SELENOP (selenoprotein P, plasma, 1)
2011-03-01 Ewa Jablonska , Jolanta Gromadzinska , Edyta Reszka , Wojciech Wasowicz AffiliationNofer Institute of Occupational Medicine, St Teresy 8, 91-348 Lodz, Poland
DNA/RNA
Description
The genomic DNA of SEPP1 spans about 12 kb. SEPP1 consists of 6 exons.
Transcription
3 alternative mRNAs exist encoding 2 different isoforms of selenoprotein P. Transcript variants 1 and 2 encode isoform 1 and transcript variant 3 encodes isoform 2.
Proteins
Note
SEPP1 belongs to selenoproteins, all of which contain selenium in the form of selenocysteine (SeC) and are being synthesized in the presence of UGA codon, specific stem loop structure in 3 UTR of mRNA called SECIS (Selenocysteine Insertion Sequence) and other specific factors. Selenoprotein P is a glycoprotein present mainly in plasma, where it accounts for 40 - 65% of total selenium in this blood compartment. Plasma SEPP1 concentration is regarded as a functional biomarker of human selenium status (Saito and Takahashi, 2002; Méplan et al., 2009; Xia et al., 2010).
Description
Selenoprotein P consists of 381 amino acids and contains ten selenocysteines: nine are located in Sec-rich C-terminal domain (suggested as the region responsible for selenium delivery) and one is present in N-terminal domain (region with redox properties responsible for enzymatic activity of the protein). Two protein isoforms were identified in human plasma: 50 kDa and 60 kDa (Méplan et al., 2007; Méplan et al., 2009).
Expression
SEPP1 is expressed mainly in the liver, from where it is exported to plasma and other tissues. Other organs expressing the protein include mainly brain, thyroid gland, prostate and mammary gland. Its expression has been found to be significantly reduced in cancer, including prostate, colon and lung (Gonzalez-Moreno et al., 2010).
SEPP1 expression is downregulated by different cytokines (Al-Taie et al., 2002). Also hepatic factors such as insulin and glucocorticoids may regulate SEPP1 expression (Speckmann et al., 2008).
SEPP1 expression is downregulated by different cytokines (Al-Taie et al., 2002). Also hepatic factors such as insulin and glucocorticoids may regulate SEPP1 expression (Speckmann et al., 2008).
Localisation
Plasma.
Function
It is supposed that SeP is responsible for the transport of selenium within body and delivering the microelement to the cells. In brain and testis (organs, in which selenium plays an important role), SEPP1 uptake is mediated by apolipoprotein E receptor-2 (apoER2). In kidneys, the uptake is regulated by another receptor, called megalin (Burk and Hill, 2009) (figure 1).
Additionally, SEPP1 is involved in the reduction of oxidative stress due to its redox properties (Saito et al., 2004).
Additionally, SEPP1 is involved in the reduction of oxidative stress due to its redox properties (Saito et al., 2004).
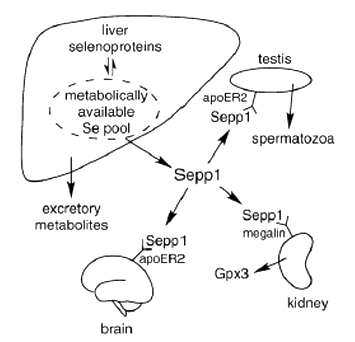
Figure 1. SEPP1 in selenium homeostasis and transport to the testis, brain and kidney. Whole - body selenium excretion is controlled by selenium excretion in the urine. SEPP1 and selenium excretory metabolites compete for metabolically available selenium in the liver, determining urinary selenium excretion. The lipoprotein receptor apoER2 binds SEPP1 and facilitates its uptake into the testis where selenium is incorporated into spermatozoa. ApoER2 also maintains brain selenium. SEPP1 is filtered by the kidney into the glomerular filtrate and binds to megalin in the brush border of proximal convoluted tubules. Those cells endocytose the SEPP1 bound to megalin and presumably use its selenium to synthesize plasma glutathione peroxidase (GPx3) (adapted from Burk and Hill, 2009, with the authors permission).
Homology
SEPP1 is conserved in chimpanzee, dog, cow, mouse, rat and zebrafish.
Mutations
Note
No mutations in SEPP1 gene have been identified yet.
Genetic variations: several SNPs were identified.
