TNFSF15 (tumor necrosis factor (ligand) superfamily, member 15)
2009-08-01 Gui-Li Yang , Jian-Wei Qi , Zhi-Song Zhang , Lu-Yuan Li AffiliationDepartment of Pathology, University of Pittsburgh School of Medicine, Pittsburgh, PA 15213, USA (LYL); College of Pharmacy, College of Life Sciences, Nankai University, 94 Wei Jin Road, 300071 Tianjin, China (LYL, GLY, JWQ, ZSZ)
DNA/RNA

Boxes with roman numerals above represent exons and horizontal lines represent intronic sequence. The putative transcription start site is indicated by a double arrowhead. R denotes the 5 untranslated sequence unique to each respective transcript, and stippled boxe represents the common 3 untranslated region.
Description
The human VEGI gene spans about 17 kb and consists of four exons.
Transcription
The size of VEGI mRNA is approximately 6.5 kb. It is unusual for a human gene of 6.5 kb to contain only a small open reading frame of 522 nucleotides. Multiple VEGI transcripts generated by the use of cryptic splice sites and alternate exons.
Pseudogene
Not known.
Proteins
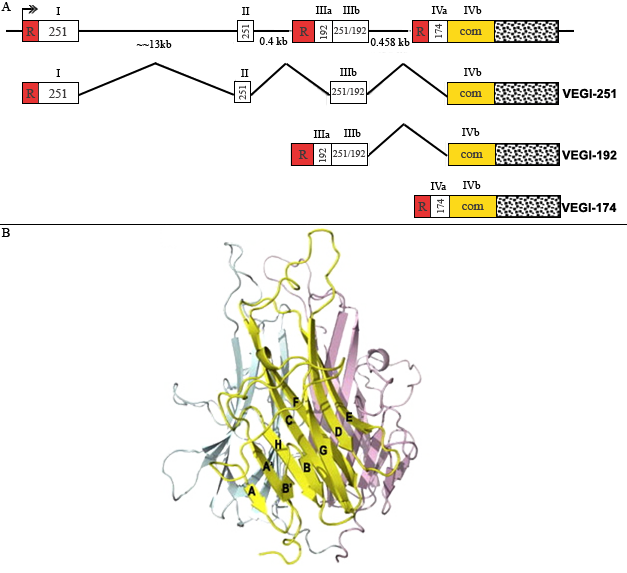
Figure A. All three isoforms.
Figure B. A ribbon diagram of the TL1A trimer. (Jin et al. BBRC 364:1, 2007).
Description
Hydrophobicity analysis of VEGI predicts a 13 amino acid hydrophobic region that follows the amino terminal segment of 12 amino acids, suggesting a structure characteristic of a type II transmembrane protein, with residues 26-174 constituting an extracellular domain analogous to domains found in other TNF family members.
VEGI isoforms exhibit a carboxyl terminal domain of 151 amino acid residues, which is encoded by part of the fourth exon, termed IVb. The initially characterized VEGI isoform, designated VEGI-174, is encoded by the fourth exon (parts IVa and IVb) alone, which includes both the putative transmembrane domain and the conserved extracellular domain. There are two additional endothelial-specific transcripts of 7.5 and 2.0 kb, which encode peptides of 251 (VEGI-251) and 192 (VEGI-192) residues, respectively. The VEGI-251 and -192 isoforms differ in their amino terminal regions, but share the conserved 151-amino acid residue carboxy terminal domain. VEGI-251 possesses a putative secretory signal peptide and its overexpression causes apoptosis of endothelial cells and inhibition of tumor growth.
VEGI isoforms exhibit a carboxyl terminal domain of 151 amino acid residues, which is encoded by part of the fourth exon, termed IVb. The initially characterized VEGI isoform, designated VEGI-174, is encoded by the fourth exon (parts IVa and IVb) alone, which includes both the putative transmembrane domain and the conserved extracellular domain. There are two additional endothelial-specific transcripts of 7.5 and 2.0 kb, which encode peptides of 251 (VEGI-251) and 192 (VEGI-192) residues, respectively. The VEGI-251 and -192 isoforms differ in their amino terminal regions, but share the conserved 151-amino acid residue carboxy terminal domain. VEGI-251 possesses a putative secretory signal peptide and its overexpression causes apoptosis of endothelial cells and inhibition of tumor growth.

