STOML2 (stomatin (EPB72)-like 2)
2010-02-01 Wenfeng Cao , Liyong Zhang , Fang Ding , Zhumei Cui , Zhihua Liu AffiliationIdentity
HGNC
LOCATION
9p13.3
LOCUSID
ALIAS
HSPC108,SLP-2
FUSION GENES
DNA/RNA

Description
The gene encoding SLP-2 was 3250 bp long and consisted of ten exons interrupted by nine introns.
Transcription
There are 5 transcripts in this gene. However, a single 1.3 kb mRNA transcript encoding SLP-2 was ubiquitously expressed, and the translation length is 356 residues (Owczarek et al., 2001).
Pseudogene
No known pseudogenes.
Proteins
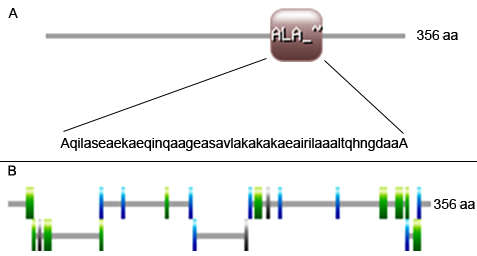
Figure A. ALA-RICH, Alanine-rich region profile: 224-274: score = 8.657.
Figure B. MYRISTYL, N-myristoylation site: 16-21: GSllAS, 31-36: GLprNT, 209-214: GTreSA, 314-319: GVvgAL, 326-331: GTpdSL, 341-346: GtdaSL; PKC-PHOSPHO-SITE, Protein kinase C phosphorylation site: 21-23: SgR, 78-80: SlK, 133-135: TmR, 335-337: SsR; CAMP-PHOSPHO-SITE, Camp-and-cGMP-dependent protein kinase phosphorylation site: 26-29: RRaS, 200-203: KRaT; CK2-PHOSPHO-SITE, Casein kinase II phosphorylation site: 78-81: SlkE, 156-159: SivD, 203-206: TvlE, 229-232: SeaE, 277-280: TvaE, 335-338: SsrD, 345-348: SldE; ASN-GLYCOSYLATION, N-glycosylation site: 96-99: NVTL, 154-157: NASI; AMIDATION, amidation site: 219-222: eGKK.
Description
NP_038470; 356 aa.
Human SLP-2 is presented on chromosome 9p13. The sequence at the 5-end of the mRNA is interesting for the presence of three potential ATG initiator sites, all sharing the same open reading frame however, commonly forms a 356 amino acid residue polypeptide with a predicted molecular weight of 38.5 kDa. Similar to other family members, SLP-2 as well as the stomatin from other species shares a characteristic NH2-terminal hydrophobic domain as well as a consensus cognate stomatin signature sequence that defines the stomatin gene family, however, it lacks the NH2-terminal hydrophobic domain (Wang et al., 2000). The SLP-2 protein contains an alanine-rich domain and a number of potential protein kinase C phosphorylation sites, cAMP-and-cGMP-dependent protein kinase phosphorylation sites and casein kinase II phosphorylation sites.
Human SLP-2 is presented on chromosome 9p13. The sequence at the 5-end of the mRNA is interesting for the presence of three potential ATG initiator sites, all sharing the same open reading frame however, commonly forms a 356 amino acid residue polypeptide with a predicted molecular weight of 38.5 kDa. Similar to other family members, SLP-2 as well as the stomatin from other species shares a characteristic NH2-terminal hydrophobic domain as well as a consensus cognate stomatin signature sequence that defines the stomatin gene family, however, it lacks the NH2-terminal hydrophobic domain (Wang et al., 2000). The SLP-2 protein contains an alanine-rich domain and a number of potential protein kinase C phosphorylation sites, cAMP-and-cGMP-dependent protein kinase phosphorylation sites and casein kinase II phosphorylation sites.
Expression
SLP-2 is widely expressed in many tissues and thought as a new component of the peripheral membrane skeleton. Especially, in the erythrocyte membrane, it also appears to exist at least partially as an oligomeric protein complex. The overexpression of SLP-2 can be found in many kinds of human tumors, such as esophageal squamous cell carcinoma, laryngeal squamous cell carcinoma, endometrial adenocarcinoma, and lung cancer.
Localisation
Predominantly on plasma membrane and in the cytoplasm.
