ASPM (asp (abnormal spindle) homolog, microcephaly associated (Drosophila))
2009-12-01 Shian-Yang Peng , Shih-Yeh Lin , Hey-Chi Hsu AffiliationIdentity
HGNC
LOCATION
1q31.3
LOCUSID
ALIAS
ASP,Calmbp1,MCPH5
FUSION GENES
DNA/RNA
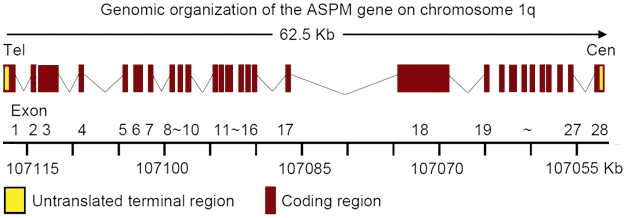
Genomic diagram of ASPM gene. Exons are represented by boxes on the diagram.
Description
ASPM, maps to NC_000001.10 in human genome, contains 28 exons with a 10,434 base pairs ORF (NCBI GenBank accession number AF509326), spanning a region of 62,568 base pairs at chromosome 1q31.
Transcription
ASPM encodes a 10906 bps mRNA, (NM_018136.4), in a telomeric to centromeric orientation and the coding region is from 258 bp to 10961bp (10434 bp). 5 part of exon 1 and 3 part of exon 28 are non-coding. The 28 exons of ASPM mRNA are 554, 144, 1480, 105, 147, 246, 68, 142, 131, 176, 146, 86, 222, 208, 143, 129, 195, 4755, 167, 97, 210, 150, 192, 193, 155, 177, 170, 300 base pairs, respectively.
Pseudogene
None.
Proteins
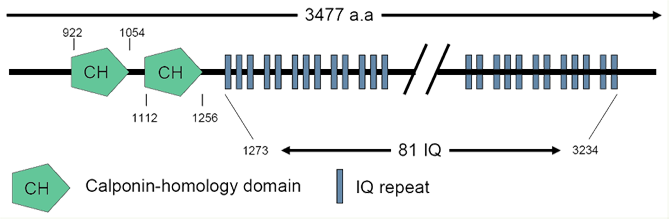
ASPM protein contains 2 calponin homology (CH) domains and 81 IQ repeats domain. Total 81 distinct IQ motifs were found at position 1273 to 3243. Numbers indicate the amino acid numbers.
Description
The 10434 bps ORF of ASPM mRNA translates a 3477 amino acid protein with a calculated molecular weight of 409.8 kDa. The main ASPM isoform protein, from N-terminal, begins with a microtubule binding domain, followed by two calponin homology (CH) domains, which are possibly responsible for transportation of the ASPM protein to the spindle poles. The third part is IQ (I for isoleucine, Q for glutamine) repeats region which is the calmodulin binding domains. The numbers of IQ repeats are identified up to 81, at position 1273 to 3243. The C-terminal of ASPM is an armadillo-like domain of unknown function.
Expression
ASPM transcripts were detected in a variety of human embryo tissues (brain, bladder, colon,heart, liver, lung, skeletal muscle, skin, spleen and stomach), and in adult tissues except for the adult brain, but with much lower amount.
ASPM isoforms were generated by alternative splicing. The brain-specific isoform, main ASPM isoform corresponds to a 3477 amino acid residue protein (410 kDa) containing 81 IQ motifs. Alternatively spliced human ASPM variants code for different numbers of IQ domains. Two major ASPM transcripts with sizes of 10.3 and 5.7 kb were identified in all tissues analyzed. The spliced variants of ASPM with variable sizes were also detected in fetus tissue.
ASPM isoforms were generated by alternative splicing. The brain-specific isoform, main ASPM isoform corresponds to a 3477 amino acid residue protein (410 kDa) containing 81 IQ motifs. Alternatively spliced human ASPM variants code for different numbers of IQ domains. Two major ASPM transcripts with sizes of 10.3 and 5.7 kb were identified in all tissues analyzed. The spliced variants of ASPM with variable sizes were also detected in fetus tissue.
Localisation
ASPM protein, a component of the mitotic spindle, localizes to the centrosome in interphase and to the spindle poles from prophase through telophase.
Function
ASPM is a spindle pole/centrosome protein, essential for neurogenic mitosis and possibly controlling the fate-switch to asymmetric cell division through the position of the centrosome at mitosis. The brain-specific main ASPM isoform protein appears to be pivotal for the expansion of cerebral cortical size. Other spliced variants contain different IQ domains or lacking both CH domains and a part of the IQ motifs may be potentially with different functions. One of the smaller proteins detected in cell extracts may be the ASPM product required for mitosis by all dividing cells.
