RANBP9 (RAN binding protein 9)
2012-03-01 Kwang-Hyun Baek , Bharathi Suresh AffiliationCHA Stem Cell Institute, CHA University, 606-16 Yeoksam 1-Dong, Gangnam-Gu, Seoul 135-081, Korea (KHB); Department of Biomedical Science, CHA University, CHA General Hospital, Seoul 135-081, Republic of Korea (BS)
Identity
HGNC
LOCATION
6p23
LOCUSID
ALIAS
BPM-L,BPM90,RANBPM,RanBP7
FUSION GENES
DNA/RNA
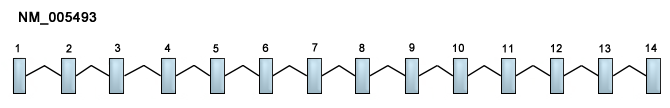
Description
Constists of 14 exons.
Transcription
The coding region of the gene starts from exon 1 to exon 14 (60th bps to 2249th bps). The length of the transcript is 2190 bps.
Pseudogene
No known pseudogenes.
Proteins
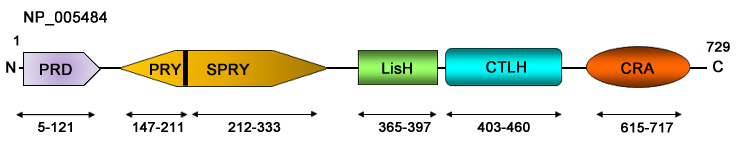
Description
RanBPM was originally identified as a centrosomal 55 kDa protein and is involved in microtubule nucleation at the centrosome (Nakamura et al., 1998). A later work by the same group revealed that the full-length RanBPM encodes for 729 amino acids and its molecular weight is 90 kDa (Nishitani et al., 2001).
RanBPM protein contains multiple conserved domains which provide potential protein-protein interaction sites such as an N-terminus proline rich domain (PRD), a splA and Ryr (SPRY) domain (212-333 aa), a lissencephaly type-I-like homology (LisH) motif (365-397 aa), a carboxy terminal to LisH (CTLH) motif (403-460 aa), and a CT11-RanBPM (CRA) motif (615-717 aa) at the C terminus of RanBPM (Murrin and Talbot, 2007). These domains play a significant role in interaction of RanBPM with a wide range of transmembrane and intracellular proteins.
RanBPM protein contains multiple conserved domains which provide potential protein-protein interaction sites such as an N-terminus proline rich domain (PRD), a splA and Ryr (SPRY) domain (212-333 aa), a lissencephaly type-I-like homology (LisH) motif (365-397 aa), a carboxy terminal to LisH (CTLH) motif (403-460 aa), and a CT11-RanBPM (CRA) motif (615-717 aa) at the C terminus of RanBPM (Murrin and Talbot, 2007). These domains play a significant role in interaction of RanBPM with a wide range of transmembrane and intracellular proteins.
Expression
RanBPM is ubiquitously expressed in human and murine cell lines and tissues of kidney, uterus, ovary, bladder, spleen, thymus, brain, skeletal muscle, lung, prostate, testes, small intestine, colon and peripheral blood lymphocytes (Rao et al., 2002; Wang et al., 2002). In contrast, reduced expression was observed in cancer cells and in several human tumor tissues.
Localisation
RanBPM is localized in the nucleus, cytoplasm, plasma membrane and cell juctions (Nishitani et al., 2001; Umeda et al., 2003; Denti et al., 2004; Chang et al., 2010).
Function
The direct binding of RanBPM with p73α, results in the nuclear translocation of cytoplasmic RanBPM. RanBPM is involved in the stabilization of p73 protein by preventing its degradation through the ubiquitin-proteasomal pathway and increases its proapoptotic function (Kramer et al., 2005). Expression of RanBPM augments T-type Ca2+ currents in HEK293/Cav3.1 cells (Kim et al., 2009). RanBPM was found to abolish the inhibitory effect of PKC on Cav3.1 currents, suggesting a key role of RanBPM in Cav3.1 channel-mediated signaling pathways. RanBPM interacts with brain-specific protein p42IP4, and modulates the p42IP4 function to regulate synaptic plasticity, actin cytoskeleton remodeling and mitogen-activated protein kinase cascade (Haase et al., 2008).