Most often studied polymorphisms within SEPP1:
- Ala234Thr (rs 3877899) - associated with a G/A transition at position 24731 of mRNA, with the amino acid change from alanine to threonine in the codon 234. This polymorphism influences the SePP1 isoform pattern. Using Western blot analysis, it was shown that in the individuals possessing Ala/Ala genotype, 60 kDa protein was a dominating isoform, whereas in those with Ala/Thr genotype, the band for 50 kDa isoform was stronger (Thr/Thr genotype was not analyzed in this study). It was also observed, that within Ala/Ala and Ala/Thr individuals, males had less 60 kDa isoform as compared to females (Méplan et al., 2009).
- r25191g/a (rs 7579) - G/A transition at position 25191 within 3-UTR. Similarly as rs 3877899 SNP, this SNP seems to influence the proportion of SEPP1 isoforms. Individuals with GG genotype had lower proportion of 60 kDa isoform as compared to those with GA or AA genotype (Méplan et al., 2009).
- (TC)5/(TC)3 repeats at promoter region. It was shown in in vitro study that TC3 allele reduced the promoter activity of reporter gene constructs in HepG2 cells (Al-Taie et al., 2002).
Genetic variations: several SNPs were identified.
Most often studied polymorphisms within SEPP1:
- Ala234Thr (rs 3877899) - associated with a G/A transition at position 24731 of mRNA, with the amino acid change from alanine to threonine in the codon 234. This polymorphism influences the SePP1 isoform pattern. Using Western blot analysis, it was shown that in the individuals possessing Ala/Ala genotype, 60 kDa protein was a dominating isoform, whereas in those with Ala/Thr genotype, the band for 50 kDa isoform was stronger (Thr/Thr genotype was not analyzed in this study). It was also observed, that within Ala/Ala and Ala/Thr individuals, males had less 60 kDa isoform as compared to females (Méplan et al., 2009).
- r25191g/a (rs 7579) - G/A transition at position 25191 within 3-UTR. Similarly as rs 3877899 SNP, this SNP seems to influence the proportion of SEPP1 isoforms. Individuals with GG genotype had lower proportion of 60 kDa isoform as compared to those with GA or AA genotype (Méplan et al., 2009).
- (TC)5/(TC)3 repeats at promoter region. It was shown in in vitro study that TC3 allele reduced the promoter activity of reporter gene constructs in HepG2 cells (Al-Taie et al., 2002).
Implicated in
Entity name
Cancer
Note
Persson-Moschos et al. (2000) conducted a nested case control study, in which 12500 middle aged men were enrolled and the follow up time was between 1974-1988. Within the studied cohort, SEPP1 plasma concentration was significantly lower in the individuals who were diagnosed with cancer during the follow up (and whose plasma samples were available for analysis, n=302) as compared to control subjects (n=604). The authors of this study suggested that plasma SEPP1 level is associated with higher risk of cancer of respiratory and digestive tract.
Entity name
Colon cancer
Note
Decreased expression of SePP1 mRNA was observed in colorectal cancer tissue as compared with normal colon mucosa (Al-Taie et al., 2004).
The study of 196 cases and 239 controls revealed no association between polymorphism at SEPP1 promoter region (TC)5/(TC)3 repeats and colon cancer risk. However, authors of the study observed genomic instability of Poly-(T)-single nucleotide repeat motif present in the SEPP1 promoter sequence in the colon cancer tissues as compared to normal colon mucosa from the same patients. This instability was observed in 10 out of 51 cases possessing two (TC)5 alleles (no instability was observed in 5 cases with (TC)5/(TC)3 genotype; (TC)3/(TC)3 homozygotes were not present in the studied group) (Al-Taie et al., 2002).
Other study, involving 772 cases and 777 controls, revealed that four SNPs within SEPP1 gene were significantly associated with risk of colorectal adenoma. The SNPs were: SEPP1 -4166G, rs12055266, rs3797310, rs2972994 (Peters et al., 2008).
In the study by Méplan et al. (2009), in which plasma samples from 20 colon cancer patients and 21 healthy individuals were analyzed, significantly lower proportion of 60 kDa isoform was observed in cases with SEPP1 GG genotype of rs 3877899 as compared to controls with the same genotype. Similar (statistically significant) difference between cases and controls was also indicated within individuals possessing GA genotype of rs7579.