Function
Human SLP-2 protein with unknown function, we hypothesize that SLP-2 may link stomatin or other integral membrane proteins to the peripheral cytoskeleton and thereby play a role in regulating ion channel conductances or the organization of sphingolipid and cholesterol-rich lipid rafts. Some recent results indicated that SLP-2 protein can significantly influence on multi-tumor progression, which allowed us to identify this unwell-known gene that maybe modulate invasion and metastasis of different cancers.
Homology
SLP-2 is one of the members of the Stomatin superfamily, among which identified vertebrate homologues are SLP-1, SLP-2, and SLP-3. SLP-1 is most abundant in brain and shares many similarities with UNC-24 (STOML1). SLP-3 is specifically expressed in olfactory sensory neurons (Seidel et al., 1998; Goldstein et al., 2003).
Mutations
Note
No mutations have been reported for SLP-2. Mutation detection of SLP-2 exons was done using PCR and automated sequencing with 30 patient-matched human esophageal cancer tissues. No mutation was found within the open-reading frame of SLP-2 after sequencing results were aligned by the procedure SeqMan of DNAStar software (Zhang et al., 2006).
Implicated in
Entity name
Various cancers
Note
SLP-2 has been shown to be over-expressed in a number of different cancers, including esophageal squamous cell carcinoma (ESCC), laryngeal squamous cell carcinoma (LSCC), endometrial adenocarcinoma (EAC), lung cancer (LC) and breast cancer (see below).
Entity name
Esophageal squamous cell carcinoma (ESCC)
Prognosis
As shown in human ESCC, a significant correlation exists between SLP-2 protein high expression and the depth of ESCC invasion (P=0.033) (Wang et al., 2009). Also, decreased cell growth and tumorigenesis in the antisense transfectants revealed that SLP-2 may be important in ESCC tumorigenesis (Zhang et al., 2006).
Entity name
Laryngeal squamous cell carcinoma (LSCC)
Prognosis
In addition, SLP-2 takes part in human LSCC malignant phenotype formation and development. High-level expression of SLP-2 protein could contribute to the prognostic characteristics of lymph node metastasis in human LSCC (Cao et al., 2007).
Entity name
Breast cancer
Prognosis
High-level expression of SLP-2 protein shows a worse prognosis, including increase in tumor size, progress in clinical stage, and appearance of lymph node and/or distant metastasis and is associated with decreased overall survival (P=0.011). Moreover, SLP-2 can be strongly associated with another important prognostic factor, HER-2/neu protein expression, which shows that they may act as dependent prognostic factors to indicate poor prognosis (Cao et al., 2007).
Entity name
Endometrial adenocarcinoma
Prognosis
Similarly, SLP-2 is also overexpressed in human endometrial adenocarcinoma (EAC) at both mRNA and protein level. Sense transfection of SLP-2 in EAC cell line accelerated cell growth whereas the antisense transfection reduced cell growth in vitro (Cui et al., 2007).
Entity name
Lung cancer
Prognosis
At last, SLP-2 was overexpressed in human lung cancer (Zhang et al., 2006). High-level SLP-2 expression was significantly correlated with distant metastasis, decreased overall survival and disease-free survival. SLP-2 overexpression was an independent prognostic factor in multivariate analysis using the Cox regression model (p
Entity name
Mitochondrial component
Note
SLP-2 localizes in mitochondria, affects mitochondrial membrane potential (MMP) and ATP production. Hence, SLP-2 is a mitochondrial protein and therefore, functions in energy process by MMP maintenance, and subsequently affecting cell motility, proliferation and chemosensitivity (Wang et al., 2009).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 17709317 | 2007 | High-level SLP-2 expression and HER-2/neu protein expression are associated with decreased breast cancer patient survival. | Cao W et al |
| 17306333 | 2007 | Prognostic significance of stomatin-like protein 2 overexpression in laryngeal squamous cell carcinoma: clinical, histologic, and immunohistochemistry analyses with tissue microarray. | Cao WF et al |
