PLCD1 (phospholipase C, delta 1)
2011-01-01 Xiaotong Hu AffiliationBiomedical Research Center, Sir Run Run Shaw Hospital, Zhejiang University, Hangzhou, China
Identity
HGNC
LOCATION
3p22.2
IMAGE

LEGEND
PLCD1 starts at 38048987 bp and ends at 38071253 bp from pter on Chr3p22-p21.3 and located between VILL and DLEC1 gene.
LOCUSID
ALIAS
NDNC3,PLC-III
FUSION GENES
DNA/RNA
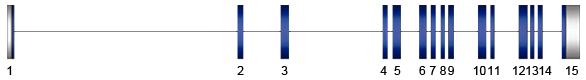

Closed and opened boxes represent coding and non-coding sequences of PLCD1 gene, respectively.
Description
The PLCD1 gene is a functioning gene and contains 15 exons and spans 22.17 kb.
Transcription
The variant 1 (NM_001130964) encodes the longer isoform 1 (NP_001124436). The variant 2 (NM_006225.3) contains an alternate 5 terminal exon compared to transcript variant 1, and initiates translation from an in-frame upstream AUG, resulting in a shorter isoform 2 (NP_006216) with a different N-terminus compared to isoform 1.
Proteins
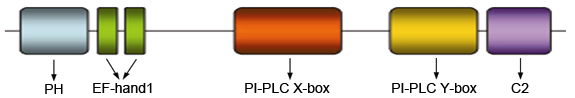
Protein domain organization of the mammalian PLCD1.
Description
The deduced 777-amino acid isoform 1 (NM_001130964) and 756-amino acid isoform 2 (NP_006216) shares 95% sequence homology with the rat protein. They contain a N-terminal PH domain, 2 EF-hand1 domains, PI-PLC X-box, PI-PLC Y-box and C2 region.
Expression
Expressed high or medium in CNS (brain), hematopoietic (blood), liver and pancreas, digestive (GI-tract), respiratory (lung), male and female tissues, placenta, urinary tract (kidney) skin and soft tissues but no expression in cardio vascular and endocrine tissues.
Localisation
Intracellular: cytoplasm and nucleus.
Function
Catalyzing the hydrolysis of phosphatidylinositol 4,5 biphosphate to generate diacylglycerol and inositol 1,4,5 triphosphate. Mediating a wide variety of cellular stimuli. Shuttling between the nucleus and the cytoplasm, and nuclear import is mediated by its Ca2+-dependent interaction with importin beta 1. Playing an important suppressive role in the development and progression of cancers such as esophageal squamous cell carcinoma (ESCC) and gastric cancer (GC).
Homology
The PLCD1 gene is conserved in dog, cow, mouse, rat, chicken, zebrafish, and A. thaliana.
Mutations

The mutation location is highlighted in red and this mutation occured in 17% (1/6) skin samples.
AA Mutation: p.E226K (Substitution - Missense).
CDS Mutation: c.676G>A (Substitution).
Implicated in
Entity name
Esophageal squamous cell carcinoma
Prognosis
Firstly, four commonly deleted regions (CDRs) at 3p26.3, 3p22, 3p21.3 and 3p14.2 were identified. Absent and down-regulated expression of several candidate TSGs, including CHL1, PCAF, RBMS3, PLCD1 and CACNA2D3, were detected in primary ESCC tumors and ESCC cell lines. These results provided evidence that minimal deleted regions at 3p26.3, 3p22, 3p21.3 and 3p14.2 containing potential TSGs may contribute to the pathogenesis of esophageal cancer.
Secondly, absent expression of PLC delta 1 was detected in 26 of 50 (52%) primary ESCCs and 4 of 9 (44.4%) ESCC cell lines, which was significantly associated with DNA copy number loss and promoter hypermethylation (P < 0.05). Functional studies showed that PLC delta 1 was able to suppress both in vitro and in vivo tumorigenic ability of ESCC cells, including foci formation, colony formation in soft agar, and tumor formation in nude mice. The tumor-suppressive mechanism of PLC delta 1 was associated with its role in the cell cycle arrest at the G(1)-S checkpoint by up-regulation of p21 and down-regulation of phosphorylated Akt (Ser(473)). In addition, down-regulation of PLC delta 1 protein was significantly correlated with ESCC metastasis (P = 0.014), which was associated with its function in increasing cell adhesion and inhibiting cell mobility. These results suggest that PLC delta 1 plays an important suppressive role in the development and progression of ESCC.
