ADAM17 (ADAM metallopeptidase domain 17)
2011-06-01 Astrid Evers , Karina Reiss AffiliationDepartment of Dermatology, Allergology, University Hospital Schleswig Holstein, Campus Kiel, Schittenhelmstrasse 7, Kiel D-24105, Germany
Identity
HGNC
LOCATION
2p25.1
LOCUSID
ALIAS
ADAM18,CD156B,CSVP,NISBD,NISBD1,TACE
FUSION GENES
DNA/RNA
Description
The genomic DNA of ADAM17 extends 66505 base pairs with 19 exons. There is only one known transcription variant.
Transcription
The mRNA of ADAM17 (NM_003183.4) contains 3572 base pairs and the open reading frame spans from bp 184 to bp 2658. Different isoforms due to differential splicing are not known.
Pseudogene
No pseudogenes have been reported for ADAM17.
Proteins
Note
ADAM17 belongs to the ADAM (a disintegrin and metalloprotease) family which is part of the adamalysin proteins.
The activity and function of several membrane proteins is regulated by the proteolytic release of their ectodomains, a process known as shedding. About 2-4% of the proteins on the cell surface are affected by ectodomain shedding (Arribas and Massague, 1995). Since there are several of functionally and structurally diverse substrates, ADAMs are involved in various cellular processes. They take part in the regulation of cell adhesion, migration, development and signalling and are also implicated in pathologic conditions such as inflammation and cancer. The consequences of ectodomain shedding can vary, depending on the function of the substrate protein. Membrane-bound precursor proteins can be released by ADAMs as mature active molecules, for example ligands of the EGF receptor or cytokines like TNFa (Peschon et al., 1998; Blobel, 2005). Receptors, for example ErbB2 and ErbB4, can also be cleaved by ADAMs and thus become inactive or may still function as soluble decoy receptors (Molina et al., 2001; Vecchi et al., 1998). Contrariwise, receptors like Notch require proteolytic cleavage to generate intracellular signalling fragments that act as transcription factors after translocation to the nucleus. In this case ectodomain shedding is the prerequisite for regulated intramembrane proteolysis (RIP) mediated by intramembrane cleaving enzymes (Pan and Rubin, 1997; Brou et al., 2000).
The activity and function of several membrane proteins is regulated by the proteolytic release of their ectodomains, a process known as shedding. About 2-4% of the proteins on the cell surface are affected by ectodomain shedding (Arribas and Massague, 1995). Since there are several of functionally and structurally diverse substrates, ADAMs are involved in various cellular processes. They take part in the regulation of cell adhesion, migration, development and signalling and are also implicated in pathologic conditions such as inflammation and cancer. The consequences of ectodomain shedding can vary, depending on the function of the substrate protein. Membrane-bound precursor proteins can be released by ADAMs as mature active molecules, for example ligands of the EGF receptor or cytokines like TNFa (Peschon et al., 1998; Blobel, 2005). Receptors, for example ErbB2 and ErbB4, can also be cleaved by ADAMs and thus become inactive or may still function as soluble decoy receptors (Molina et al., 2001; Vecchi et al., 1998). Contrariwise, receptors like Notch require proteolytic cleavage to generate intracellular signalling fragments that act as transcription factors after translocation to the nucleus. In this case ectodomain shedding is the prerequisite for regulated intramembrane proteolysis (RIP) mediated by intramembrane cleaving enzymes (Pan and Rubin, 1997; Brou et al., 2000).
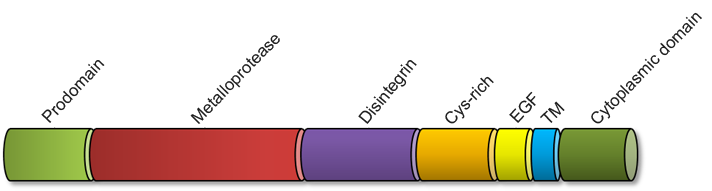
The structure of "a disintegrin and metalloproteases" consists of a N-terminal prodomain (aa 1-214) followed by a metalloprotease domain (aa 223-477), a disintegrin domain (aa 484-560) with a cystein-rich region, an epidermal growth factor-like domain (EGF) (aa 571-602), a transmembrane domain (TM) (aa 672-694) and a cytoplasmic tail (aa 695-824) (Arribas and Esselens, 2009 - modified). Amino acid specifications according to Pubmed graphics for ADAM17 (NP_003174.3).