| 19839737 | 2010 | SLP-2 overexpression is associated with tumour distant metastasis and poor prognosis in pulmonary squamous cell carcinoma. | Chang D et al |
| 17342323 | 2007 | Stomatin-like protein 2 is overexpressed and related to cell growth in human endometrial adenocarcinoma. | Cui Z et al |
| 12239636 | 2003 | Cloning and characterization of SLP3: a novel member of the stomatin family expressed by olfactory receptor neurons. | Goldstein BJ et al |
| 11435687 | 2001 | A novel member of the STOMATIN/EPB72/mec-2 family, stomatin-like 2 (STOML2), is ubiquitously expressed and localizes to HSA chromosome 9p13.1. | Owczarek CM et al |
| 9931417 | 1998 | Molecular cloning of hSLP-1, a novel human brain-specific member of the band 7/MEC-2 family similar to Caenorhabditis elegans UNC-24. | Seidel G et al |
| 19597348 | 2009 | Downregulation of a mitochondria associated protein SLP-2 inhibits tumor cell motility, proliferation and enhances cell sensitivity to chemotherapeutic reagents. | Wang Y et al |
| 10713127 | 2000 | Identification and characterization of human SLP-2, a novel homologue of stomatin (band 7.2b) present in erythrocytes and other tissues. | Wang Y et al |
| 16533792 | 2006 | Stomatin-like protein 2 is overexpressed in cancer and involved in regulating cell growth and cell adhesion in human esophageal squamous cell carcinoma. | Zhang L et al |
Other Information
Locus ID:
NCBI: 30968
MIM: 608292
HGNC: 14559
Ensembl: ENSG00000165283
Variants:
dbSNP: 30968
ClinVar: 30968
TCGA: ENSG00000165283
COSMIC: STOML2
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000165283 | ENST00000356493 | Q9UJZ1 |
| ENSG00000165283 | ENST00000452248 | Q9UJZ1 |
| ENSG00000165283 | ENST00000488050 | F2Z2I8 |
| ENSG00000165283 | ENST00000619795 | A0A087WYB4 |
Expression (GTEx)
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38073379 | 2024 | Diagnostic and prognostic utility of TROP-2, SLP-2, and CXCL12 expression in papillary thyroid carcinoma. | 0 |
| 38214751 | 2024 | Overexpressing lipid raft protein STOML2 modulates the tumor microenvironment via NF-κB signaling in colorectal cancer. | 0 |
| 38073379 | 2024 | Diagnostic and prognostic utility of TROP-2, SLP-2, and CXCL12 expression in papillary thyroid carcinoma. | 0 |
| 38214751 | 2024 | Overexpressing lipid raft protein STOML2 modulates the tumor microenvironment via NF-κB signaling in colorectal cancer. | 0 |
| 36906621 | 2023 | STOML2 restricts mitophagy and increases chemosensitivity in pancreatic cancer through stabilizing PARL-induced PINK1 degradation. | 4 |
| 36906621 | 2023 | STOML2 restricts mitophagy and increases chemosensitivity in pancreatic cancer through stabilizing PARL-induced PINK1 degradation. | 4 |
| 35656794 | 2022 | CHCHD10 and SLP2 control the stability of the PHB complex: a key factor for motor neuron viability. | 9 |
| 35656794 | 2022 | CHCHD10 and SLP2 control the stability of the PHB complex: a key factor for motor neuron viability. | 9 |
| 33412331 | 2021 | Characterization of the interactome of c-Src within the mitochondrial matrix by proximity-dependent biotin identification. | 6 |
| 33446239 | 2021 | STOML2 potentiates metastasis of hepatocellular carcinoma by promoting PINK1-mediated mitophagy and regulates sensitivity to lenvatinib. | 42 |
| 33846782 | 2021 | Stomatin‑like protein 2 induces metastasis by regulating the expression of a rate‑limiting enzyme of the hexosamine biosynthetic pathway in pancreatic cancer. | 8 |
| 34088631 | 2021 | Stomatin-Like Protein-2: A Potential Target to Treat Mitochondrial Cardiomyopathy. | 6 |
| 33412331 | 2021 | Characterization of the interactome of c-Src within the mitochondrial matrix by proximity-dependent biotin identification. | 6 |
| 33446239 | 2021 | STOML2 potentiates metastasis of hepatocellular carcinoma by promoting PINK1-mediated mitophagy and regulates sensitivity to lenvatinib. | 42 |
| 33846782 | 2021 | Stomatin‑like protein 2 induces metastasis by regulating the expression of a rate‑limiting enzyme of the hexosamine biosynthetic pathway in pancreatic cancer. | 8 |
Citation
Wenfeng Cao ; Liyong Zhang ; Fang Ding ; Zhumei Cui ; Zhihua Liu
STOML2 (stomatin (EPB72)-like 2)
Atlas Genet Cytogenet Oncol Haematol. 2010-02-01
Online version: http://atlasgeneticsoncology.org/gene/44346/stoml2
