ING2 (inhibitor of growth family, member 2)
2009-05-01 Susanne Jennek , Aria Baniahmad AffiliationInstitute of Human Genetics, Anthropology, Jena University Hospital, Kollegiengasse 10, 07743 Jena, Germany
DNA/RNA
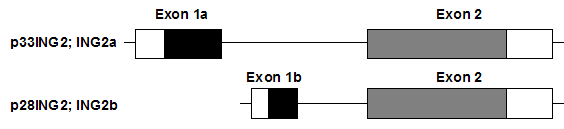
Gene structure of ING2a and ING2b (modified according to Unoki et al., 2008).
Description
The two isoforms share exon 2 but have different exon 1. The exon 1a of ING2a encodes 58 amino acids. Exon 1b of ING2b encodes 18 amino acids. There is no significant homology between the N-terminal part of ING2b and the N-terminal regions of ING1 isoforms (Unoki et al., 2008).
Transcription
The promoter region of ING2a possesses two p53 binding sites in contrast to the promoter of ING2b. Binding of p53 to these sites suppresses the ING2a expression. The promoter region of ING2b harbors a HSF1 and HSF2 binding site, a c-Rel, a SP1 and five MZF1 binding sites (Unoki et al., 2008).
Proteins
Description
Expression
ING2 is widely expressed in normal tissues (Shimada et al., 1998).
Localisation
ING2 is predominantly localized in the nucleus to chromatin and the nuclear matrix (Gozani et al., 2003). Through reduced levels of phosphoinositide PtdIns5P, ING2 might be released from chromatin and translocates partially to the cytoplasm (Gozani et al., 2003).
Function
The tumor suppressor protein ING2 has been described to regulate cell cycle, cellular senescence and gene regulation including chromatin level. In response to DNA damage ING2 enhances the acetylation of p53, which negatively regulates cell proliferation (Nagashima et al., 2001). In addition the level of ING2 expression directly regulates the onset of replicative senescence through the induction of p300-dependent acetylation of p53 (Pedeux et al., 2005). ING2 also induces the global histone H4 acetylation and chromatin relaxation and thereby enhances the nucleotide excision repair (Wang et al., 2006). Furthermore, ING2 is also a part of two related mSin3/HDAC1/HDAC2 corepressor complexes (Doyon et al., 2006). Further, chromatin association of ING2 is linked to ING2 ability to bind to trimethylated K4 of histone 3 (H3K4me3) via its plant homeodomain (PHD) region (Shi et al., 2006). p33ING2 is also associated with histone methyltranferase (HMT-) activity in vitro and in vivo, methylating specifically histone H3 and histone H1 (Goeman et al., 2008). In addition, ING2, as a transcriptional repressor, directly interacts with the corepressor Alien and enhances the Alien-mediated gene silencing (Fegers et al., 2007). Furthermore, ING2 interacts with the Smad-interaction transcriptional modulator SnoN mediating TGF-beta-induced Smad-dependent transcription and cellular responses (Sarker et al., 2008). The activity of ING2, as a nuclear phosphatidylinositol receptor can be modulated by phosphoinositides (Gozani et al., 2003).
Mutations
Note
So far natural occurring point mutations of ING2 in association with cancer were not yet described. However, loss of heterozygosity (LOH) and aberrant ING2-mRNA levels were associated with cancer.
Implicated in
Entity name
Colon cancer
Oncogenesis
ING2 expression level in human colon tumors is significantly higher than in normal colon tissue. In conclusion, ING2 might be involved in colon cancers (Shimada et al., 1998).
Entity name
Hepatocellular carcinoma (HCC)
Oncogenesis
ING2 transcription and post-transcription level is downregulated in the majority of HCC tumors compared with non-tumors liver tissue. Furthermore, ING2 expression level is reduced in 44 of 84 (52.4%) HCC cases and the ING2 expression level correlated with tumor size, histopathologic classification and serum AFP (Zhang et al., 2008). It is also shown that HCC patients with reduced ING2 expression have a significantly increased risk exhibiting a shorter survival time. In conclusion, ING2 may be involved in the progression of HCC (Zhang et al., 2008).
Entity name
Head and neck squamous cell carcinoma (HNSCC)
Oncogenesis
There is a loss of heterozygosity (LOH) in the region 4q32 in the long arm of the chromosome 4 in 20% of the cases (Borkovsky et. al., 2008). This region includes ING2 and SAP30 genes that are parts of the two related mSin3/ HDAC1/2 corepressor complexes. LOH on region 4q35.1 was detected in 30 (54.6%) out of 55 informative cases (Borkovsky et. al., 2008). High LOH frequency is associated with advanced tumor stages, therefore, ING2 LOH is likely to be a late event in HNSCC.
Entity name
Lung cancer
Oncogenesis
Although, there are no ING2 mutations identified in 30 human lung cancer cell lines and 31 primary lung cancer tumors. The ING2 mRNA expression is reduced in 6 out of 7 lung cancer cell lines (Okano et al., 2006).
