RPL10 (ribosomal protein L10)
2010-08-01 Mohit Goel , Ranjan Tamuli AffiliationDepartment of Biotechnology, Indian Institute of Technology Guwahati, Guwahati-781 039, Assam, India
Identity
HGNC
LOCATION
Xq28
LOCUSID
ALIAS
DKFZp686J1851,DXS648,DXS648E,FLJ23544,FLJ27072,NOV,QM
FUSION GENES
DNA/RNA
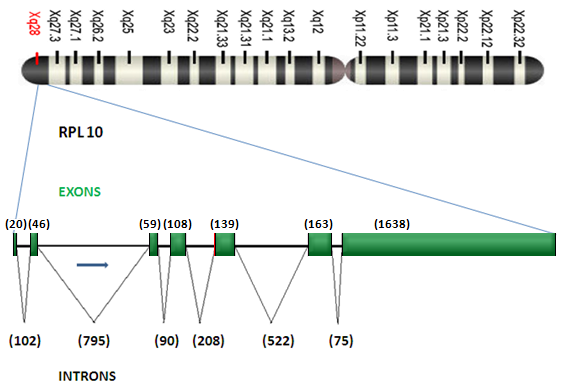
Description
DNA size 3.96 kb, mRNA size 2172 bp, 7 exons. The RPL10 gene is co-transcribed with the small nucleolar RNA gene U70 that is located in its fifth intron. Multiple processed pseudogenes of the gene RPL10 are dispersed in the genome. Moreover, transcript variants utilizing alternative polyA signals exist; the variant with the longest 3 UTR overlaps the deoxyribonuclease I-like 1 gene on the opposite strand.
Proteins
Expression
Ubiquitous. RPL10 is expressed in a wide variety of embryonic and adult tissues, down-regulated during adipocyte, kidney, and heart differentiation.
Localisation
Cytoplasm.
Function
The ribosomal protein L10 (RPL10), a member of the L10E family of ribosomal proteins, is a key protein in assembling 60S ribosomal subunit and organizes the architecture of the aminoacyl-tRNA binding site. RPL10 was originally identified as QM, a candidate for a Wilms tumor suppressor; however, later studies did not support the original hypothesis. In vitro studies have shown the interaction of RPL10 with the transcript regulator the c-Jun, as well as with the proto-oncogene c-Yes; however, these interactions yet to demonstrate in vivo.
Homology
The percent identity below represents identity of RPL10 over an aligned region in UniGene.
- Mus musculus: 100 (percent identity)
- Xenopus tropicalis: 99.5
- Monodelphis domestica: 99.5
- Pan troglodytes: 99.5
- Xenopus laevis: 99.1
- Danio rerio: 97.7
- Drosophila melanogaster: 88.9
- Caenorhabditis elegans: 85.5
- Neurospora crassa: 84.7
- Saccharomyces cerevisiae: 78.3
- Mus musculus: 100 (percent identity)
- Xenopus tropicalis: 99.5
- Monodelphis domestica: 99.5
- Pan troglodytes: 99.5
- Xenopus laevis: 99.1
- Danio rerio: 97.7
- Drosophila melanogaster: 88.9
- Caenorhabditis elegans: 85.5
- Neurospora crassa: 84.7
- Saccharomyces cerevisiae: 78.3
Mutations
Note
Two missense mutations L206M and H213Q at the C-terminal end of RPL10 were identified in two independent families with autism, a disorder of neural development.
Implicated in
Entity name
Prostatic adenocarcinoma
Note
RPL10 gene showed up-regulation in androgen-independent C81 passage cells, derived from the LNCaP cell model that recapitulates prostate cancer progression. In a study using immunohistochemical technique, human prostatic tissues showed expression of RPL10 protein in all normal prostate glands adjacent to prostate cancer and in various intraepithelial neoplasia (PIN). However, in prostate cancer, the staining intensity and stained areas were decreased, compared to the normal glands and PIN lesions. There was an inverse correlation from normal to low-grade tumors and then to high-grade tumors. In high-grade tumors, the positive areas were mostly confined to peripheral aspects of tumors and were particularly strong in foci of perineural invasion. These results suggested that decreased RPL10 expression may be associated with early development of prostate cancer, but later a high level of RPL10 may facilitate progression of the tumors to a more aggressive phenotype.
