NRAS (neuroblastoma RAS viral oncogene homolog)
1999-02-01 Franz Watzinger , Thomas Lion AffiliationChildrens Cancer Research Institute (CCRI), Kinderspitalgasse 6, A-1090 Vienna, Austria
Identity
HGNC
LOCATION
1p13.2
LOCUSID
ALIAS
ALPS4,CMNS,N-ras,NCMS,NRAS1,NS6
FUSION GENES
DNA/RNA
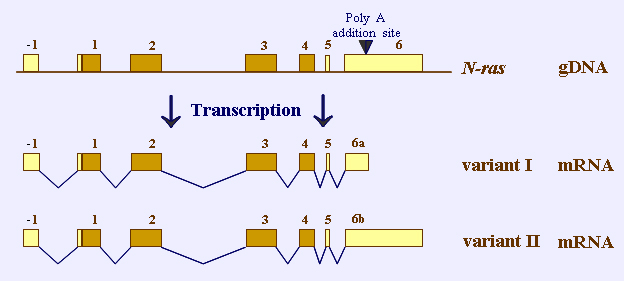
evolution of two N-ras mRNA transcripts
the slightly differing consensus sequence of the poly A addition site (AATATA instead of AATAAA) leads to inefficient processing and to two different RNA transcripts; exons that encode protein are shown as black boxes, untranslated exons as white boxes; the upstream untranslated exon is indicated as Exon -1
Description
consists of seven exons, spread over 8kb of genomic DNA
Transcription
inefficient processing of pre-mRNA reveals two different transcripts of 4.3kb and 2kb (see Fig), differing in the extension of the 3 end of the smaller message; the longer transcript is probably a result of an RNA-extension through the termination site
Proteins
Description
p21N-ras; regular RAS protein - characterized in the page
Expression
ubiquitously expressed
Localisation
anchored to the inner surface of the plasma membrane
Function
analogously to other GTP-binding proteins (such as Translation Elongation Factor EFTu or signal transducing G-Proteins) RAS proteins are involved in signal transduction pathways
Homology
ras gene family is part of the ras superfamily including the mammalian RAS, RAL, RAC, RHO, RAP, and RAB gene families and the yeast homologs like SEC4 and YPT1 genes; genes encode small monomeric proteins of low molecular mass (20-30 kDa) which share at least 30% homology to RAS proteins
Implicated in
Entity name
tumor (frequency of N-RAS mutations); references in Full Bibliography
Entity name
acute non lymphocytic leukemia and myelodysplasia (20-40%)
Entity name
chronic myelogenous leukemia, acute lymphocytic leukemia (0-10%)
Entity name
brain (0-15%)
Entity name
skin (0-20%)
Entity name
thyroid (0-60%)
Entity name
testis (0-40%)
Entity name
stomach (gastric tumors) (5%)
Entity name
testis liver (0-15%)
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 6700714 | 1984 | Transforming ras genes from human melanoma: a manifestation of tumour heterogeneity? | Albino AP et al |
| 3292251 | 1988 | Incidence of ras gene mutations in neuroblastoma. | Ballas K et al |
| 2114981 | 1990 | ras oncogenes: their role in neoplasia. | Barbacid M et al |
| 2547513 | 1989 | ras oncogenes in human cancer: a review. | Bos JL et al |
| 8919545 | 1996 | Ras oncogene mutations in thyroid tumors: polymerase chain reaction-restriction-fragment-length polymorphism analysis from paraffin-embedded tissues. | Capella G et al |
| 1323601 | 1992 | Infrequent point mutations in codons 12 and 61 of ras oncogenes in human hepatocellular carcinomas. | Challen C et al |
| 9156660 | 1997 | N-ras gene point mutations in Brazilian acute myelogenous leukemia patients correlate with a poor prognosis. | De Melo MB et al |
| 3600795 | 1987 | A new oncogene in human thyroid papillary carcinomas and their lymph-nodal metastases. | Fusco A et al |
| 9137494 | 1997 | Mutation of ras oncogene in gastric adenocarcinoma: association with histological phenotype. | Kim TY et al |
| 2648253 | 1989 | High frequency of ras oncogene activation in all stages of human thyroid tumorigenesis. | Lemoine NR et al |