Description
The preproform of ADAM17 contains 824 amino acids (134 kDa). After cleavage of the prodomain by furin or other proprotein convertases, the mature form has a predicted molecular weight of 98 kDa. The prodomain serves as a chaperone and keeps the enzyme in an inactive state during maturation in the trans-Golgi network (Schlöndorff et al., 2000). The catalytic domain contains the characteristic zinc binding motif (HEXXHXXGXXH) and three potential N-glycosylation sites. The disintegrin domain is discussed to be involved in regulation of the catalytic activity of ADAM17 (Li and Fan, 2004; Smith et al., 2002), but might also be involved in interaction with integrins and therefore mediate cell-cell adhesion or cell-matrix interaction. The cytoplasmic tail of ADAM17 was thought to be important in regulation of the enzyme activity because of its potential tyrosine phosphorylation site, but recent studies showed that overexpressed ADAM17 lacking its cytoplasmic domain was able to cleave TNFalpha upon phorbolester stimulation (Reddy et al., 2000; Horiuchi et al., 2007). The transmembrane domain might play a role in regulation of ADAM activity as studies with ADAM-chimera with exchanged transmembrane domains and experiments with plasma membrane modulating stimuli indicate (Le Gall et al., 2010; Reiss et al., 2011).
Expression
ADAM17 is ubiquitously expressed in various tissues, for example in the brain, kidney, heart and skeletal muscles. The expression pattern changes during embryonic development and the adult life (Black et al., 1997).
Localisation
ADAM17 is a type I transmembrane protein, but only 10% of the total protein can be found in the plasma membrane on the cell surface (Schlöndorff et al., 2000). Most of the protein is localised in the endoplasmic reticulum and trans-Golgi network. It is suggested that ADAM17 accumulates in cholesterol rich departments of the plasma membrane (lipid rafts) (Tellier et al., 2006).
Function
ADAM17 knockout mice display a severe phenotype that resembles mice lacking the epidermal growth factor receptor (EGFR) or one of its ligands (TGFalpha, HB-EGF or amphiregulin). Perinatal mortality is probably due to defects in heart development.
Because of their catalytic and their disintegrin domain, ADAMs can have both adhesive and proteolytic functions. ADAM17 is best studied for its proteolytic properties and cleaves its substrates in the juxtamembrane region. The proteins processed by ADAM17 have different functions and therefore proteolytic processing can either activate or abrogate processes. For example both receptors and ligands can be cleaved, thus signalling can be either initiated or stopped by ADAM17.
Because of their catalytic and their disintegrin domain, ADAMs can have both adhesive and proteolytic functions. ADAM17 is best studied for its proteolytic properties and cleaves its substrates in the juxtamembrane region. The proteins processed by ADAM17 have different functions and therefore proteolytic processing can either activate or abrogate processes. For example both receptors and ligands can be cleaved, thus signalling can be either initiated or stopped by ADAM17.
Homology
Homologs of the human ADAM17 are found in various species ranging from other mammalians to primitive chordates like zebrafish. See table 1 for results of NCBI-Blast/blastp with human ADAM17 (NP_003174.3) as query sequence. Its closest relative in the family of human ADAM metalloproteases is ADAM10 (NP_001101.1), but there is only less than 30% amino acid sequence identity according to NCBI-Blast (blastp).
| Species | Accession # | identity to human ADAM17 | similarity to human ADAM17 |
| Sus scrofa | NP_001093396.1 | 92% | 97% |
| Mus musculus | NP_033745.4 | 92% | 96% |
| Rattus norwegicus | NP_064702.1 | 92% | 96% |
| Gallus gallus | NP_001008682.1 | 77% | 87% |
| Xenopus laevis | NP_001089130.1 | 73% | 85% |
| Danio rerio | NP_955967.1 | 61% | 74% |
| Drosophila melanogaster | NP_733334.1 | 42% | 59% |
Mutations
Note
So far no common germinal or somatic mutation variants are known.
Implicated in
Entity name
Cancer
Note
Various growth factors necessary for tumor progression and growth are shed by ADAM17 and increased shedding of EGFR ligands was observed in tissues developing a malignant phenotype (Katakowski et al., 2009). ADAM17 is supposed to play a role in different malignancies: increased levels of the enzyme were detected in gastric carcinoma (Yoshimura et al., 2002), primary colon carcinoma (Blanchot-Jossic et al., 2005), skin malignancies (Oh et al., 2009) and ovarian cancer (Tanaka et al., 2005) but its role is best studied in breast cancer (see below).
Entity name
Breast cancer
Disease
It was shown that overexpression of ADAM17 in breast cancer correlated with TGFalpha expression (Borrell-Pages et al., 2003), metastasis and tumor progression (McGowan et al., 2007) and shorter survival of patients (McGowan et al., 2008). Recent studies by Kenny and Bissell demonstrated that the malignant phenotype of a breast cancer cell line was reverted to a normal phenotype using siRNA against ADAM17 (Kenny and Bissell, 2007).