Homology
According to protein identities compared with Human ASPM:
Pan troglodytes: ASPM; (3443/3477, 99%)
Canis lupus familiaris: ASPM; (2738/3392, 80%)
Bos taurus: ASPM; (2557/3484, 73%)
Mus musculus: Aspm; (1882/2948, 63%)
Rattus norvegicus: Aspm; (1851/2939, 62%)
Danio rerio: aspm; (1390/3554, 39%)
Pan troglodytes: ASPM; (3443/3477, 99%)
Canis lupus familiaris: ASPM; (2738/3392, 80%)
Bos taurus: ASPM; (2557/3484, 73%)
Mus musculus: Aspm; (1882/2948, 63%)
Rattus norvegicus: Aspm; (1851/2939, 62%)
Danio rerio: aspm; (1390/3554, 39%)
Mutations

Mutation of ASPM in MCPH. According to Nicholas et al. 2009, all reported autosomal recessive primary microcephaly (MCPH) mutations in ASPM have been presented in this table by different kind of mutations, including nonsense mutations, deletions, insertions and mutations in the splicing region. The reference sequence of mutation sites is according to NM_018136.3.
Implicated in
Entity name
Primary autosomal recessive microcephaly (MCPH)
Note
Primary autosomal recessive microcephaly is a neurodevelopment disorder due to the consequence of deficient neurogenesis within the neurogenic epithelium, resulting in congenital microcephaly (reduced brain size) and mental retardation. MCPH is the consequence of impairment in mitotic spindle regulation in cortical progenitors due to mutations in ASPM.
Homozygous mutations in the ASPM gene, located at MCPH5 locus on chromosome 1q31, are the most common cause of MCPH particularly in the Pakistani population. ASPM mutations are restricted to individuals with an MCPH, no defects other than microcephaly are found in patients carrying mutations in this gene. So far, the phenotypic differences in people with different versions of these genes were not found.
Homozygous mutations in the ASPM gene, located at MCPH5 locus on chromosome 1q31, are the most common cause of MCPH particularly in the Pakistani population. ASPM mutations are restricted to individuals with an MCPH, no defects other than microcephaly are found in patients carrying mutations in this gene. So far, the phenotypic differences in people with different versions of these genes were not found.
Entity name
Hepatocellular carcinoma
Note
ASPM, a component of the mitotic spindle, is shown to express in many human malignant cells nearly and all transformed human cell lines, suggesting that ASPM play an important role in cell proliferation in tumorigenesis.
ASPM mRNA was overexpressed in human hepatocellular carcinoma (HCC), but was very low or undetectable in adult liver, and in benign hepatic tumors, such as hepatocellular adenoma and focal nodular hyperplasia. The overexpression of ASPM correlated with higher-grade (grade II-IV), high-stage (stage IIIA-IV which had vascular invasion) and poor prognosis of HCC. In addition, ASPM overexpression is the most important molecular factor associated with ETR (early tumor recurrence) (intrahepatic tumor recurrence and/or distant metastasis within 12 months after HCC tumor resection), and could be used as a molecular marker predicting enhanced invasive/metastatic potential of HCC.
ASPM mRNA was overexpressed in human hepatocellular carcinoma (HCC), but was very low or undetectable in adult liver, and in benign hepatic tumors, such as hepatocellular adenoma and focal nodular hyperplasia. The overexpression of ASPM correlated with higher-grade (grade II-IV), high-stage (stage IIIA-IV which had vascular invasion) and poor prognosis of HCC. In addition, ASPM overexpression is the most important molecular factor associated with ETR (early tumor recurrence) (intrahepatic tumor recurrence and/or distant metastasis within 12 months after HCC tumor resection), and could be used as a molecular marker predicting enhanced invasive/metastatic potential of HCC.
Entity name
Glioblastoma
Note
ASPM is essential for normal mitotic spindle function in embryonic neuroblasts, and is recognized as a critical regulator of brain size, it may play a role in promoting neuroblast proliferation. Using gene coexpression module in glioblastoma, ASPM was identified as a key gene of glioblastoma, its overexpression was demonstrated in glioblastoma relative to normal brain. siRNA-mediated ASPM knockdown inhibits neural stem cell self renewal and turn forward neural stem cell differentiation. ASPM knockdown also inhibits cell growth of U87-EGFRvIII cells (glioblastoma cells) and the low passage explant culture from a glioblastoma patient. Those result suggesting that ASPM is a potential molecular target in glioblastoma that resulting in the overexpression of ASPM in glioblastoma.
