinoma by Targeting PEG3. | 2 |
| 34192708 | 2021 | miR-155-5p Promotes Cell Proliferation and Migration of Clear Cell Renal Cell Carcinoma by Targeting PEG3. | 2 |
| 32729618 | 2020 | PEG3 mutation is associated with elevated tumor mutation burden and poor prognosis in breast cancer. | 8 |
| 33023834 | 2020 | HPV-CCDC106 integration alters local chromosome architecture and hijacks an enhancer by three-dimensional genome structure remodeling in cervical cancer. | 33 |
| 32729618 | 2020 | PEG3 mutation is associated with elevated tumor mutation burden and poor prognosis in breast cancer. | 8 |
| 33023834 | 2020 | HPV-CCDC106 integration alters local chromosome architecture and hijacks an enhancer by three-dimensional genome structure remodeling in cervical cancer. | 33 |
| 29080840 | 2019 | Decorin is a devouring proteoglycan: Remodeling of intracellular catabolism via autophagy and mitophagy. | 39 |
| 31420931 | 2019 | Long noncoding RNA ZNF667-AS1 reduces tumor invasion and metastasis in cervical cancer by counteracting microRNA-93-3p-dependent PEG3 downregulation. | 29 |
| 29080840 | 2019 | Decorin is a devouring proteoglycan: Remodeling of intracellular catabolism via autophagy and mitophagy. | 39 |
| 31420931 | 2019 | Long noncoding RNA ZNF667-AS1 reduces tumor invasion and metastasis in cervical cancer by counteracting microRNA-93-3p-dependent PEG3 downregulation. | 29 |
| 28174297 | 2017 | Decorin-inducible Peg3 Evokes Beclin 1-mediated Autophagy and Thrombospondin 1-mediated Angiostasis. | 31 |
| 28798237 | 2017 | Decorin-evoked paternally expressed gene 3 (PEG3) is an upstream regulator of the transcription factor EB (TFEB) in endothelial cell autophagy. | 30 |
| 28852087 | 2017 | Reduced expression of Paternally Expressed Gene-3 enhances somatic cell reprogramming through mitochondrial activity perturbation. | 7 |
Citation
Yinhua Yu ; Weiwei Feng ; Zhen Lu ; Robert C Bast Jr
PEG3 (paternally expressed 3)
Atlas Genet Cytogenet Oncol Haematol. 2010-08-01
Online version: http://atlasgeneticsoncology.org/gene/41690/peg3
