STAT2 (Signal Transducer and Activator of Transcription 2)
2015-05-01 Ming Li AffiliationDepartment of Immunology, Xiangya Medical College, Basic Medical College, Central South University, Changsha 410078, Hunan, P. R. China
Identity
HGNC
LOCATION
12q13.3
LOCUSID
ALIAS
IMD44,ISGF-3,P113,PTORCH3,STAT113
FUSION GENES
Abstract
Review on STAT2, with data on DNA, on the protein encoded, and where the gene is implicated.
DNA/RNA
Description
24 exons spanning 18657 bp
Transcription
There are two major transcripts. In transcript variant 1, mRNA is 4576 bp. Transcript variant 2 uses an alternate in-frame splice site in exon 3; as a result, it lacks an internal 12bp insertion compared to transcript variant 1. Another spliced form is generated by reading through the intron between exons 20 and 21, which correspond to the region encoding the SH2 domain. The spliced form contains a stop codon in the SH2 domain, giving rise to a short form of STAT2 when the mRNAs are translated. The putative translated proteins lack half of the SH2 domain, the tyrosine phosphorylation site required for dimerization and DNA binding, and the C-terminal activation domain.
Proteins
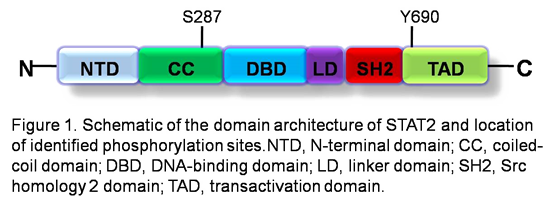
Description
There are two major isoforms of STAT2. The long form is known as isoform 1 and is a 851 amino acid protein (113KDa on gels). Isoform 2 lacks an internal four amino acid segment compared to isoform 1.
The STAT2 gene product contains 6 domains: an N-terminal domain (NTD), a coiled-coil domain (CC), a DNA-binding domain (DBD), a linker domain (LD), a Src homology-2 domain (SH2), and a transactivation domain (TAD) (figure 1). NTD is required for tyrosine phosphorylation of STAT2 in response to type I IFNs, binding of STAT2 to IFN receptors and cooperative binding of STAT1:STAT2 heterodimers or ISGF3 to promoters that contain tandem GAS or ISRE, respectively. CC mediates protein interactions and is the domain IRF9 binds. DBD does not bind DNA as a part of ISGF3. DBD contains a bipartite nuclear localization signal (NLS) when ISGF3 forms. No function for LD is known. SH2 serves 2 main functions: binding to a phosphorylated IFN receptor, thereby making STAT2 available for Tyk2-mediated tyrosine phosphorylation; and binding to tyrosine phosphorylated STAT1 to form an active heterodimer. The TAD is essential for recruitment of transcription regulators. TAD also contains the nuclear export signal (NES). Nucleocytoplasmic shuttling of STAT2 is attributed to the constitutive binding of STAT2 to the NLS-containing IRF9 to transport STAT2 into the nucleus, while the STAT2 NES exports STAT2 back to the cytosol (Steen and Gamero, 2012).
Interferons (IFNs) activate the Janus kinase (JAK)/STAT pathway by binding to their corresponding receptor complex. Jak1 and TYK2 are pre-associated with type I and type III IFN receptors, and phosphorylate specific tyrosine residues within the receptor chain, which serve as docking sites for the recruitment of STATs. JAKs phosphorylate a conserved tyrosine residue situated in the C-terminal region of STAT2 (Y690) and STAT1 , thus allowing STAT1 and STAT2 to interact via reciprocal SH2-phosphotyrosyl interactions. Formation of the interferon-stimulated gene factor 3 (ISGF3) complex takes place when activated STAT1:STAT2 heterodimers are released from receptor chains to bind the DNA-binding adaptor protein, IRF9 (p48/ISGF3G). ISGF3 translocates to the nucleus and binds the DNA containing an IFN-stimulated response element (ISRE) by STAT1 and IRF9 to activate gene transcription of IFN-stimulated genes (ISGs). In addition, STAT2 can form heterodimers individually with either STAT1 or IRF9. Each of these complexes will bind IRSE or IFN-gamma activation sequences (GAS) to activate gene transcription of ISGs. Serine 287 phosphorylation can negatively regulate the biological activities of type I IFNs (Steen et al., 2013). IFNs are the only cytokines known to date that can activate STAT2.