The study of 196 cases and 239 controls revealed no association between polymorphism at SEPP1 promoter region (TC)5/(TC)3 repeats and colon cancer risk. However, authors of the study observed genomic instability of Poly-(T)-single nucleotide repeat motif present in the SEPP1 promoter sequence in the colon cancer tissues as compared to normal colon mucosa from the same patients. This instability was observed in 10 out of 51 cases possessing two (TC)5 alleles (no instability was observed in 5 cases with (TC)5/(TC)3 genotype; (TC)3/(TC)3 homozygotes were not present in the studied group) (Al-Taie et al., 2002).
Other study, involving 772 cases and 777 controls, revealed that four SNPs within SEPP1 gene were significantly associated with risk of colorectal adenoma. The SNPs were: SEPP1 -4166G, rs12055266, rs3797310, rs2972994 (Peters et al., 2008).
In the study by Méplan et al. (2009), in which plasma samples from 20 colon cancer patients and 21 healthy individuals were analyzed, significantly lower proportion of 60 kDa isoform was observed in cases with SEPP1 GG genotype of rs 3877899 as compared to controls with the same genotype. Similar (statistically significant) difference between cases and controls was also indicated within individuals possessing GA genotype of rs7579.
Entity name
Prostate cancer
Note
Expression of SEPP1 mRNA was down regulated in human prostate tumours as well as prostate carcinoma cell lines (Calvo et al., 2002).
Further investigation revealed that down-regulation of SEPP1 in prostate cancer cells leads to an increased production of reactive oxygen species (Gonzalez-Moreno et al., 2010).
In the study conducted on 90 males with prostate cancer and 100 control men, it was observed that proteins concentration measured in serum was lower in cases as compared to controls (Meyer et al., 2009).
The interaction between polymorphic variants of SePP1 and SOD2 genes in prostate cancer risk was found by Cooper et al. (2008). According to the results based on CAPS study (Prostate CAncer in Sweden), males being homozygous for SEPP1 Ala234 allele (rs 3877899) and who possessed at least one SOD2 Ala16 allele (rs 4880), were at significantly higher risk of prostate cancer.
In a study of 248 prostate cancer cases and 492 controls, borderline significant association between prostate cancer risk and SEPP1 polymorphism (rs 7579) was found (Steinbrecher et al., 2010).
Further investigation revealed that down-regulation of SEPP1 in prostate cancer cells leads to an increased production of reactive oxygen species (Gonzalez-Moreno et al., 2010).
In the study conducted on 90 males with prostate cancer and 100 control men, it was observed that proteins concentration measured in serum was lower in cases as compared to controls (Meyer et al., 2009).
The interaction between polymorphic variants of SePP1 and SOD2 genes in prostate cancer risk was found by Cooper et al. (2008). According to the results based on CAPS study (Prostate CAncer in Sweden), males being homozygous for SEPP1 Ala234 allele (rs 3877899) and who possessed at least one SOD2 Ala16 allele (rs 4880), were at significantly higher risk of prostate cancer.
In a study of 248 prostate cancer cases and 492 controls, borderline significant association between prostate cancer risk and SEPP1 polymorphism (rs 7579) was found (Steinbrecher et al., 2010).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15203372 | 2004 | Expression profiling and genetic alterations of the selenoproteins GI-GPx and SePP in colorectal carcinogenesis. | Al-Taie OH et al |
| 19345254 | 2009 | Selenoprotein P-expression, functions, and roles in mammals. | Burk RF et al |
| 12235003 | 2002 | Alterations in gene expression profiles during prostate cancer progression: functional correlations to tumorigenicity and down-regulation of selenoprotein-P in mouse and human tumors. | Calvo A et al |
| 19074884 | 2008 | Interaction between single nucleotide polymorphisms in selenoprotein P and mitochondrial superoxide dismutase determines prostate cancer risk. | Cooper ML et al |
| 20181815 | 2010 | Establishing optimal selenium status: results of a randomized, double-blind, placebo-controlled trial. | Hurst R et al |
| 17536041 | 2007 | Genetic polymorphisms in the human selenoprotein P gene determine the response of selenoprotein markers to selenium supplementation in a gender-specific manner (the SELGEN study). | Méplan C et al |
