RHOBTB1 (Rho-related BTB domain containing 1)
2016-02-01 Kristina Schenková , Shuo Cai , Francisco Rivero AffiliationAbstract
RhoBTB1 is one of the three members of the RhoBTB family. All RhoBTB proteins are characterized by a GTPase domain followed by a proline-rich region, a tandem of two BTB domains and a C-terminal putative RING finger domain. RHOBTB1 is a putative tumour suppressor gene. Expression of RHOBTB1 has been found decreased in diverse tumors including kidney, breast, stomach and colon cancers and head and neck squamous cell carcinomas. RhoBTB1 is a component of Cullin 3-dependent ubiquitin ligase complexes but its role in tumorigenesis is unknown.
DNA/RNA

Description
Transcription
Proteins
Note
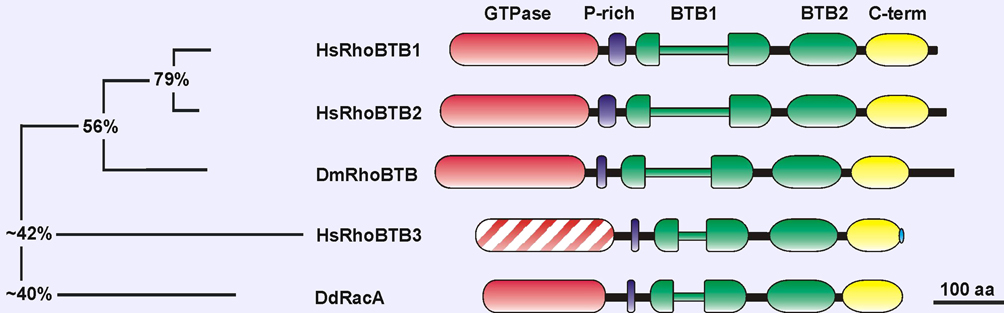
Description
The proline-rich region links the GTPase to the first BTB domain. This region could act as a SH3 domain-binding site.
The BTB domain (broad complex, tramtract and bric-a-brac) is an evolutionary conserved protein-protein interaction domain that participates in homomeric and heteromeric associations with other BTB domains. The BTB domain was also identified as a component of multimeric cullin3-dependent ubiquitin ligase complexes. The first BTB domain is bipartite, being interrupted by an insertion of unknown function. The BTB domains of RhoBTB allow the formation of homodimers and of heterodimers with other proteins of the RhoBTB family (Berthold et al., 2008).
The C-terminus is a region conserved in all members of the RhoBTB subfamily. It predictably folds as 4 consecutive alpha-helices and one beta-strand and may constitute a RING finger domain (Manjarrez et al., 2014). Many RING finger domains function as ubiquitin ligases. RhoBTB1 does not bear a CAAX motif that is typical for classical Rho GTPases and serves for localization of the protein to membranes.
Expression
Expression of RHOBTB1 has been found decreased in kidney, breast and stomach tumors in a cancer profiling array (Berthold et al., 2008), in 37% of 46 head and neck squamous cell carcinomas (Beder et al., 2006) and in colon cancer tissues (Xu et al., 2013).
RHOBTB1 is a target of the microRNA MIR31- (Alder et al., 2012; Xu et al., 2013).
Localisation
Function
RhoBTB1 binds to cullin3 and by analogy to RhoBTB2 and RhoBTB3 may constitute ubiquitin ligase complexes. RhoBTB proteins appear to exist in an inactive state through an intramolecular interaction of the BTB domain region with the GTPase domain (Berthold et al., 2008). This model has been refined recently for RhoBTB2 to show that the HSP90AA1 (Hsp90) chaperone machinery unlocks RhoBTB, enabling GTP binding and interaction with Cullin 3 and the COPS8 (COP9) signalosome. COP9 deneddylates Cullin 3 and stabilizes the complex (Manjarrez et al. 2014). Considering the high degree of similarity between RhoBTB2 and RhoBTB1, this mechanism is very likely to apply to RhoBTB1 too.
RHOBTB1 has been identified as a target gene of the nuclear hormone receptor PPARG. RhoBTB1 mRNA and protein levels are decreased in the aorta of mice expressing a dominant negative PPARG. These mice also present a concomitant decrease in Cullin 3, and it has been proposed that RhoBTB1 regulates Cullin 3 levels or activity, which in turn regulates RHOA. turnover in smooth muscle. RhoBTB1 emerges as a component of a signaling mechanism that regulates vascular function and blood pressure (Pelham et al., 2012).
RhoBTB1, like RhoBTB2, is required for expression of the chemokine CXCL14 in keratinocytes independently of Cullin3-mediated protein degradation (McKinnon et al., 2008).
RhoBTB1 displays only a moderate influence on the morphology and actin organisation of porcine aortic endothelial cells upon ectopic expression. It does not interact with the GTPase-binding domain of WASP, PAK1 or RTKN (Rhotekin), which are well-known effectors of many typical Rho GTPases (Aspenstrom et al., 2004).