| 9107428 | 1997 | Ras oncogene mutations in childhood brain tumors. | Maltzman TH et al |
| 1421167 | 1992 | ras activation in experimental carcinogenesis. | Mangues R et al |
| 1892776 | 1991 | No N-ras mutations in human uveal melanoma: the role of ultraviolet light revisited. | Mooy CM et al |
| 1381946 | 1992 | Detection of RAS mutations in archival testicular germ cell tumors by polymerase chain reaction and oligonucleotide hybridization. | Moul JW et al |
| 2682461 | 1989 | Activated ras genes in human seminoma: evidence for tumor heterogeneity. | Mulder MP et al |
| 1574246 | 1992 | Multiple point mutation of N-ras and K-ras oncogenes in myelodysplastic syndrome and acute myelogenous leukemia. | Nakagawa T et al |
| 8123851 | 1994 | Prognostic importance of mutations in the ras proto-oncogenes in de novo acute myeloid leukemia. | Neubauer A et al |
| 3307760 | 1987 | Human cancer and cellular oncogenes. | Nishimura S et al |
| 9639416 | 1998 | RAS, FMS and p53 mutations and poor clinical outcome in myelodysplasias: a 10-year follow-up. | Padua RA et al |
| 8143614 | 1993 | K-ras oncogene codon 12 point mutations in testicular cancer. | Ridanpää M et al |
| 8505202 | 1993 | Investigation of the role of the ras protooncogene point mutation in human uveal melanomas. | Soparker CN et al |
| 3283656 | 1988 | Detection of activated ras oncogenes in human thyroid carcinomas. | Suárez HG et al |
| 1544704 | 1992 | N-ras gene mutations in acute myeloid leukemia: accurate detection by solid-phase minisequencing. | Syvänen AC et al |
| 1739910 | 1992 | High incidence of ras gene mutation in intrahepatic cholangiocarcinoma. | Tada M et al |
| 8033117 | 1994 | Absence of N-ras mutations in myeloid and lymphoid blast crisis of chronic myeloid leukemia. | Watzinger F et al |
| 9593469 | 1998 | Correlation of N-ras point mutations with specific chromosomal abnormalities in primary myelodysplastic syndrome. | de Souza Fernandez T et al |
| 2674680 | 1989 | N-ras mutations in human cutaneous melanoma from sun-exposed body sites. | van 't Veer LJ et al |
Other Information
Locus ID:
NCBI: 4893
MIM: 164790
HGNC: 7989
Ensembl: ENSG00000213281
Variants:
dbSNP: 4893
ClinVar: 4893
TCGA: ENSG00000213281
COSMIC: NRAS
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000213281 | ENST00000369535 | P01111 |
| ENSG00000213281 | ENST00000369535 | Q5U091 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
PharmGKB
| Entity ID | Name | Type | Evidence | Association | PK | PD | PMIDs |
|---|---|---|---|---|---|---|---|
| PA10040 | cetuximab | Chemical | LabelAnnotation, VipGene | associated | 26438111 | ||
| PA153561371 | egfr inhibitors | Chemical | VipGene | associated | 26438111 | ||
| PA162373091 | panitumumab | Chemical | LabelAnnotation, VipGene | associated | 26438111 | ||
| PA165946873 | vemurafenib | Chemical | LabelAnnotation, VipGene | associated | |||
| PA166114911 | dabrafenib | Chemical | LabelAnnotation, VipGene | associated | |||
| PA166115364 | trametinib | Chemical | LabelAnnotation | associated | |||
| PA24928 | ARAF | Gene | Pathway | associated | |||
| PA25408 | BRAF | Gene | Pathway | associated | 28362716 | ||
| PA26880 | CRK | Gene | Pathway | associated | 20124951 | ||
| PA28127 | FGFR1 | Gene | Pathway | associated | 28362716 | ||
| PA28180 | FLT1 | Gene | Pathway | associated | 28362716 | ||
| PA28181 | FLT3 | Gene | Pathway | associated | 28362716 | ||
