ACLY (ATP citrate lyase)
2012-10-01 Marie E Beckner AffiliationDepartment of Pathology, Louisiana State University Health Sciences Center - Shreveport, USA
DNA/RNA
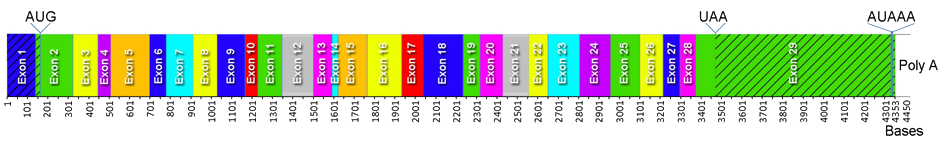
Homo sapiens ATP citrate lyase (ACLY), transcript variant 1, 4450 bp mRNA, encodes a 1101 aa protein. NCBI Reference Sequence: NM_001096. Locus NM_001096.
Description
Two transcript variants have been identified and this variant (1) represents the longer ACLY transcript. It encodes the longer isoform of ACLY. Placement of code for the initiating methionine, stop codon, poly adenylation signal, boundaries of the 29 exons, and the untranslated region (hatched) are shown. Location of missing sequence in variant 2 compared to variant 1 is indicated in the diagram of the ACLY protein shown below. The sequence for ACLY has been conserved in evolution, putatively from an ancient single gene present prior to separation of animals and fungi with some fungi subsequently developing two genes to code for complete ACLY whereas animals have retained a single gene.
Transcription
4450 bp mRNA (NCBI RefSeq, May-2012). Multiple Sp1 binding sites and CAAT are present in the promoter of rat ACLY and it can be induced by a low fat/high carbohydrate diet.
Pseudogene
None known at this time.
Proteins
Note
ACLY is a metabolic enzyme found as a tetramer of apparently identical subunits (440000 molecular weight). It was discovered in 1950s. ACLY cleaves citric acid in a multistep process with participation of cofactors to form the products, acetyl-CoA and oxaloacetate. Functional domains of ACLY resemble regions of related enzymes that can play similar roles in metabolism of other substrates.
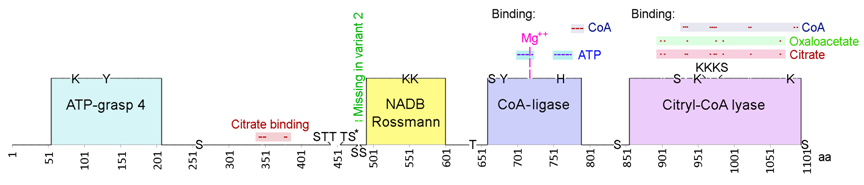
ATP citrate lyase (ACL or ACLY), variant 1. GenBank: AAH06195 protein sequence with locations of functional domains, multiple binding regions, Rossman fold (492-601), and post-translational modifications, including potential phosphorylation of tyrosines (131, 682), serines (260, 442, 455, 478, 481, 663, 839, 922, 979, 1100), threonines (445, 447, 453, 639), and a histidine (760), and N6-acetylysine (86, 546, 554, 948, 962, 968, 978, 1077). The missing sequence (476-485) in variant 2 results in the loss of 2 serines as indicated in the diagram.
Description
Four of these subunits form a homotetramer.
Expression
Prokaryotes and eukaryotes. The association between increased expression for ACLY and the gene encoding enolase, ENO1, is highly statistically significant. Greater expression of ACLY can be found in mammalian cells under hypoxic conditions. It is more highly expressed in many malignant tissues when compared to their benign counterparts. Aberrant expression can be found in breast, liver, colon, lung, and prostate cancers and is inversely correlated with tumor stage and differentiation so that increased ACLY expression is a negative prognostic factor. ACLYs knockdown in non-small cell lung carcinoma (NSCLC) can lead to apoptosis and differentiation in vitro and less growth in vivo.
Localisation
ACLY is a relatively abundant cytoplasmic protein and can be associated with outer surfaces of mitochondria and it is also preferentially distributed to pseudopodia in migrating cells. Relatively small amounts have been found in the nuclei. Also, ACLY is found in synaptosomes.