Amino acid sequence alignment of three VEGI isoforms. The putative hydrophobic regions of VEGI-251 and VEGI-174 are underlined. Asterisk denotes the start of shared sequences for all three isoforms.
Expression
VEGI is specifically expressed in endothelial cells. Analysis of total RNA preparations from many cell lines and primary cell cultures by Northern blot analysis confirmed the specificity of VEGI expression, with only HUVEC and human venous endothelial cells demonstrating detectable levels of expression. Using multiple tissue Northern blots, the VEGI transcript was found in many adult human tissues, including placenta, lung, skeletal muscle, kidney, pancreas, spleen, prostate, small intestine, and colon, suggesting that the gene product may play a role in the function of a normal vasculature.The failure to detect the transcripts of this new gene in some of the human tissues probably is due to relatively small proportion of endothelial cells in these tissues. Using isoform-specific probes, we have determined that the distribution profiles of VEGI isoforms in human organs and tissues appear to be different. The 7.5 kb transcript encoding VEGI-251 was expressed at high levels in the placenta, kidney, lung and liver, whereas the 2 kb transcript corresponding to VEGI-174 was observed in liver, kidney, skeletal muscle and heart. VEGI-174 mRNA was more abundant in heart, skeletal muscle, pancreas, adrenal gland, and liver, while VEGI-251 was more abundant in fetal kidney and fetal lung. Overlapping expression of VEGI-251 and VEGI-174 mRNA was detected in prostate, salivary gland and placenta, whereas VEGI-192 mRNA was not readily detected by Northern blot. These expression patterns suggest the possibility of tissue or developmentally specific functions for VEGI isoforms. Alternatively, this expression pattern also supports the view that one VEGI isoform is the functional cytokine,while the others act in regulatory roles to modulate the activity of the active isoform. In this case, it is possible that the non-functional isoforms do not exist at the protein level. VEGI isoform expression has also been examined in cultured cells by RNase protection assay. All three known VEGI isoforms were detected in human endothelial cells, including coronary artery endothelial (HCAE), HUVE cells, and human microvascular endothelial (HMVE) cells. Very low levels are sometimes detected in adult bovine aortic endothelial (ABAE) cells. Little VEGI expression was detectable in human coronary artery smooth muscle (CASM) and mouse endothelioma bEND.3 cells. More than one isoform is detectable simultaneously, with VEGI-251 being the most abundant. The expression of this protein is inducible by TNF and IL-1 alpha, but not by gamma-interferon.
Localisation
Endothelial cells and monocytes. However, VEGI was not expressed in either B or T cells.
Function
VEGI is an endogenous inhibitor of angiogenesis produced largely by vascular endothelial cells and exerts a specific inhibitory activity on the proliferation of endothelial cells. VEGI enforces growth arrest of endothelial cells in G0 and early G1 phases of the cell cycle but induces apoptosis in proliferating endothelial cells. The MAPKs p38 and jun N-terminal kinase (JNK) are required for VEGI-mediated endothelial inhibition. Engineered overexpression of secreted VEGI by cancer cells or systemic administration of recombinant VEGI to tumor-bearing mice inhibits tumor growth in numerous tumor models. Recent studies show that VEGI helps modulate the immune system by activating T cells and stimulating dendritic cell maturation, suggesting that VEGI is directly involved in modulating the interaction between the endothelium and the immune system. Recombinant VEGI has an inhibitory activity on mouse bone marrow-derived EPCs in culture, preventing their differentiation toward endothelial cells.
Interaction of TL1A with DR3 promotes T cell expansion during an immune response (Migone et al., 2002).
Interaction of TL1A with DR3 promotes T cell expansion during an immune response (Migone et al., 2002).
Homology
VEGI exhibits 20-30% sequence homology to human TNF-alpha, TNF-beta, and the Fas ligand, similar to that among other TNF family members.
Implicated in
Entity name
Colon carcinoma
Note
Local production of a secreted form of VEGI via gene transfer caused complete suppression of the growth of MC-38 murine colon cancers in syngeneic C57BL/6mice. Histological examination showed marked reduction of vascularization in MC-38 tumors that expressed soluble but not membrane-bound VEGI or were transfected with control vector. The conditioned media from soluble VEGI-expressing cells showed marked inhibitory effect on in vitro proliferation of adult bovine aortic endothelial cells.