RanBPM also exhibits negative regulation. It binds with Mu opioid receptor (MOP), and inhibits the agonist-induced receptor internalization without altering MOP signaling through adenylyl cyclase, suggesting the regulatory effect of RanBPM on MOP activity in mammalian cells (Talbot et al., 2009). CD39, a prototypic member of the NTPDase family, forms a complex with RanBPM (Wu et al., 2006). This association substantially down-regulates the NTPDase activity of CD39.
RanBPM also acts as a ligand for the Rho-GEF domain (Bowman et al., 2008). RanBPM along with Rho-GEF domain of obscurin regulates the assembly of titin during the formation of the Z-disk and A/I junction, showing a vital role of RanBPM in myofibrillogenesis.
RanBPM stimulates the transcriptional activity of different proteins. RanBPM interacts with androgen receptor (AR) (Rao et al., 2002). AR belongs to a large steroid receptor family which also includes glucocorticoid (GR), progesterone, and mineralocorticoid receptors. RanBPM was found to enhance AR transactivation in a ligand-dependent fashion. Similar results were observed with GR activity, whereas the estrogen receptor activity remains unaffected. Furthermore, RanBPM was found to enhance the activity of thyroid hormone receptors (TRs), a member of nuclear receptor superfamily in a ligand-independent manner (Poirier et al., 2006), which confirms the selective action of RanBPM on receptors activity.
Expression of RanBPM influences the activity of Rta, which activates the transcription of Epstein-Barr virus (EBV) lytic genes and the lytic cycle, indicating a new role of RanBPM as a viral protein regulator. Sumoylation of Rta enhances its activity to complete the EBV lytic development in an efficient way (Chang et al., 2008). RanBPM promotes Rta sumoylation by interacting with Ubc9, which states the participation of RanBPM in EBV lytic activation.
RanBPM interacts with several proteins which are involved in neurological function. By interacting with PlexinA receptor, RanBPM mediates Semaphorin 3A signaling which is involved in axonal growth (Togashi et al., 2006). It also interacts with calbindin D28K (Lutz et al., 2003) and TAF4 to regulate neuritogenesis in neural stem cells (Brunkhorst et al., 2005). Expression of RanBPM in primary neurons decreased L1-dependent neurite outgrowth and extension (Cheng et al., 2005). RanBPM showed high expression in the Kenyon cells of the larval mushroom body (MB), and its expression is sufficient to rescue all behavioral phenotypes (Scantlebury et al., 2010). By performing genetic epistasis experiments, authors observed the participation of RanBPM with the FMRP (Fragile X Mental Retardation Protein) in the development of neuromuscular junction. Citron kinase (CITK) plays an important role in neurogenic mitoses. RanBPM potentially interacts with CITK and plays a novel role in the progression of neocortical precursors through M-phase at the ventricular surface (Chang et al., 2010).
RanBPM also exhibits negative regulation. It binds with Mu opioid receptor (MOP), and inhibits the agonist-induced receptor internalization without altering MOP signaling through adenylyl cyclase, suggesting the regulatory effect of RanBPM on MOP activity in mammalian cells (Talbot et al., 2009). CD39, a prototypic member of the NTPDase family, forms a complex with RanBPM (Wu et al., 2006). This association substantially down-regulates the NTPDase activity of CD39.
RanBPM also acts as a ligand for the Rho-GEF domain (Bowman et al., 2008). RanBPM along with Rho-GEF domain of obscurin regulates the assembly of titin during the formation of the Z-disk and A/I junction, showing a vital role of RanBPM in myofibrillogenesis.
RanBPM stimulates the transcriptional activity of different proteins. RanBPM interacts with androgen receptor (AR) (Rao et al., 2002). AR belongs to a large steroid receptor family which also includes glucocorticoid (GR), progesterone, and mineralocorticoid receptors. RanBPM was found to enhance AR transactivation in a ligand-dependent fashion. Similar results were observed with GR activity, whereas the estrogen receptor activity remains unaffected. Furthermore, RanBPM was found to enhance the activity of thyroid hormone receptors (TRs), a member of nuclear receptor superfamily in a ligand-independent manner (Poirier et al., 2006), which confirms the selective action of RanBPM on receptors activity.