Entity name
Inflammation
Note
The identification of ADAM17 as TNFalpha converting enzyme suggested an important role of this metalloprotease in inflammatory diseases with elevated levels of soluble TNFalpha, such as rheumatoid arthritis and inflammatory bowel disease. Indeed, increased enzymatic activity of ADAM17 was shown in tissues of patients with osteoarthritis (Amin, 1999) and rheumatoid arthritis (Ohta et al., 2001). Furthermore ADAM17 was shown to process several factors that are involved in leukocyte recruitment to the site of inflammation. Moreover ADAM17 regulates leukocyte transmigration through the vascular endothelium for example by cleaving adhesion molecules between endothelial cells. The vascular cell adhesion molecule (CC: TXT: V-CAM ID: 42784>) is released by ADAM17 and functions as the ligand of the leukocyte very late antigen 4 (VLA-4 or alpha4beta1 integrin), which is implicated in the leukocyte adhesion to the vascular endothelium (Garton et al., 2003). L-Selectin cleavage by ADAM17 promotes leukocyte migration through the basal membrane after adhering to the endothelium (Faveeuw et al., 2001; Peschon et al., 1998). ADAM17 was also identified as the sheddase of the tight junction molecule JAM-1 between endothelial cells (Koenen et al., 2009). Cleavage of JAM-1 contributes to the passage of leukocytes through the endothelial cell layer and the processed molecule serves as a biomarker of inflammation.
Disease
- rheumatoid arthritis (see above),
- osteoarthritis (see above),
- inflammatory bowel disease (high levels of ADAM17 expression were reported in epithelial cells during the active phase of Crohns disease (Cesaro et al., 2009)),
- psoriasis (ADAM17 expression was upregulated in keratinocytes, blood vessels and mast cells from patients (Kawaguchi et al., 2005)),
- pulmonary inflammation (reduced invasion of eosinophils in a model of acute allergic lung inflammation after treatment with ADAM17/MMP inhibitors (Trifilieff et al., 2002)).
- osteoarthritis (see above),
- inflammatory bowel disease (high levels of ADAM17 expression were reported in epithelial cells during the active phase of Crohns disease (Cesaro et al., 2009)),
- psoriasis (ADAM17 expression was upregulated in keratinocytes, blood vessels and mast cells from patients (Kawaguchi et al., 2005)),
- pulmonary inflammation (reduced invasion of eosinophils in a model of acute allergic lung inflammation after treatment with ADAM17/MMP inhibitors (Trifilieff et al., 2002)).
Entity name
Alzheimers disease
Note
The majority of the amyloid precursor protein (APP) is physiologically cleaved by alpha-secretase (ADAM10 or ADAM17). This cleavage leads to the generation of a soluble non-amyloidogenic fragment (sAPPalpha), whereas shedding of APP by beta-secretase produces the amyloidogenic Abeta peptide (Allinson et al., 2003). Accordingly, decreased activity of alpha-secretase results in the formation of amyloid plaques. While ADAM10 represents the most important alpha-secretase, ADAM17 could contribute to APP-processing under certain conditions (Buxbaum et al., 1998; Jorissen et al., 2010).