Entity name
Breast cancer
Note
The anticancer potential of VEGI was examined in a breast cancer xenograft tumor model in which the cancer cells were co-injected with Chinese hamster ovary cells overexpressing a secreted form of the protein. The co-injection resulted in potent inhibition of xenograft tumor growth. Our findings are consistent with the view that VEGI is an endothelial cell-specific negative regulator of angiogenesis.
Entity name
Mucosal vaccine adjuvant
Note
Kayamuro et al., (2009) reported that TL1A induced the strongest immune response and augmented OVA-specific IgG and IgA responses in serum and mucosal compartments, respectively. The OVA-specific immune response of TL1A was characterized by high levels of serum IgG1 and increased production of IL-4 and IL-5 from splenocytes of immunized mice, suggesting that TL1A might induce Th2-type responses. These findings indicate that TL1A has the most potential as a mucosal adjuvant among the TNFS cytokines.
Entity name
Inflammatory bowel disease
Note
Bamias et al., (2003) provided evidence that the novel cytokine TL1A may play an important role in a Th1-mediated disease such as Crohns disease. Takedatsu et al., (2008) revealed that TL1A is an important modulator in the development of chronic mucosal inflammation by enhancing T(H)1 and T(H)17 effector functions. The central role of TL1A represents an attractive, novel therapeutic target for the treatment of Crohns disease patients.
Entity name
Inflammatory arthritis
Note
Bull et al., (2008) demonstrated that the DR3-TL1A pathway regulates joint destruction in two murine models of arthritis and represents a potential novel target for therapeutic intervention in inflammatory joint disease. Bamias et al., (2008) concluded that TL1A may serve as an inflammatory marker in rheumatoid arthritis. Interactions between TL1A and its receptors may be important in the pathogenesis of rheumatoid arthritis.
Entity name
Renal inflammation and injury
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 18287561 | 2008 | TL1A both promotes and protects from renal inflammation and injury. | Al-Lamki RS et al |
| 15308462 | 2004 | Endothelial progenitor cells for postnatal vasculogenesis. | Asahara T et al |
| 14568967 | 2003 | Expression, localization, and functional activity of TL1A, a novel Th1-polarizing cytokine in inflammatory bowel disease. | Bamias G et al |
| 16698931 | 2006 | Role of TL1A and its receptor DR3 in two models of chronic murine ileitis. | Bamias G et al |
| 18757243 | 2008 | Circulating levels of TNF-like cytokine 1A (TL1A) and its decoy receptor 3 (DcR3) in rheumatoid arthritis. | Bamias G et al |
| 18824582 | 2008 | The Death Receptor 3-TNF-like protein 1A pathway drives adverse bone pathology in inflammatory arthritis. | Bull MJ et al |
| 18769489 | 2008 | Preparation and characterization of a novel chimeric protein VEGI-CTT in Escherichia coli. | Cai J et al |
| 11923219 | 2002 | A novel secreted splice variant of vascular endothelial cell growth inhibitor. | Chew LJ et al |
| 18187653 | 2008 | Endothelial progenitor cells control the angiogenic switch in mouse lung metastasis. | Gao D et al |
| 16061878 | 2005 | VEGI-192, a new isoform of TNFSF15, specifically eliminates tumor vascular endothelial cells and suppresses tumor growth. | Hou W et al |
| 17905596 | 2007 | Purification and crystallization of recombinant human TNF-like ligand TL1A. | Jin T et al |
| 19406102 | 2009 | TNF superfamily member, TL1A, is a potential mucosal vaccine adjuvant. | Kayamuro H et al |
| 16517446 | 2006 | Vascular endothelial growth inhibitor (VEGI), an endogenous negative regulator of angiogenesis. | Metheny-Barlow LJ et al |
| 11911831 | 2002 | TL1A is a TNF-like ligand for DR3 and TR6/DcR3 and functions as a T cell costimulator. | Migone TS et al |
| 18598698 | 2008 | TL1A (TNFSF15) regulates the development of chronic colitis by modulating both T-helper 1 and T-helper 17 activation. | Takedatsu H et al |