Expression of RanBPM influences the activity of Rta, which activates the transcription of Epstein-Barr virus (EBV) lytic genes and the lytic cycle, indicating a new role of RanBPM as a viral protein regulator. Sumoylation of Rta enhances its activity to complete the EBV lytic development in an efficient way (Chang et al., 2008). RanBPM promotes Rta sumoylation by interacting with Ubc9, which states the participation of RanBPM in EBV lytic activation.
RanBPM interacts with several proteins which are involved in neurological function. By interacting with PlexinA receptor, RanBPM mediates Semaphorin 3A signaling which is involved in axonal growth (Togashi et al., 2006). It also interacts with calbindin D28K (Lutz et al., 2003) and TAF4 to regulate neuritogenesis in neural stem cells (Brunkhorst et al., 2005). Expression of RanBPM in primary neurons decreased L1-dependent neurite outgrowth and extension (Cheng et al., 2005). RanBPM showed high expression in the Kenyon cells of the larval mushroom body (MB), and its expression is sufficient to rescue all behavioral phenotypes (Scantlebury et al., 2010). By performing genetic epistasis experiments, authors observed the participation of RanBPM with the FMRP (Fragile X Mental Retardation Protein) in the development of neuromuscular junction. Citron kinase (CITK) plays an important role in neurogenic mitoses. RanBPM potentially interacts with CITK and plays a novel role in the progression of neocortical precursors through M-phase at the ventricular surface (Chang et al., 2010).
Homology
Human RanBPM shows 90% nucleotide homology with mouse RanBPM.
Implicated in
Entity name
Various cancers
Note
RanBPM binds to and modulates the function of a wide range of proteins. RanBPM interacts with oncoprotein Mgl-1 (Suresh et al., 2010) in mammalian cultured cells and modulates stability of Mgl-1 and functionally extends the half-life of Mgl-1 by preventing its protein turnover through the ubiquitin-proteasomal pathway. Furthermore, the overexpression of RanBPM inhibits the activity of Mgl-1 both in cell migration and colony formation assays, which reveal the novel role of RanBPM as an activator of tumor suppressor.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 18579686 | 2008 | The rho-guanine nucleotide exchange factor domain of obscurin regulates assembly of titin at the Z-disk through interactions with Ran binding protein 9. | Bowman AL et al |
| 18455188 | 2008 | Enhancement of transactivation activity of Rta of Epstein-Barr virus by RanBPM. | Chang LK et al |
| 19790105 | 2010 | RanBPM regulates the progression of neuronal precursors through M-phase at the surface of the neocortical ventricular zone. | Chang Y et al |
| 16000162 | 2005 | RanBPM is an L1-interacting protein that regulates L1-mediated mitogen-activated protein kinase activation. | Cheng L et al |
| 14722085 | 2004 | RanBPM is a phosphoprotein that associates with the plasma membrane and interacts with the integrin LFA-1. | Denti S et al |
| 18298663 | 2008 | RanBPM, a novel interaction partner of the brain-specific protein p42IP4/centaurin alpha-1. | Haase A et al |
| 18801335 | 2009 | Modulation of Ca(v)3.1 T-type Ca2+ channels by the ran binding protein RanBPM. | Kim T et al |
| 15558019 | 2005 | Protein stability and function of p73 are modulated by a physical interaction with RanBPM in mammalian cultured cells. | Kramer S et al |
| 12684061 | 2003 | Calbindin D28K interacts with Ran-binding protein M: identification of interacting domains by NMR spectroscopy. | Lutz W et al |
| 18040864 | 2007 | RanBPM, a scaffolding protein in the immune and nervous systems. | Murrin LC et al |
| 9817760 | 1998 | When overexpressed, a novel centrosomal protein, RanBPM, causes ectopic microtubule nucleation similar to gamma-tubulin. | Nakamura M et al |