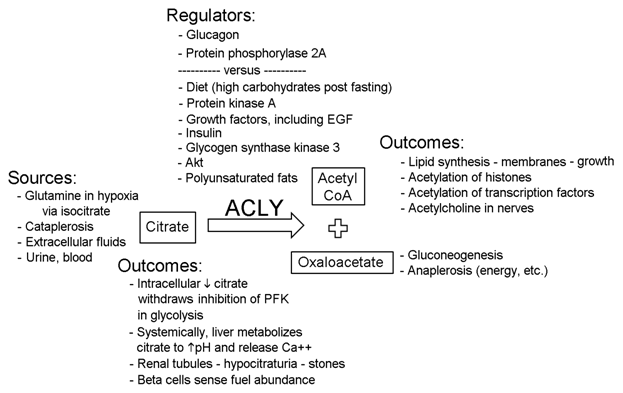
Function
ACLY catalyzes the following reaction: ATP + citrate + CoA = ADP + phosphate + acetyl-CoA + oxaloacetate.
ACLY is well-known for linking carbohydrate and lipid metabolism which can lead to membrane production during cell growth. However, a myriad of other consequences from the breakdown of citrate also occur and are indicated above. Systemically and locally the effects of ACLYs activity can have a powerfull impact. These include alteration of transcription. Citrate passes through nuclear pores and undergoes cleavage by the small amounts of ACLY in the nucleus to generate acetyl CoA that affects transcription via acetylation of histones and transcription factors. Cataplerosis includes citrates transport from the mitochondria via a transporter to provide cytosolic citrate. The transfer of metabolites into mitochondria via shuttles, transporters, etc. constitutes anaplerosis so that either energy or amino acids can be formed, depending on the oxygenation state and the cells needs. Regulation of ACLY is complex and appears to resemble that of glycogen synthase in regard to phosphorylations occurring sequentially in a hierarchical manner. The multiple sources of citrate help to explain the varying effects of ACLY. Note that exogenous citrate can come from anticoagulants. Although ACLY is susceptible to proteolysis, the lower weight (53 kDa) digestion product of ACLY retains its activity.
Loss of ACLY function in plants can result in a bonsai phenotype. Hydroxycitrate, found in the fruit of a tropical tree, Garcinia cambogia (bitter kola) that grows in Southeast Asia and southern India, is a competitive inhibitor and has been extensively used in functional studies. However, solubility issues and the large quantities of hydroxycitrate needed for inhibition of ACLY are disadvantages for using it in clinical studies. Proprietary formulations are available as weight loss supplements and at least one of these has been combined with established anti-cancer agents. The cell-penetrant gamma-lactone, SB-204990, a prodrug of SB-201076, was described in 1998 as an oral drug to inhibit ACLY and has been used in several cancer studies more recently. Radicicol and tartrate are also inhibitors of ACLY. Multiple agents, such as α-lipoic acid, statins, capsaicin, a Met kinase inhibitor (SU11274), etc., have been found to enhance the effects of ACLY inhibitors in small studies of tumors.
ACLY is well-known for linking carbohydrate and lipid metabolism which can lead to membrane production during cell growth. However, a myriad of other consequences from the breakdown of citrate also occur and are indicated above. Systemically and locally the effects of ACLYs activity can have a powerfull impact. These include alteration of transcription. Citrate passes through nuclear pores and undergoes cleavage by the small amounts of ACLY in the nucleus to generate acetyl CoA that affects transcription via acetylation of histones and transcription factors. Cataplerosis includes citrates transport from the mitochondria via a transporter to provide cytosolic citrate. The transfer of metabolites into mitochondria via shuttles, transporters, etc. constitutes anaplerosis so that either energy or amino acids can be formed, depending on the oxygenation state and the cells needs. Regulation of ACLY is complex and appears to resemble that of glycogen synthase in regard to phosphorylations occurring sequentially in a hierarchical manner. The multiple sources of citrate help to explain the varying effects of ACLY. Note that exogenous citrate can come from anticoagulants. Although ACLY is susceptible to proteolysis, the lower weight (53 kDa) digestion product of ACLY retains its activity.