Secondly, absent expression of PLC delta 1 was detected in 26 of 50 (52%) primary ESCCs and 4 of 9 (44.4%) ESCC cell lines, which was significantly associated with DNA copy number loss and promoter hypermethylation (P < 0.05). Functional studies showed that PLC delta 1 was able to suppress both in vitro and in vivo tumorigenic ability of ESCC cells, including foci formation, colony formation in soft agar, and tumor formation in nude mice. The tumor-suppressive mechanism of PLC delta 1 was associated with its role in the cell cycle arrest at the G(1)-S checkpoint by up-regulation of p21 and down-regulation of phosphorylated Akt (Ser(473)). In addition, down-regulation of PLC delta 1 protein was significantly correlated with ESCC metastasis (P = 0.014), which was associated with its function in increasing cell adhesion and inhibiting cell mobility. These results suggest that PLC delta 1 plays an important suppressive role in the development and progression of ESCC.
Entity name
Breast carcinoma
Prognosis
PLCD1 are more highly expressed in the transformed cell lines compared to MCF-10A. To test whether PLCd1 or PLCd3 played any role in tumor cell proliferation or cell migration. RNAi mediated knockdown of PLCD1 reduced proliferation of the MDA-MB-231 cells. Morphological changes including cell rounding, and surface blebbing and nuclear fragmentation were observed. These changes were accompanied by reductions in cell migration activities. On the other hand, PLCD1 knockdown failed to cause comparable morphological changes in the normal MCF-10A line, but did reduce cell proliferation and migration. Taken together, these data are consistent with the idea that PLCD1 support the growth and migration of normal and neoplastic mammary epithelial cells in vitro.
However there is contrasted results published in another paper. Their results suggested that PLCD1 is a functional tumor suppressor inducing G(2)/M arrest and frequently methylated in breast cancer.
However there is contrasted results published in another paper. Their results suggested that PLCD1 is a functional tumor suppressor inducing G(2)/M arrest and frequently methylated in breast cancer.
Entity name
Gastric cancer
Prognosis
Located at the important tumor suppressor locus, 3p22, PLCD1 encodes an enzyme that mediates regulatory signaling of energy metabolism, calcium homeostasis and intracellular movements. We identified PLCD1 as a downregulated gene in aerodigestive carcinomas through expression profiling and epigenetic characterization. We found that PLCD1 was expressed in all normal adult tissues but low or silenced in 84% (16/19) gastric cancer cell lines, well correlated with its CpG island (CGI) methylation status. Methylation was further detected in 62% (61/98) gastric primary tumors, but none of normal gastric mucosa tissues. PLCD1 methylation was significantly correlated with tumor high stage. Detailed methylation analysis of 37 CpG sites at the PLCD1 CGI by bisulfite genomic sequencing confirmed its methylation. PLCD1 silencing could be reversed by pharmacological demethylation with 5-aza-2-deoxycytidine, indicating a direct epigenetic silencing. Ectopic expression of PLCD1 in silenced gastric tumor cells dramatically inhibited their clonogenicity and migration, possibly through downregulating MMP7 expression and hampering the reorganization of cytoskeleton through cofilin inactivation by phosphorylation. Thus, epigenetic inactivation of PLCD1 is common and tumor-specific in gastric cancer, and PLCD1 acts as a functional tumor suppressor involved in gastric carcinogenesis.
Entity name
Colon carcinomas
Prognosis
Decreased levels of the PLC delta 1 protein were seen in most colon carcinomas (12 of 13 paired samples) and PLC delta 1 protein was not detected in any of the carcinoma cell lines.