Entity name
Kidney diseases
Note
In polycystic kidney disease (PDK) increased activation of the EGF receptor leads to cyst formation and loss of kidney function. These symptoms could be significantly decreased by treatment with ADAM17 inhibitors (Richards et al., 1998; Dell et al., 2001). Which EGFR ligand is responsible in this context remains unclear, but TGFalpha knockout mice were shown to still develop PDK (Nemo et al., 2005). ADAM17 is also involved in chronic kidney disease (CDK) and it is suggested that this is due to the transactivation of EGFR through the GPCR agonist angiotensin-II (Lautrette et al., 2005). In this process ADAM17 is activated by the GPCR and releases TGFalpha, which in turn activates the EGFR.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 14598310 | 2003 | ADAMs family members as amyloid precursor protein alpha-secretases. | Allinson TM et al |
| 10419777 | 1999 | Regulation of tumor necrosis factor-alpha and tumor necrosis factor converting enzyme in human osteoarthritis. | Amin AR et al |
| 19601834 | 2009 | ADAM17 as a therapeutic target in multiple diseases. | Arribas J et al |
| 7844156 | 1995 | Transforming growth factor-alpha and beta-amyloid precursor protein share a secretory mechanism. | Arribas J et al |
| 9034190 | 1997 | A metalloproteinase disintegrin that releases tumour-necrosis factor-alpha from cells. | Black RA et al |
| 16041691 | 2005 | Up-regulated expression of ADAM17 in human colon carcinoma: co-expression with EGFR in neoplastic and endothelial cells. | Blanchot-Jossic F et al |
| 15688065 | 2005 | ADAMs: key components in EGFR signalling and development. | Blobel CP et al |
| 12606576 | 2003 | TACE is required for the activation of the EGFR by TGF-alpha in tumors. | Borrell-Pagès M et al |
| 10882063 | 2000 | A novel proteolytic cleavage involved in Notch signaling: the role of the disintegrin-metalloprotease TACE. | Brou C et al |
| 9774383 | 1998 | Evidence that tumor necrosis factor alpha converting enzyme is involved in regulated alpha-secretase cleavage of the Alzheimer amyloid protein precursor. | Buxbaum JD et al |
| 19299578 | 2009 | Differential expression and regulation of ADAM17 and TIMP3 in acute inflamed intestinal epithelia. | Cesaro A et al |
| 11576338 | 2001 | A novel inhibitor of tumor necrosis factor-alpha converting enzyme ameliorates polycystic kidney disease. | Dell KM et al |
| 11468168 | 2001 | Transendothelial migration of lymphocytes across high endothelial venules into lymph nodes is affected by metalloproteinases. | Faveeuw C et al |
| 12878595 | 2003 | Stimulated shedding of vascular cell adhesion molecule 1 (VCAM-1) is mediated by tumor necrosis factor-alpha-converting enzyme (ADAM 17). | Garton KJ et al |
| 20184396 | 2010 | ADAM-17: the enzyme that does it all. | Gooz M et al |
| 17079736 | 2007 | Substrate selectivity of epidermal growth factor-receptor ligand sheddases and their regulation by phorbol esters and calcium influx. | Horiuchi K et al |
| 20371803 | 2010 | The disintegrin/metalloproteinase ADAM10 is essential for the establishment of the brain cortex. | Jorissen E et al |
| 19515085 | 2009 | Tumorigenicity of cortical astrocyte cell line induced by the protease ADAM17. | Katakowski M et al |
| 15888146 | 2005 | Overexpression of tumour necrosis factor-alpha-converting enzyme in psoriasis. | Kawaguchi M et al |
| 17218988 | 2007 | Targeting TACE-dependent EGFR ligand shedding in breast cancer. | Kenny PA et al |
| 19258599 | 2009 | Regulated release and functional modulation of junctional adhesion molecule A by disintegrin metalloproteinases. | Koenen RR et al |
| 16041383 | 2005 | Angiotensin II and EGF receptor cross-talk in chronic kidney diseases: a new therapeutic approach. | Lautrette A et al |
| 20980382 | 2010 | ADAM17 is regulated by a rapid and reversible mechanism that controls access to its catalytic site. | Le Gall SM et al |
| 15075334 | 2004 | Loss of ectodomain shedding due to mutations in the metalloprotease and cysteine-rich/disintegrin domains of the tumor necrosis factor-alpha converting enzyme (TACE). | Li X et al |
| 18238782 | 2008 | ADAM-17 predicts adverse outcome in patients with breast cancer. | McGowan PM et al |
| 11406546 | 2001 | Trastuzumab (herceptin), a humanized anti-Her2 receptor monoclonal antibody, inhibits basal and activated Her2 ectodomain cleavage in breast cancer cells. | Molina MA et al |
| 15774823 | 2005 | Transforming growth factor alpha (TGF-alpha) and other targets of tumor necrosis factor-alpha converting enzyme (TACE) in murine polycystic kidney disease. | Nemo R et al |
| 19278423 | 2009 | The disintegrin-metalloproteinases ADAM 10, 12 and 17 are upregulated in invading peripheral tumor cells of basal cell carcinomas. | Oh ST et al |