Entity name
Cutaneous melanomas
Oncogenesis
The nuclear expression level of ING2 is significantly reduced in human melanomas compared to dysplastic nevi. It is suggested that reduced ING2 expression may be involved in the initiation of melanoma (Lu et al., 2006; Ythier et al., 2008).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 18998165 | 2009 | Frequent deletion of ING2 locus at 4q35.1 associates with advanced tumor stage in head and neck squamous cell carcinoma. | Borkosky SS et al |
| 15526165 | 2004 | Biological functions of the ING family tumor suppressors. | Campos EI et al |
| 16387653 | 2006 | ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. | Doyon Y et al |
| 17929852 | 2007 | The tumor suppressors p33ING1 and p33ING2 interact with alien in vivo and enhance alien-mediated gene silencing. | Fegers I et al |
| 18513492 | 2008 | ING2 recruits histone methyltransferase activity with methylation site specificity distinct from histone H3 lysines 4 and 9. | Goeman F et al |
| 12859901 | 2003 | The PHD finger of the chromatin-associated protein ING2 functions as a nuclear phosphoinositide receptor. | Gozani O et al |
| 16755297 | 2006 | Nuclear ING2 expression is reduced in human cutaneous melanomas. | Lu F et al |
| 11481424 | 2001 | DNA damage-inducible gene p33ING2 negatively regulates cell proliferation through acetylation of p53. | Nagashima M et al |
| 16465410 | 2006 | Alterations in novel candidate tumor suppressor genes, ING1 and ING2 in human lung cancer. | Okano T et al |
| 16024799 | 2005 | ING2 regulates the onset of replicative senescence by induction of p300-dependent p53 acetylation. | Pedeux R et al |
| 16644882 | 2006 | Molecular biology of squamous cell carcinoma of the head and neck. | Perez-Ordoñez B et al |
| 18334480 | 2008 | ING2 as a novel mediator of transforming growth factor-beta-dependent responses in epithelial cells. | Sarker KP et al |
| 16728974 | 2006 | ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. | Shi X et al |
| 10072587 | 1998 | Cloning of a novel gene (ING1L) homologous to ING1, a candidate tumor suppressor. | Shimada Y et al |
| 17949986 | 2007 | After a decade of study-ING, a PHD for a versatile family of proteins. | Soliman MA et al |
| 18951897 | 2008 | A novel ING2 isoform, ING2b, synergizes with ING2a to prevent cell cycle arrest and apoptosis. | Unoki M et al |
| 16488987 | 2006 | The novel tumor suppressor p33ING2 enhances nucleotide excision repair via inducement of histone H4 acetylation and chromatin relaxation. | Wang J et al |
| 18636562 | 2008 | The new tumor suppressor genes ING: genomic structure and status in cancer. | Ythier D et al |
| 18160212 | 2008 | Decreased expression of ING2 gene and its clinicopathological significance in hepatocellular carcinoma. | Zhang HK et al |
Other Information
Locus ID:
NCBI: 3622
MIM: 604215
HGNC: 6063
Ensembl: ENSG00000168556
Variants:
dbSNP: 3622
ClinVar: 3622
TCGA: ENSG00000168556
COSMIC: ING2
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000168556 | ENST00000302327 | Q9H160 |
| ENSG00000168556 | ENST00000412117 | C9J4X5 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 34017078 | 2021 | ING2 tumor suppressive protein translocates into mitochondria and is involved in cellular metabolism homeostasis. | 4 |
| 34017078 | 2021 | ING2 tumor suppressive protein translocates into mitochondria and is involved in cellular metabolism homeostasis. | 4 |
| 32015217 | 2020 | Integrated analysis of DNA methylation and mRNA expression profiles to identify key genes involved in the regrowth of clinically non-functioning pituitary adenoma. | 8 |
| 32424454 | 2020 | Inhibitor of growth 2 regulates the high glucose-induced cell cycle arrest and epithelial-to-mesenchymal transition in renal proximal tubular cells. | 8 |
| 32015217 | 2020 | Integrated analysis of DNA methylation and mRNA expression profiles to identify key genes involved in the regrowth of clinically non-functioning pituitary adenoma. | 8 |
| 32424454 | 2020 | Inhibitor of growth 2 regulates the high glucose-induced cell cycle arrest and epithelial-to-mesenchymal transition in renal proximal tubular cells. | 8 |
| 31180538 | 2019 | miR‑92a contributes to cell proliferation, apoptosis and doxorubicin chemosensitivity in gastric carcinoma cells. | 8 |
| 31180538 | 2019 | miR‑92a contributes to cell proliferation, apoptosis and doxorubicin chemosensitivity in gastric carcinoma cells. | 8 |
| 26717876 | 2016 | Expression profiles of inhibitor of growth protein 2 in normal and cancer tissues: An immunohistochemical screening analysis. | 3 |
| 27305909 | 2016 | A novel crosstalk between the tumor suppressors ING1 and ING2 regulates androgen receptor signaling. | 13 |
| 26717876 | 2016 | Expression profiles of inhibitor of growth protein 2 in normal and cancer tissues: An immunohistochemical screening analysis. | 3 |
| 27305909 | 2016 | A novel crosstalk between the tumor suppressors ING1 and ING2 regulates androgen receptor signaling. | 13 |
| 25613071 | 2015 | A novel tumor suppressor gene in basal cell carcinoma: inhibition of growth factor-2. | 1 |
| 25613071 | 2015 | A novel tumor suppressor gene in basal cell carcinoma: inhibition of growth factor-2. | 1 |
| 24712846 | 2014 | Decreased expression of ING2 gene and its clinicopathological significance in Chinese NSCLC patients. | 3 |
Citation
Susanne Jennek ; Aria Baniahmad
ING2 (inhibitor of growth family, member 2)
Atlas Genet Cytogenet Oncol Haematol. 2009-05-01
Online version: http://atlasgeneticsoncology.org/gene/40975/ing2