Entity name
Rat colon neoplasms
Prognosis
The expression of PLC-delta expression in rat colon neoplasms induced by methylazoxymethanol (MAM) acetate was examined. Large-bowel neoplasms were observed in five of 10 rats given MAM acetate 40 weeks after treatment. PLC-delta expression in the neoplasms was not detected by northern blot analysis, and a low level of expression was detected by immunoblot analysis, although PLC-delta expression was apparent in the non-neoplastic colon mucosae of MAM acetate-treated rats as well as in the colon mucosae of control rats.
Entity name
Insulinoma
Note
Insulinoma MIN6 cells.
Prognosis
To study the effects of enhanced phosphoinositide hydrolysis on insulin secretion, phosphoinositide-specific phospholipase Cbeta1 (PLCbeta1) or PLCdelta1 was overexpressed in insulinoma MIN6 cells via adenoviral vectors. Inositol phosphate production stimulated by KCl or glucose in both PLCbeta1- and PLCdelta1-overexpressing cells were greater than that in control cells, reduced phosphatidylinositol-4,5-bisphosphate levels were observed in these cells stimulated by NaF or KCl. These data suggest that excessive phosphoinositide hydrolysis inhibits secretagogue-induced insulin release in MIN6 cells.
Entity name
Pheochromocytoma
Note
Pheochromocytoma PC12 cells.
Prognosis
PLCD1 is recruited from the cytoplasm to lipid rafts after CCH-induced Ca2+ mobilization following the activation of PLCbeta by GPCR and PLCD1 is activated only in lipid rafts by localized capacitative entry of extracellular Ca2+. PLCD1, p122RhoGAP and RhoA in combination could constitute a unique functional unit for the regulation of both phosphoinositide/Ca2+ signaling and the actin cytoskeleton in the periphery of specified membrane domains. This would provide further insights into the molecular mechanisms of cancer development.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 18006814 | 2007 | Characterization of a novel tumor-suppressor gene PLC delta 1 at 3p22 in esophageal squamous cell carcinoma. | Fu L et al |
| 19448674 | 2009 | Phospholipase C delta 1 is a novel 3p22.3 tumor suppressor involved in cytoskeleton organization, with its epigenetic silencing correlated with high-stage gastric cancer. | Hu XT et al |
| 9920735 | 1999 | Enhanced phosphoinositide hydrolysis via overexpression of phospholipase C beta1 or delta1 inhibits stimulus-induced insulin release in insulinoma MIN6 cells. | Ishihara H et al |
| 9345909 | 1997 | Genomic structure of the human PLCD1 (phospholipase C delta 1) locus on 3p22-->p21.3. | Ishikawa S et al |
| 7893368 | 1995 | Expression of phospholipases gamma 1, beta 1, and delta 1 in primary human colon carcinomas and colon carcinoma cell lines. | Nomoto K et al |
| 18508313 | 2008 | Single-nucleotide polymorphism-mass array reveals commonly deleted regions at 3p22 and 3p14.2 associate with poor clinical outcome in esophageal squamous cell carcinoma. | Qin YR et al |
| 19344632 | 2009 | Expression and function of phospholipase C in breast carcinoma. | Rebecchi MJ et al |
| 20657189 | 2010 | PLCD1 is a functional tumor suppressor inducing G(2)/M arrest and frequently methylated in breast cancer. | Xiang T et al |
| 18157946 | 2008 | Recruitment and activation of phospholipase C (PLC)-delta1 in lipid rafts by muscarinic stimulation of PC12 cells: contribution of p122RhoGAP/DLC1, a tumor-suppressing PLCdelta1 binding protein. | Yamaga M et al |
| 7528022 | 1994 | Reduced expression of phospholipase C-delta, a signal-transducing enzyme, in rat colon neoplasms induced by methylazoxymethanol acetate. | Yoshimi N et al |