| 9434163 | 1997 | Characterization of a novel TNF-like ligand and recently described TNF ligand and TNF receptor superfamily genes and their constitutive and inducible expression in hematopoietic and non-hematopoietic cells. | Tan KB et al |
| 17785811 | 2007 | The endothelial cell-produced antiangiogenic cytokine vascular endothelial growth inhibitor induces dendritic cell maturation. | Tian F et al |
| 19329781 | 2009 | Inhibition of endothelial progenitor cell differentiation by VEGI. | Tian F et al |
| 11739281 | 2001 | Modulation of endothelial cell growth arrest and apoptosis by vascular endothelial growth inhibitor. | Yu J et al |
| 9880523 | 1999 | TL1, a novel tumor necrosis factor-like cytokine, induces apoptosis in endothelial cells. Involvement of activation of stress protein kinases (stress-activated protein kinase and p38 mitogen-activated protein kinase) and caspase-3-like protease. | Yue TL et al |
| 9872942 | 1999 | VEGI, a novel cytokine of the tumor necrosis factor family, is an angiogenesis inhibitor that suppresses the growth of colon carcinomas in vivo. | Zhai Y et al |
| 10360832 | 1999 | Inhibition of angiogenesis and breast cancer xenograft tumor growth by VEGI, a novel cytokine of the TNF superfamily. | Zhai Y et al |
| 12837752 | 2003 | Structure and inhibitory effects on angiogenesis and tumor development of a new vascular endothelial growth inhibitor. | Zilberberg L et al |
Other Information
Locus ID:
NCBI: 9966
MIM: 604052
HGNC: 11931
Ensembl: ENSG00000181634
Variants:
dbSNP: 9966
ClinVar: 9966
TCGA: ENSG00000181634
COSMIC: TNFSF15
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000181634 | ENST00000374044 | X6R8I9 |
| ENSG00000181634 | ENST00000374045 | O95150 |
| ENSG00000181634 | ENST00000374045 | A0A0U5JA19 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37392242 | 2024 | The Relation of VEGFA, VEGFR2, VEGI, and HIF1A Genetic Variants and Their Serum Protein Levels with Breast Cancer in Egyptian Patients. | 0 |
| 38597952 | 2024 | TL1A is an epithelial alarmin that cooperates with IL-33 for initiation of allergic airway inflammation. | 0 |
| 38866875 | 2024 | MicroRNA-141-regulated KLK10 and TNFSF-15 gene expression in hepatoblastoma cells as a novel mechanism in liver carcinogenesis. | 0 |
| 37392242 | 2024 | The Relation of VEGFA, VEGFR2, VEGI, and HIF1A Genetic Variants and Their Serum Protein Levels with Breast Cancer in Egyptian Patients. | 0 |
| 38597952 | 2024 | TL1A is an epithelial alarmin that cooperates with IL-33 for initiation of allergic airway inflammation. | 0 |
| 38866875 | 2024 | MicroRNA-141-regulated KLK10 and TNFSF-15 gene expression in hepatoblastoma cells as a novel mechanism in liver carcinogenesis. | 0 |
| 36768135 | 2023 | Increased Serum Levels of Tumor Necrosis Factor-like Ligand 1A in Atopic Dermatitis. | 0 |
| 37156999 | 2023 | Genetic architecture of the inflammatory bowel diseases across East Asian and European ancestries. | 28 |
| 36768135 | 2023 | Increased Serum Levels of Tumor Necrosis Factor-like Ligand 1A in Atopic Dermatitis. | 0 |
| 37156999 | 2023 | Genetic architecture of the inflammatory bowel diseases across East Asian and European ancestries. | 28 |
| 34022066 | 2022 | Genome-wide association study identifies TNFSF15 associated with childhood asthma. | 6 |
| 34232416 | 2022 | The relationship between TNF-like protein 1A and coronary artery aneurysms in children with Kawasaki disease. | 1 |
| 34974051 | 2022 | Nonylphenol regulates TL1A through the AhR/HDAC2/HNF4α pathway in endothelial cells to promote the angiogenesis of colorectal cancer. | 1 |
| 36226563 | 2022 | Role of TNFSF15 variants in oral cancer development and clinicopathologic characteristics. | 5 |
| 36429070 | 2022 | Immune Checkpoint Inhibitor (ICI) Genes and Aging in Clear Cell Renal Cell Carcinoma (ccRCC): Clinical and Genomic Study. | 4 |
Citation
Gui-Li Yang ; Jian-Wei Qi ; Zhi-Song Zhang ; Lu-Yuan Li
TNFSF15 (tumor necrosis factor (ligand) superfamily, member 15)
Atlas Genet Cytogenet Oncol Haematol. 2009-08-01
Online version: http://atlasgeneticsoncology.org/gene/42638/tnfsf15