Entity name
Ovarian cancer
Note
Both adenine (A)/guanine (G) replacement was detected at the 605th nucleotide which changes the coding from serine to asparagines in 17 (58.6%) of the 29 ovarian tumors studied. The frequencies of A/A, G/G and A/G homo- or hetero-zygosity were 3.5%, 37.9% and 58.6%, respectively in cancer tissues but they were 26.1%, 52.2% and 21.7%, respectively in the adjacent normal tissues, indicating a higher heterozygous rate in cancer (58.6% vs 21.7%, p
Entity name
Wilms tumor
Note
RPL10 was originally isolated by subtractive hybridization between a tumorigenic cell line (deleted for part of 11p) and a non-tumorigenic cell line (the tumorigenic cell line carrying an extra t(X;11) translocation chromosome). The RPL10 mRNA level was found modulated between the tumorigenic and nontumorigenic cell lines and suspected to be involved in the maintenance of the nontumorigenic phenotype. However, later study had shown that the RPL10 gene is X-linked and therefore not involved in suppression of tumorigenesis in Wilms tumor.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 16331298 | 2006 | Reduction of QM protein expression correlates with tumor grade in prostatic adenocarcinoma. | Altinok G et al |
| 1658743 | 1991 | The isolation and characterization of a novel cDNA demonstrating an altered mRNA level in nontumorigenic Wilms' microcell hybrid cells. | Dowdy SF et al |
| 19166581 | 2009 | An investigation of ribosomal protein L10 gene in autism spectrum disorders. | Gong X et al |
| 12082018 | 2002 | Expression profile of differentially-regulated genes during progression of androgen-independent growth in human prostate cancer cells. | Karan D et al |
| 16940977 | 2006 | Mutations in the ribosomal protein gene RPL10 suggest a novel modulating disease mechanism for autism. | Klauck SM et al |
| 16627977 | 2006 | Loss of heterozygosity and microsatellite instability at the Xq28 and the A/G heterozygosity of the QM gene are associated with ovarian cancer. | Shen XJ et al |
| 1330878 | 1992 | The QM gene is X-linked and therefore not involved in suppression of tumorigenesis in Wilms' tumor. | van den Ouweland AM et al |
Other Information
Locus ID:
NCBI: 6134
MIM: 312173
HGNC: 10298
Ensembl: ENSG00000147403
Variants:
dbSNP: 6134
ClinVar: 6134
TCGA: ENSG00000147403
COSMIC: RPL10
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37280198 | 2023 | The ufmylation modification of ribosomal protein L10 in the development of pancreatic adenocarcinoma. | 5 |
| 37280198 | 2023 | The ufmylation modification of ribosomal protein L10 in the development of pancreatic adenocarcinoma. | 5 |
| 29930300 | 2019 | The ribosomal RPL10 R98S mutation drives IRES-dependent BCL-2 translation in T-ALL. | 35 |
| 29930300 | 2019 | The ribosomal RPL10 R98S mutation drives IRES-dependent BCL-2 translation in T-ALL. | 35 |
| 28744013 | 2018 | The T-cell leukemia-associated ribosomal RPL10 R98S mutation enhances JAK-STAT signaling. | 40 |
| 29066376 | 2018 | A de novo mutation in RPL10 causes a rare X-linked ribosomopathy characterized by syndromic intellectual disability and epilepsy: A new case and review of the literature. | 14 |
| 30172100 | 2018 | Ribosomal protein L10 in mitochondria serves as a regulator for ROS level in pancreatic cancer cells. | 19 |
| 28744013 | 2018 | The T-cell leukemia-associated ribosomal RPL10 R98S mutation enhances JAK-STAT signaling. | 40 |
| 29066376 | 2018 | A de novo mutation in RPL10 causes a rare X-linked ribosomopathy characterized by syndromic intellectual disability and epilepsy: A new case and review of the literature. | 14 |
| 30172100 | 2018 | Ribosomal protein L10 in mitochondria serves as a regulator for ROS level in pancreatic cancer cells. | 19 |
| 28428269 | 2017 | Low frequency mutations in ribosomal proteins RPL10 and RPL5 in multiple myeloma. | 18 |
| 28428269 | 2017 | Low frequency mutations in ribosomal proteins RPL10 and RPL5 in multiple myeloma. | 18 |
| 27726420 | 2016 | Mitochondrial Ribosomal Protein L10 Associates with Cyclin B1/Cdk1 Activity and Mitochondrial Function. | 18 |
| 27726420 | 2016 | Mitochondrial Ribosomal Protein L10 Associates with Cyclin B1/Cdk1 Activity and Mitochondrial Function. | 18 |
| 25846674 | 2015 | RPL10 mutation segregating in a family with X-linked syndromic Intellectual Disability. | 17 |
Citation
Mohit Goel ; Ranjan Tamuli
RPL10 (ribosomal protein L10)
Atlas Genet Cytogenet Oncol Haematol. 2010-08-01
Online version: http://atlasgeneticsoncology.org/gene/42148/rpl10