Entity name
Cancers of the uterus and ovary
Note
From high-density oligonucleotide microarrays and using quantitative real-time RT-PCR, ASPM is found more highly expressed in cancers of the uterus and ovary when compared with their normal endometrium counterparts.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 12355089 | 2002 | ASPM is a major determinant of cerebral cortical size. | Bond J et al |
| 14574646 | 2003 | Protein-truncating mutations in ASPM cause variable reduction in brain size. | Bond J et al |
| 16673149 | 2006 | Genetic studies of autosomal recessive primary microcephaly in 33 Pakistani families: Novel sequence variants in ASPM gene. | Gul A et al |
| 17849285 | 2007 | Novel protein-truncating mutations in the ASPM gene in families with autosomal recessive primary microcephaly. | Gul A et al |
| 17090670 | 2006 | Analysis of oncogenic signaling networks in glioblastoma identifies ASPM as a molecular target. | Horvath S et al |
| 15972725 | 2005 | The microcephaly ASPM gene is expressed in proliferating tissues and encodes for a mitotic spindle protein. | Kouprina N et al |
| 15355437 | 2004 | Genetic analysis of primary microcephaly in Indian families: novel ASPM mutations. | Kumar A et al |
| 18676753 | 2008 | ASPM is a novel marker for vascular invasion, early recurrence, and poor prognosis of hepatocellular carcinoma. | Lin SY et al |
| 19028728 | 2009 | The molecular landscape of ASPM mutations in primary microcephaly. | Nicholas AK et al |
| 17534152 | 2007 | ASPM and citron kinase co-localize to the midbody ring during cytokinesis. | Paramasivam M et al |
| 16141009 | 2005 | ASPM mutations identified in patients with primary microcephaly and seizures. | Shen J et al |
| 16123590 | 2005 | The abnormal spindle-like, microcephaly-associated (ASPM) gene encodes a centrosomal protein. | Zhong X et al |
Other Information
Locus ID:
NCBI: 259266
MIM: 605481
HGNC: 19048
Ensembl: ENSG00000066279
Variants:
dbSNP: 259266
ClinVar: 259266
TCGA: ENSG00000066279
COSMIC: ASPM
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000066279 | ENST00000294732 | Q8IZT6 |
| ENSG00000066279 | ENST00000367408 | Q5VYL4 |
| ENSG00000066279 | ENST00000367409 | Q8IZT6 |
Expression (GTEx)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38279565 | 2024 | ASPM stabilizes the NOTCH intracellular domain 1 and promotes oncogenesis by blocking FBXW7 binding in hepatocellular carcinoma cells. | 0 |
| 38576051 | 2024 | RBM10 C761Y mutation induced oncogenic ASPM isoforms and regulated β-catenin signaling in cholangiocarcinoma. | 0 |
| 38279565 | 2024 | ASPM stabilizes the NOTCH intracellular domain 1 and promotes oncogenesis by blocking FBXW7 binding in hepatocellular carcinoma cells. | 0 |
| 38576051 | 2024 | RBM10 C761Y mutation induced oncogenic ASPM isoforms and regulated β-catenin signaling in cholangiocarcinoma. | 0 |
| 36638332 | 2023 | ASPM Activates Hedgehog and Wnt Signaling to Promote Small Cell Lung Cancer Stemness and Progression. | 8 |
| 36883909 | 2023 | ASPM promotes migration and invasion of anaplastic thyroid carcinoma by stabilizing KIF11. | 4 |
| 36980263 | 2023 | The Multiple Mitotic Roles of the ASPM Orthologous Proteins: Insight into the Etiology of ASPM-Dependent Microcephaly. | 4 |
| 37309171 | 2023 | ASPM overexpression enhances cellular proliferation and migration and predicts worse prognosis for papillary renal cell carcinoma. | 2 |
| 36638332 | 2023 | ASPM Activates Hedgehog and Wnt Signaling to Promote Small Cell Lung Cancer Stemness and Progression. | 8 |
| 36883909 | 2023 | ASPM promotes migration and invasion of anaplastic thyroid carcinoma by stabilizing KIF11. | 4 |
| 36980263 | 2023 | The Multiple Mitotic Roles of the ASPM Orthologous Proteins: Insight into the Etiology of ASPM-Dependent Microcephaly. | 4 |
| 37309171 | 2023 | ASPM overexpression enhances cellular proliferation and migration and predicts worse prognosis for papillary renal cell carcinoma. | 2 |
| 34693918 | 2022 | Two novel truncating variants of the ASPM gene identified in a nonconsanguineous Chinese family associated with primary microcephaly. | 2 |
| 34826358 | 2022 | Mutation screening of multiple Pakistani MCPH families revealed novel and recurrent protein-truncating mutations of ASPM. | 4 |
| 35089071 | 2022 | A Two-Base Pair Deletion in IQ Repeats in ASPM Underlies Microcephaly in a Pakistani Family. | 4 |
Citation
Shian-Yang Peng ; Shih-Yeh Lin ; Hey-Chi Hsu
ASPM (asp (abnormal spindle) homolog, microcephaly associated (Drosophila))
Atlas Genet Cytogenet Oncol Haematol. 2009-12-01
Online version: http://atlasgeneticsoncology.org/gene/44463/aspm