| 20378690 | 2010 | Genetic variants in selenoprotein genes increase risk of colorectal cancer. | Méplan C et al |
| 19453253 | 2009 | Relative abundance of selenoprotein P isoforms in human plasma depends on genotype, se intake, and cancer status. | Méplan C et al |
| 19690186 | 2009 | Reduced serum selenoprotein P concentrations in German prostate cancer patients. | Meyer HA et al |
| 10798212 | 2000 | Selenoprotein P in plasma in relation to cancer morbidity in middle-aged Swedish men. | Persson-Moschos ME et al |
| 18483336 | 2008 | Variation in the selenoenzyme genes and risk of advanced distal colorectal adenoma. | Peters U et al |
| 15117283 | 2004 | Domain structure of bi-functional selenoprotein P. | Saito Y et al |
| 12423375 | 2002 | Characterization of selenoprotein P as a selenium supply protein. | Saito Y et al |
| 18972406 | 2008 | Selenoprotein P expression is controlled through interaction of the coactivator PGC-1alpha with FoxO1a and hepatocyte nuclear factor 4alpha transcription factors. | Speckmann B et al |
| 20852007 | 2010 | Effects of selenium status and polymorphisms in selenoprotein genes on prostate cancer risk in a prospective study of European men. | Steinbrecher A et al |
| 20573787 | 2010 | Optimization of selenoprotein P and other plasma selenium biomarkers for the assessment of the selenium nutritional requirement: a placebo-controlled, double-blind study of selenomethionine supplementation in selenium-deficient Chinese subjects. | Xia Y et al |
Other Information
Locus ID:
NCBI: 6414
MIM: 601484
HGNC: 10751
Ensembl: ENSG00000250722
Variants:
dbSNP: 6414
ClinVar: 6414
TCGA: ENSG00000250722
COSMIC: SELENOP
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37391553 | 2024 | Association of Serum Selenium and Selenoprotein P with Oxidative Stress Biomarkers in Patients with Polycystic Ovary Syndrome. | 0 |
| 37495908 | 2024 | Selenoprotein P, peroxiredoxin-5, renalase, and total antioxidant status in patients with suspected obstructive sleep apnea. | 0 |
| 38182643 | 2024 | Selenoprotein P expression in glioblastoma as a regulator of ferroptosis sensitivity: preservation of GPX4 via the cycling-selenium storage. | 0 |
| 38198038 | 2024 | Strong associations of serum selenoprotein P with all-cause mortality and mortality due to cancer, cardiovascular, respiratory and gastrointestinal diseases in older German adults. | 1 |
| 37391553 | 2024 | Association of Serum Selenium and Selenoprotein P with Oxidative Stress Biomarkers in Patients with Polycystic Ovary Syndrome. | 0 |
| 37495908 | 2024 | Selenoprotein P, peroxiredoxin-5, renalase, and total antioxidant status in patients with suspected obstructive sleep apnea. | 0 |
| 38182643 | 2024 | Selenoprotein P expression in glioblastoma as a regulator of ferroptosis sensitivity: preservation of GPX4 via the cycling-selenium storage. | 0 |
| 38198038 | 2024 | Strong associations of serum selenoprotein P with all-cause mortality and mortality due to cancer, cardiovascular, respiratory and gastrointestinal diseases in older German adults. | 1 |
| 37108730 | 2023 | SELENOP rs3877899 Variant Affects the Risk of Developing Advanced Stages of Retinopathy of Prematurity (ROP). | 1 |
| 37166989 | 2023 | SELENOP modifies sporadic colorectal carcinogenesis and WNT signaling activity through LRP5/6 interactions. | 4 |
| 37234930 | 2023 | Lower Plasma Selenoprotein P Levels in Regularly Exercising Young Adults. | 1 |
| 37423160 | 2023 | Autoantibodies to selenoprotein P in chronic fatigue syndrome suggest selenium transport impairment and acquired resistance to thyroid hormone. | 7 |
| 37423559 | 2023 | Selenoprotein P deficiency is associated with higher risk of incident heart failure. | 0 |
| 37108730 | 2023 | SELENOP rs3877899 Variant Affects the Risk of Developing Advanced Stages of Retinopathy of Prematurity (ROP). | 1 |
| 37166989 | 2023 | SELENOP modifies sporadic colorectal carcinogenesis and WNT signaling activity through LRP5/6 interactions. | 4 |
Citation
Ewa Jablonska ; Jolanta Gromadzinska ; Edyta Reszka ; Wojciech Wasowicz
SELENOP (selenoprotein P, plasma, 1)
Atlas Genet Cytogenet Oncol Haematol. 2011-03-01
Online version: http://atlasgeneticsoncology.org/gene/46513/selenop