The STAT2 gene product contains 6 domains: an N-terminal domain (NTD), a coiled-coil domain (CC), a DNA-binding domain (DBD), a linker domain (LD), a Src homology-2 domain (SH2), and a transactivation domain (TAD) (figure 1). NTD is required for tyrosine phosphorylation of STAT2 in response to type I IFNs, binding of STAT2 to IFN receptors and cooperative binding of STAT1:STAT2 heterodimers or ISGF3 to promoters that contain tandem GAS or ISRE, respectively. CC mediates protein interactions and is the domain IRF9 binds. DBD does not bind DNA as a part of ISGF3. DBD contains a bipartite nuclear localization signal (NLS) when ISGF3 forms. No function for LD is known. SH2 serves 2 main functions: binding to a phosphorylated IFN receptor, thereby making STAT2 available for Tyk2-mediated tyrosine phosphorylation; and binding to tyrosine phosphorylated STAT1 to form an active heterodimer. The TAD is essential for recruitment of transcription regulators. TAD also contains the nuclear export signal (NES). Nucleocytoplasmic shuttling of STAT2 is attributed to the constitutive binding of STAT2 to the NLS-containing IRF9 to transport STAT2 into the nucleus, while the STAT2 NES exports STAT2 back to the cytosol (Steen and Gamero, 2012).
Interferons (IFNs) activate the Janus kinase (JAK)/STAT pathway by binding to their corresponding receptor complex. Jak1 and TYK2 are pre-associated with type I and type III IFN receptors, and phosphorylate specific tyrosine residues within the receptor chain, which serve as docking sites for the recruitment of STATs. JAKs phosphorylate a conserved tyrosine residue situated in the C-terminal region of STAT2 (Y690) and STAT1 , thus allowing STAT1 and STAT2 to interact via reciprocal SH2-phosphotyrosyl interactions. Formation of the interferon-stimulated gene factor 3 (ISGF3) complex takes place when activated STAT1:STAT2 heterodimers are released from receptor chains to bind the DNA-binding adaptor protein, IRF9 (p48/ISGF3G). ISGF3 translocates to the nucleus and binds the DNA containing an IFN-stimulated response element (ISRE) by STAT1 and IRF9 to activate gene transcription of IFN-stimulated genes (ISGs). In addition, STAT2 can form heterodimers individually with either STAT1 or IRF9. Each of these complexes will bind IRSE or IFN-gamma activation sequences (GAS) to activate gene transcription of ISGs. Serine 287 phosphorylation can negatively regulate the biological activities of type I IFNs (Steen et al., 2013). IFNs are the only cytokines known to date that can activate STAT2.
Expression
STAT2 is ubiquitously expressed in most cell types.
Localisation
STAT2 predominantly resides in the cytoplasm. Nucleocytoplasmic shuttling occurs in the absence of IFN stimulation, but translocates to the nucleus upon tyrosine phosphorylation when stimulated by IFNs (Reich, 2013).
Function
Transcription factor. STAT2 mediates the transcription of numerous IFN-induced genes involved in linking adaptive and innate immunity and exerting antiviral, antiproliferative, apoptotic, and antitumor effects.
Homology
Shares homology with the other 6 mammalian STAT genes: STAT1, STAT3, STAT4, STAT5A, STAT5B, STAT6. Human STAT2 is relatively well conserved with macaque and chimpanzee. Human and murine STAT2 are highly homologous (76% identity over the first 712 amino acids).