| PA28183 | FLT4 | Gene | Pathway | associated | 28362716 | ||
| PA30086 | KDR | Gene | Pathway | associated | 28362716 | ||
| PA30128 | KIT | Gene | Pathway | associated | 28362716 | ||
| PA30592 | MAP3K1 | Gene | Pathway | associated | |||
| PA33148 | PDGFRB | Gene | Pathway | associated | 28362716 | ||
| PA34183 | RAF1 | Gene | Pathway | associated | 20124951, 28362716 | ||
| PA34335 | RET | Gene | Pathway | associated | 28362716 | ||
| PA443622 | Carcinoma, Non-Small-Cell Lung | Disease | ClinicalAnnotation | associated | PD | 21636554 | |
| PA444760 | Leukemia, Myeloid, Acute | Disease | VipGene | associated | |||
| PA444903 | Melanoma | Disease | VipGene | associated | |||
| PA446108 | Colorectal Neoplasms | Disease | VipGene | associated | 21305640 | ||
| PA448803 | carboplatin | Chemical | ClinicalAnnotation | associated | PD | 21636554 | |
| PA449014 | cisplatin | Chemical | ClinicalAnnotation | associated | PD | 21636554 | |
| PA449748 | gemcitabine | Chemical | ClinicalAnnotation | associated | PD | 21636554 | |
| PA7360 | EGFR | Gene | Pathway | associated |
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37953090 | 2024 | Targeted next-generation sequencing of Japanese patients with sinonasal mucosal melanomas identifies frequent NRAS and CTNNB1 mutations. | 0 |
| 38031763 | 2024 | Mechanism of co-operation of mutant IL-7Rα and mutant NRAS in acute lymphoblastic leukemia: role of MYC. | 2 |
| 38254245 | 2024 | Missense mutation of NRAS is associated with malignant progression in neurocutaneous melanosis. | 0 |
| 38294692 | 2024 | NRAS Mutant Dictates AHCYL1-Governed ER Calcium Homeostasis for Melanoma Tumor Growth. | 0 |
| 38325046 | 2024 | miR-196b-5p regulates inflammatory process and migration via targeting Nras in trabecular meshwork cells. | 0 |
| 38579072 | 2024 | Epidemiological and clinicopathological features of KRAS, NRAS, BRAF mutations and MSI in Chinese patients with stage I-III colorectal cancer. | 0 |
| 38759471 | 2024 | Deep neural network for the prediction of KRAS, NRAS, and BRAF genotypes in left-sided colorectal cancer based on histopathologic images. | 0 |
| 38809881 | 2024 | Changes in cell morphology and function induced by the NRAS Q61R mutation in lymphatic endothelial cells. | 0 |
| 38875469 | 2024 | KRAS, NRAS and BRAF Mutational Profile of Colorectal Cancer in a Series of Moroccan Patients. | 0 |
| 37953090 | 2024 | Targeted next-generation sequencing of Japanese patients with sinonasal mucosal melanomas identifies frequent NRAS and CTNNB1 mutations. | 0 |
| 38031763 | 2024 | Mechanism of co-operation of mutant IL-7Rα and mutant NRAS in acute lymphoblastic leukemia: role of MYC. | 2 |
| 38254245 | 2024 | Missense mutation of NRAS is associated with malignant progression in neurocutaneous melanosis. | 0 |
| 38294692 | 2024 | NRAS Mutant Dictates AHCYL1-Governed ER Calcium Homeostasis for Melanoma Tumor Growth. | 0 |
| 38325046 | 2024 | miR-196b-5p regulates inflammatory process and migration via targeting Nras in trabecular meshwork cells. | 0 |
| 38579072 | 2024 | Epidemiological and clinicopathological features of KRAS, NRAS, BRAF mutations and MSI in Chinese patients with stage I-III colorectal cancer. | 0 |
Citation
Franz Watzinger ; Thomas Lion
NRAS (neuroblastoma RAS viral oncogene homolog)
Atlas Genet Cytogenet Oncol Haematol. 1999-02-01
Online version: http://atlasgeneticsoncology.org/gene/92/nras