Other Information
Locus ID:
NCBI: 5333
MIM: 602142
HGNC: 9060
Ensembl: ENSG00000187091
Variants:
dbSNP: 5333
ClinVar: 5333
TCGA: ENSG00000187091
COSMIC: PLCD1
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000187091 | ENST00000334661 | P51178 |
| ENSG00000187091 | ENST00000334661 | A0A384MR47 |
| ENSG00000187091 | ENST00000463876 | P51178 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
PharmGKB
| Entity ID | Name | Type | Evidence | Association | PK | PD | PMIDs |
|---|---|---|---|---|---|---|---|
| PA26097 | CASR | Gene | Pathway | associated | |||
| PA33759 | PRKCA | Gene | Pathway | associated | |||
| PA33761 | PRKCB | Gene | Pathway | associated | |||
| PA33766 | PRKCG | Gene | Pathway | associated | |||
| PA33767 | PRKCH | Gene | Pathway | associated | |||
| PA33768 | PRKCI | Gene | Pathway | associated | |||
| PA33771 | PRKD1 | Gene | Pathway | associated | |||
| PA33775 | PRKCZ | Gene | Pathway | associated |
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36794559 | 2023 | One recurrent heterozygous mutation of the PLCD1 gene in a Chinese family with hereditary leukonychia: A case report and genotype-phenotype correlation analysis. | 1 |
| 36849889 | 2023 | Phospholipase C delta 1 inhibits WNT/β-catenin and EGFR-FAK-ERK signaling and is disrupted by promoter CpG methylation in renal cell carcinoma. | 1 |
| 36987828 | 2023 | Identification of a Novel PLCD1 Variant in a Danish Family with Hereditary Leukonychia. | 0 |
| 37228901 | 2023 | lncRNA MIR503HG Targets miR-191-5p/PLCD1 Axis and Negatively Modulates Apoptosis, Extracellular Matrix Disruption, and Inflammation in Abdominal Aortic Aneurysm. | 0 |
| 37460940 | 2023 | MicroRNA-191 regulates oral squamous cell carcinoma cells growth by targeting PLCD1 via the Wnt/β-catenin signaling pathway. | 1 |
| 36794559 | 2023 | One recurrent heterozygous mutation of the PLCD1 gene in a Chinese family with hereditary leukonychia: A case report and genotype-phenotype correlation analysis. | 1 |
| 36849889 | 2023 | Phospholipase C delta 1 inhibits WNT/β-catenin and EGFR-FAK-ERK signaling and is disrupted by promoter CpG methylation in renal cell carcinoma. | 1 |
| 36987828 | 2023 | Identification of a Novel PLCD1 Variant in a Danish Family with Hereditary Leukonychia. | 0 |
| 37228901 | 2023 | lncRNA MIR503HG Targets miR-191-5p/PLCD1 Axis and Negatively Modulates Apoptosis, Extracellular Matrix Disruption, and Inflammation in Abdominal Aortic Aneurysm. | 0 |
| 37460940 | 2023 | MicroRNA-191 regulates oral squamous cell carcinoma cells growth by targeting PLCD1 via the Wnt/β-catenin signaling pathway. | 1 |
| 32236884 | 2021 | PLCD1 Suppressed Cellular Proliferation, Invasion, and Migration via Inhibition of Wnt/β-Catenin Signaling Pathway in Esophageal Squamous Cell Carcinoma. | 8 |
| 32795530 | 2021 | Germline Mutation of PLCD1 Contributes to Human Multiple Pilomatricomas through Protein Kinase D/Extracellular Signal-Regulated Kinase1/2 Cascade and TRPV6. | 2 |
| 33236921 | 2021 | Phospholipase C controls chloride-dependent short-circuit current in human bronchial epithelial cells. | 1 |
| 33786625 | 2021 | Identification of two novel mutations in the PLCD1 gene in Chinese patients with hereditary leukonychia. | 1 |
| 34727930 | 2021 | The tumor suppressor activity of DLC1 requires the interaction of its START domain with Phosphatidylserine, PLCD1, and Caveolin-1. | 6 |
Citation
Xiaotong Hu
PLCD1 (phospholipase C, delta 1)
Atlas Genet Cytogenet Oncol Haematol. 2011-01-01
Online version: http://atlasgeneticsoncology.org/gene/43927/plcd1