Mutations
Note
Two mutations in intron 4 (Hambleton,et al., 2013) as well as the intron between exon 20 and 21 were found to prevent correct splicing of STAT2 (Sugiyama et al., 1996).
Implicated in
Note
To evade the antiviral protective effects of IFNs, certain viruses have developed strategies to impair the IFN signaling pathway by specifically targeting STAT2. The strategies are to reduce STAT2 levels, to sequester STAT2 in the cytoplasm, or to prevent its tyrosine phosphorylation.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 12114405 | 2002 | Suppression of type I interferon signaling proteins is an early event in squamous skin carcinogenesis. | Clifford JL et al |
| 18999875 | 2008 | Expression of STATs and their inhibitors SOCS and PIAS in brain tumors. In vitro and in vivo study. | Ehrmann J et al |
| 20233899 | 2010 | STAT2 contributes to promotion of colorectal and skin carcinogenesis. | Gamero AM et al |
| 23391734 | 2013 | STAT2 deficiency and susceptibility to viral illness in humans. | Hambleton S et al |
| 26061044 | 2015 | The Tumor Suppressive Effects of HPP1 Are Mediated Through JAK-STAT-Interferon Signaling Pathways. | Hernandez JM et al |
| 16698995 | 2006 | Differentially regulated interferon response determines the outcome of Newcastle disease virus infection in normal and tumor cell lines. | Krishnamurthy S et al |
| 16523488 | 2006 | cDNA-array profiling of melanomas and paired melanocyte cultures. | Mischiati C et al |
| 11912448 | 2002 | Selective activation of members of the signal transducers and activators of transcription family in prostate carcinoma. | Ni Z et al |
| 24470978 | 2013 | STATs get their move on. | Reich NC et al |
| 23139419 | 2013 | Identification of STAT2 serine 287 as a novel regulatory phosphorylation site in type I interferon-induced cellular responses. | Steen HC et al |
| 8601453 | 1996 | Identification of alternative splicing form of Stat2. | Sugiyama T et al |
| 24895110 | 2015 | Host STAT2/type I interferon axis controls tumor growth. | Yue C et al |
Other Information
Locus ID:
NCBI: 6773
MIM: 600556
HGNC: 11363
Ensembl: ENSG00000170581
Variants:
dbSNP: 6773
ClinVar: 6773
TCGA: ENSG00000170581
COSMIC: STAT2
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
PharmGKB
| Entity ID | Name | Type | Evidence | Association | PK | PD | PMIDs |
|---|---|---|---|---|---|---|---|
| PA134885765 | SOCS3 | Gene | Pathway | associated | 26111151 | ||
| PA134992438 | CAMK1D | Gene | Pathway | associated | |||
| PA24684 | AKT1 | Gene | Pathway | associated | |||
| PA24685 | AKT2 | Gene | Pathway | associated | |||
| PA24686 | AKT3 | Gene | Pathway | associated | |||
| PA248 | NFKB1 | Gene | Pathway | associated | |||
| PA26048 | CAMK1 | Gene | Pathway | associated | |||
| PA26049 | CAMK1G | Gene | Pathway | associated | |||
| PA27749 | ELK1 | Gene | Pathway | associated | |||
| PA28212 | FOS | Gene | Pathway | associated | |||
| PA283 | MAPK8 | Gene | Pathway | associated | |||
| PA296 | RELA | Gene | Pathway | associated | |||
| PA29988 | JAK1 | Gene | Pathway | associated | |||
| PA30006 | JUN | Gene | Pathway | associated | |||
| PA30616 | MAPK1 | Gene | Pathway | associated | |||
| PA30622 | MAPK3 | Gene | Pathway | associated | |||
| PA31353 | MYC | Gene | Pathway | associated | |||
| PA31600 | NFKB2 | Gene | Pathway | associated | |||