RhoBTB1, like RhoBTB2 and RhoBTB3, interacts with LLRC41 (leucine rich repeat containing 41, MUF1). MUF-1 is a nuclear protein and carries a BC-box that functions as a linker in multicomponent Cullin 5-dependent ubiquitin ligase complexes (Schenková et al., 2012). MUF1 may be a substrate for RhoBTB-Cullin 3 ubiquitin ligase complexes. The function of MUF1 is unknown, but it is suspected to be involved in the DNA damage response
Homology
Mutations
Note
Implicated in
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 22689922 | 2012 | Dysregulation of miR-31 and miR-21 induced by zinc deficiency promotes esophageal cancer. | Alder H et al |
| 14521508 | 2004 | Rho GTPases have diverse effects on the organization of the actin filament system. | Aspenström P et al |
| 16170569 | 2006 | Identification of a candidate tumor suppressor gene RHOBTB1 located at a novel allelic loss region 10q21 in head and neck cancer. | Beder LB et al |
| 18298893 | 2008 | Rho GTPases of the RhoBTB subfamily and tumorigenesis. | Berthold J et al |
| 24608665 | 2014 | Hsp90-dependent assembly of the DBC2/RhoBTB2-Cullin3 E3-ligase complex. | Manjarrez JR et al |
| 18762809 | 2008 | The atypical Rho GTPase RhoBTB2 is required for expression of the chemokine CXCL14 in normal and cancerous epithelial cells. | McKinnon CM et al |
| 9872452 | 1998 | Prediction of the coding sequences of unidentified human genes. XI. The complete sequences of 100 new cDNA clones from brain which code for large proteins in vitro. | Nagase T et al |
| 23040068 | 2012 | Cullin-3 regulates vascular smooth muscle function and arterial blood pressure via PPARγ and RhoA/Rho-kinase. | Pelham CJ et al |
| 12426103 | 2002 | Genomic organization and expression profile of the small GTPases of the RhoBTB family in human and mouse. | Ramos S et al |
| 11222756 | 2001 | The Dictyostelium discoideum family of Rho-related proteins. | Rivero F et al |
| 22709582 | 2012 | MUF1/leucine-rich repeat containing 41 (LRRC41), a substrate of RhoBTB-dependent cullin 3 ubiquitin ligase complexes, is a predominantly nuclear dimeric protein. | Schenková K et al |
| 23258531 | 2013 | The tumor suppressor gene RhoBTB1 is a novel target of miR-31 in human colon cancer. | Xu RS et al |
Other Information
Locus ID:
NCBI: 9886
MIM: 607351
HGNC: 18738
Ensembl: ENSG00000072422
Variants:
dbSNP: 9886
ClinVar: 9886
TCGA: ENSG00000072422
COSMIC: RHOBTB1
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000072422 | ENST00000337910 | O94844 |
| ENSG00000072422 | ENST00000337910 | A0A024QZL4 |
| ENSG00000072422 | ENST00000357917 | O94844 |
| ENSG00000072422 | ENST00000357917 | A0A024QZL4 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37575477 | 2023 | Structure and Function of RhoBTB1 Required for Substrate Specificity and Cullin-3 Ubiquitination. | 1 |
| 37575477 | 2023 | Structure and Function of RhoBTB1 Required for Substrate Specificity and Cullin-3 Ubiquitination. | 1 |
| 32354068 | 2020 | RNA Interference Screening Identifies Novel Roles for RhoBTB1 and RhoBTB3 in Membrane Trafficking Events in Mammalian Cells. | 4 |
| 32354068 | 2020 | RNA Interference Screening Identifies Novel Roles for RhoBTB1 and RhoBTB3 in Membrane Trafficking Events in Mammalian Cells. | 4 |
| 31431478 | 2019 | RhoBTB1 interacts with ROCKs and inhibits invasion. | 8 |
| 31431478 | 2019 | RhoBTB1 interacts with ROCKs and inhibits invasion. | 8 |
| 28219369 | 2017 | The tumor suppressor RhoBTB1 controls Golgi integrity and breast cancer cell invasion through METTL7B. | 44 |
| 28219369 | 2017 | The tumor suppressor RhoBTB1 controls Golgi integrity and breast cancer cell invasion through METTL7B. | 44 |
| 23258531 | 2013 | The tumor suppressor gene RhoBTB1 is a novel target of miR-31 in human colon cancer. | 36 |
| 23258531 | 2013 | The tumor suppressor gene RhoBTB1 is a novel target of miR-31 in human colon cancer. | 36 |
| 16170569 | 2006 | Identification of a candidate tumor suppressor gene RHOBTB1 located at a novel allelic loss region 10q21 in head and neck cancer. | 18 |
| 16385451 | 2006 | A scan of chromosome 10 identifies a novel locus showing strong association with late-onset Alzheimer disease. | 83 |
| 16170569 | 2006 | Identification of a candidate tumor suppressor gene RHOBTB1 located at a novel allelic loss region 10q21 in head and neck cancer. | 18 |
| 16385451 | 2006 | A scan of chromosome 10 identifies a novel locus showing strong association with late-onset Alzheimer disease. | 83 |
Citation
Kristina Schenková ; Shuo Cai ; Francisco Rivero
RHOBTB1 (Rho-related BTB domain containing 1)
Atlas Genet Cytogenet Oncol Haematol. 2016-02-01
Online version: http://atlasgeneticsoncology.org/gene/42981/rhobtb1
Historical Card
2008-12-01 RHOBTB1 (Rho-related BTB domain containing 1) by Kristina Schenková,Francisco Rivero Affiliation