| 11470507 | 2001 | Full-sized RanBPM cDNA encodes a protein possessing a long stretch of proline and glutamine within the N-terminal region, comprising a large protein complex. | Nishitani H et al |
| 16595702 | 2006 | Identification and characterization of RanBPM, a novel coactivator of thyroid hormone receptors. | Poirier MB et al |
| 12361945 | 2002 | RanBPM, a nuclear protein that interacts with and regulates transcriptional activity of androgen receptor and glucocorticoid receptor. | Rao MA et al |
| 20498842 | 2010 | The Drosophila gene RanBPM functions in the mushroom body to regulate larval behavior. | Scantlebury N et al |
| 20829363 | 2010 | Stability and function of mammalian lethal giant larvae-1 oncoprotein are regulated by the scaffolding protein RanBPM. | Suresh B et al |
| 19788913 | 2009 | Regulation of mu opioid receptor internalization by the scaffold protein RanBPM. | Talbot JN et al |
| 16672672 | 2006 | RanBPM contributes to Semaphorin3A signaling through plexin-A receptors. | Togashi H et al |
| 12559565 | 2003 | A novel nuclear protein, Twa1, and Muskelin comprise a complex with RanBPM. | Umeda M et al |
| 12147692 | 2002 | Activation of Ras/Erk pathway by a novel MET-interacting protein RanBPM. | Wang D et al |
| 16478441 | 2006 | RanBPM associates with CD39 and modulates ecto-nucleotidase activity. | Wu Y et al |
Other Information
Locus ID:
NCBI: 10048
MIM: 603854
HGNC: 13727
Ensembl: ENSG00000010017
Variants:
dbSNP: 10048
ClinVar: 10048
TCGA: ENSG00000010017
COSMIC: RANBP9
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000010017 | ENST00000011619 | Q96S59 |
| ENSG00000010017 | ENST00000011619 | A0A024QZW3 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37676377 | 2024 | ATM-Mediated translocation of RanBPM regulates DNA damage response by stabilizing p21 in non-small cell lung cancer cells. | 1 |
| 37676377 | 2024 | ATM-Mediated translocation of RanBPM regulates DNA damage response by stabilizing p21 in non-small cell lung cancer cells. | 1 |
| 29723489 | 2018 | Scaffolding protein RanBPM and its interactions in diverse signaling pathways in health and disease. | 3 |
| 30076413 | 2018 | RANBP9 affects cancer cells response to genotoxic stress and its overexpression is associated with worse response to platinum in NSCLC patients. | 10 |
| 29723489 | 2018 | Scaffolding protein RanBPM and its interactions in diverse signaling pathways in health and disease. | 3 |
| 30076413 | 2018 | RANBP9 affects cancer cells response to genotoxic stress and its overexpression is associated with worse response to platinum in NSCLC patients. | 10 |
| 28027932 | 2017 | RanBPM inhibits BLT2-mediated IL-8 production and invasiveness in aggressive breast cancer cells. | 3 |
| 28547582 | 2017 | Ran binding protein 9 (RanBPM) binds IFN-λR1 in the IFN-λ signaling pathway. | 2 |
| 28668087 | 2017 | Inhibition of HDAC6 activity through interaction with RanBPM and its associated CTLH complex. | 8 |
| 28739732 | 2017 | Reduced Expression of RanBPM Is Associated with Poorer Survival from Lung Cancer and Increased Proliferation and Invasion of Lung Cancer Cells In Vitro. | 6 |
| 28027932 | 2017 | RanBPM inhibits BLT2-mediated IL-8 production and invasiveness in aggressive breast cancer cells. | 3 |
| 28547582 | 2017 | Ran binding protein 9 (RanBPM) binds IFN-λR1 in the IFN-λ signaling pathway. | 2 |
| 28668087 | 2017 | Inhibition of HDAC6 activity through interaction with RanBPM and its associated CTLH complex. | 8 |
| 28739732 | 2017 | Reduced Expression of RanBPM Is Associated with Poorer Survival from Lung Cancer and Increased Proliferation and Invasion of Lung Cancer Cells In Vitro. | 6 |
| 26919101 | 2016 | Ubiquitin-specific protease 11 functions as a tumor suppressor by modulating Mgl-1 protein to regulate cancer cell growth. | 18 |
Citation
Kwang-Hyun Baek ; Bharathi Suresh
RANBP9 (RAN binding protein 9)
Atlas Genet Cytogenet Oncol Haematol. 2012-03-01
Online version: http://atlasgeneticsoncology.org/gene/42040/ranbp9