| 11508576 | 2001 | Tumor necrosis factor-alpha (TNF-alpha) converting enzyme contributes to production of TNF-alpha in synovial tissues from patients with rheumatoid arthritis. | Ohta S et al |
| 9244301 | 1997 | Kuzbanian controls proteolytic processing of Notch and mediates lateral inhibition during Drosophila and vertebrate neurogenesis. | Pan D et al |
| 9812885 | 1998 | An essential role for ectodomain shedding in mammalian development. | Peschon JJ et al |
| 10799547 | 2000 | Functional analysis of the domain structure of tumor necrosis factor-alpha converting enzyme. | Reddy P et al |
| 21642425 | 2011 | Unsaturated fatty acids drive disintegrin and metalloproteinase (ADAM)-dependent cell adhesion, proliferation, and migration by modulating membrane fluidity. | Reiss K et al |
| 9486961 | 1998 | Epidermal growth factor receptor activity mediates renal cyst formation in polycystic kidney disease. | Richards WG et al |
| 21194787 | 2011 | The "A Disintegrin And Metalloproteases" ADAM10 and ADAM17: novel drug targets with therapeutic potential? | Saftig P et al |
| 10727411 | 2000 | Intracellular maturation and localization of the tumour necrosis factor alpha convertase (TACE). | Schlöndorff J et al |
| 12460986 | 2002 | The cysteine-rich domain regulates ADAM protease function in vivo. | Smith KM et al |
| 16000575 | 2005 | Clinical significance of heparin-binding epidermal growth factor-like growth factor and a disintegrin and metalloprotease 17 expression in human ovarian cancer. | Tanaka Y et al |
| 17010968 | 2006 | The shedding activity of ADAM17 is sequestered in lipid rafts. | Tellier E et al |
| 11934805 | 2002 | Pharmacological profile of PKF242-484 and PKF241-466, novel dual inhibitors of TNF-alpha converting enzyme and matrix metalloproteinases, in models of airway inflammation. | Trifilieff A et al |
| 9685416 | 1998 | Tyrosine phosphorylation and proteolysis. Pervanadate-induced, metalloprotease-dependent cleavage of the ErbB-4 receptor and amphiregulin. | Vecchi M et al |
| 11807715 | 2002 | ADAMs (a disintegrin and metalloproteinase) messenger RNA expression in Helicobacter pylori-infected, normal, and neoplastic gastric mucosa. | Yoshimura T et al |
Other Information
Locus ID:
NCBI: 6868
MIM: 603639
HGNC: 195
Ensembl: ENSG00000151694
Variants:
dbSNP: 6868
ClinVar: 6868
TCGA: ENSG00000151694
COSMIC: ADAM17
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37540030 | 2024 | The protease ADAM17 at the crossroads of disease: revisiting its significance in inflammation, cancer, and beyond. | 7 |
| 37999645 | 2024 | Neuraminidase-induced externalization of phosphatidylserine activates ADAM17 and impairs insulin signaling in endothelial cells. | 0 |
| 38063205 | 2024 | m(6)A‑modified HOXC10 promotes HNSCC progression via co‑activation of ADAM17/EGFR and Wnt/β‑catenin signaling. | 0 |
| 38287826 | 2024 | MiR-145 Alleviates Sepsis-Induced Inflammatory Responses and Organ Injury by Targeting ADAM17. | 0 |
| 38335437 | 2024 | Expression of ADAM17 and its clinical value for patients with pernicious placenta previa: A retrospective study of 148 PPP patients underwent cesarean section. | 0 |
| 38413999 | 2024 | Tumor-derived exosomal ADAM17 promotes pre-metastatic niche formation by enhancing vascular permeability in colorectal cancer. | 1 |
| 38550672 | 2024 | The Important Role of Phosphatidylserine, ADAM17, TNF-Alpha, and Soluble MER on Efferocytosis Activity in Central Obesity. | 0 |
| 38673989 | 2024 | Regulation of Mertk Surface Expression via ADAM17 and γ-Secretase Proteolytic Processing. | 0 |
| 38781971 | 2024 | Cryo-EM reveals that iRhom2 restrains ADAM17 protease activity to control the release of growth factor and inflammatory signals. | 0 |
| 38863060 | 2024 | Downregulation of the metalloproteinases ADAM10 or ADAM17 promotes osteoclast differentiation. | 0 |
| 38897568 | 2024 | Ectodomain shedding of PLA2R1 is mediated by the metalloproteases ADAM10 and ADAM17. | 0 |
| 37540030 | 2024 | The protease ADAM17 at the crossroads of disease: revisiting its significance in inflammation, cancer, and beyond. | 7 |
| 37999645 | 2024 | Neuraminidase-induced externalization of phosphatidylserine activates ADAM17 and impairs insulin signaling in endothelial cells. | 0 |
| 38063205 | 2024 | m(6)A‑modified HOXC10 promotes HNSCC progression via co‑activation of ADAM17/EGFR and Wnt/β‑catenin signaling. | 0 |
| 38287826 | 2024 | MiR-145 Alleviates Sepsis-Induced Inflammatory Responses and Organ Injury by Targeting ADAM17. | 0 |
Citation
Astrid Evers ; Karina Reiss
ADAM17 (ADAM metallopeptidase domain 17)
Atlas Genet Cytogenet Oncol Haematol. 2011-06-01
Online version: http://atlasgeneticsoncology.org/gene/572/adam17