| PA337 | STAT3 | Gene | Pathway | associated | |||
| PA33759 | PRKCA | Gene | Pathway | associated | |||
| PA33761 | PRKCB | Gene | Pathway | associated | |||
| PA33766 | PRKCG | Gene | Pathway | associated | |||
| PA338 | STAT5A | Gene | Pathway | associated | |||
| PA36042 | SP1 | Gene | Pathway | associated | |||
| PA36183 | STAT1 | Gene | Pathway | associated | |||
| PA36185 | STAT4 | Gene | Pathway | associated | |||
| PA36186 | STAT5B | Gene | Pathway | associated | |||
| PA37242 | USP18 | Gene | Pathway | associated | 26111151 | ||
| PA90 | CAMK2A | Gene | Pathway | associated | |||
| PA91 | CAMK2B | Gene | Pathway | associated | |||
| PA92 | CAMK2D | Gene | Pathway | associated | |||
| PA93 | CAMK2G | Gene | Pathway | associated |
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38514903 | 2024 | The lack of either IRF9, or STAT2, has surprisingly little effect on human natural killer cell development and function. | 0 |
| 38879005 | 2024 | African swine fever virus pB475L evades host antiviral innate immunity via targeting STAT2 to inhibit IFN-I signaling. | 0 |
| 38514903 | 2024 | The lack of either IRF9, or STAT2, has surprisingly little effect on human natural killer cell development and function. | 0 |
| 38879005 | 2024 | African swine fever virus pB475L evades host antiviral innate immunity via targeting STAT2 to inhibit IFN-I signaling. | 0 |
| 36328987 | 2023 | circCAPRIN1 interacts with STAT2 to promote tumor progression and lipid synthesis via upregulating ACC1 expression in colorectal cancer. | 13 |
| 36976641 | 2023 | Human inherited complete STAT2 deficiency underlies inflammatory viral diseases. | 6 |
| 37115061 | 2023 | STAT2 act a prognostic biomarker and associated with immune infiltration in kidney renal clear cell carcinoma. | 0 |
| 37526179 | 2023 | Investigation of the impact of AXL, TLR3, and STAT2 in congenital Zika syndrome through genetic polymorphisms and protein-protein interaction network analyses. | 0 |
| 38139463 | 2023 | ISGF3 and STAT2/IRF9 Control Basal and IFN-Induced Transcription through Genome-Wide Binding of Phosphorylated and Unphosphorylated Complexes to Common ISRE-Containing ISGs. | 1 |
| 36328987 | 2023 | circCAPRIN1 interacts with STAT2 to promote tumor progression and lipid synthesis via upregulating ACC1 expression in colorectal cancer. | 13 |
| 36976641 | 2023 | Human inherited complete STAT2 deficiency underlies inflammatory viral diseases. | 6 |
| 37115061 | 2023 | STAT2 act a prognostic biomarker and associated with immune infiltration in kidney renal clear cell carcinoma. | 0 |
| 37526179 | 2023 | Investigation of the impact of AXL, TLR3, and STAT2 in congenital Zika syndrome through genetic polymorphisms and protein-protein interaction network analyses. | 0 |
| 38139463 | 2023 | ISGF3 and STAT2/IRF9 Control Basal and IFN-Induced Transcription through Genome-Wide Binding of Phosphorylated and Unphosphorylated Complexes to Common ISRE-Containing ISGs. | 1 |
| 34643427 | 2022 | The Human STAT2 Coiled-Coil Domain Contains a Degron for Zika Virus Interferon Evasion. | 8 |
Citation
Ming Li
STAT2 (Signal Transducer and Activator of Transcription 2)
Atlas Genet Cytogenet Oncol Haematol. 2015-05-01
Online version: http://atlasgeneticsoncology.org/gene/42429/stat2-(signal-transducer-and-activator-of-transcription-2)