Loss of ACLY function in plants can result in a bonsai phenotype. Hydroxycitrate, found in the fruit of a tropical tree, Garcinia cambogia (bitter kola) that grows in Southeast Asia and southern India, is a competitive inhibitor and has been extensively used in functional studies. However, solubility issues and the large quantities of hydroxycitrate needed for inhibition of ACLY are disadvantages for using it in clinical studies. Proprietary formulations are available as weight loss supplements and at least one of these has been combined with established anti-cancer agents. The cell-penetrant gamma-lactone, SB-204990, a prodrug of SB-201076, was described in 1998 as an oral drug to inhibit ACLY and has been used in several cancer studies more recently. Radicicol and tartrate are also inhibitors of ACLY. Multiple agents, such as α-lipoic acid, statins, capsaicin, a Met kinase inhibitor (SU11274), etc., have been found to enhance the effects of ACLY inhibitors in small studies of tumors.
Homology
ACLY is a member of the acyl-CoA synthetase superfamily (ADP-forming). ACLYs amino terminal region, 1-419, resembles ATP citrate (pro-S)-lyase and the region, 1-424, is homologous to the β-subunit of succinyl-CoA synthetase and the region, 486-818, is homologous to the α-subunit of succinyl-CoA synthetase. Also, the hierarchy of multiple, sequential serine/threonine phosphorylations responsible for the complex regulation of glycogen synthase is similar to the serine/threonine phosphorylations in ACLY. Sequence surrounding the histidine in ACLYs catalytic site, that is phosphorylated by nucleoside diphosphate kinase (NDPK or nm23), is similar to sequence around phosphorylation sites in other substrates of nm23, such as aldolase C. ACLY has homology with citrate synthase that catalyzes its reverse reaction. Rat ACLY is 96,3% identical to human ACLY.
Mutations
Germinal
Homozygous knock-out of ACLY in mice is lethal. Heterozygous knock-out mice appear to be normal.
Implicated in
Entity name
Bladder (transitional cell) cancer
Note
A bladder cancer cell line (MBT-2) studied in a mouse syngenic cancer model has demonstrated efficacy of calcium hydrocitrate when it was used to inhibit ACLY, combined with other drugs and agents, in several small studies.
Entity name
Breast cancer
Note
Increased expression of ACLY may play a role in the agressive breast cancers. Elevated levels were found in both primary and metastatic cell lines compared to normal cell lines and the highest expression levels occurred in metastatic cell lines.
Entity name
Colon carcinoma
Note
Silencing ACLY in human colon carcinoma cells (HCT116) has been shown to suppress histone acetylation.
Entity name
Gliomas (glial brain tumors)
Note
ACLY has been demonstrated to localize preferentially to pseudopodia in U87 human glioblastoma cells. Inhibition of ACLY in U87 cells with a soluble form of hydroxycitrate suppressed their cell migration, clonogenicity and brain invasion under glycolytic conditions and enhanced the suppressive effects of a Met kinase inhibitor on cell migration. Queries of the NIHs REMBRANDT brain tumor database based on Affymetrix array data indicated that decreased patient survival correlated with increased gene expression of ACLY in gliomas.
Entity name
Liver (hepatocellular) carcinoma
Note
Markedly increased expression of mRNA for ACLY and genes for other lipogenic enzymes have been found in hepatocellular carcinoma compared to surrounding non-cancerous liver tissue.
Entity name
Lung carcinoma
Note
A Lewis lung cancer cell line (LL/2) studied in a mouse syngenic cancer model has demonstrated efficacy of using calcium hydrocitrate combined with other drugs and agents in several small studies. In one of these studies, results were confirmed using a human xenograft model, NCI-H69, small cell lung carcinoma, with tumor development reduced and prolonged animal survival observed. In another study with ACLY knockdown, there was inhibition of growth in vivo for non-small cell lung carcinoma along with apoptosis and differentiation. Enhancement of the anti-tumor effects was achieved by adding statins with regression of established tumors reported. Human lung adenocarcinoma samples have been shown to have significantly increased ACLY activity compared to normal lung tissue and phosphorylated ACLY overexpression correlated with stage, grade, and poorer prognosis. Growth arrest in A549 cells was achieved with RNA interference for ACLY. Inhibitory results were also achieved in A549 cells with the ACLY inhibitor, SB-204990.
Entity name
Melanoma
Note
A melanoma cell line (B16-F10) studied in a mouse syngenic cancer model has demonstrated efficacy of using calcium hydrocitrate combined with other drugs and agents in several small studies.
Entity name
Ovarian carcinoma
Note
Higher ACLY expression has been found in malignant ovarian tissue compared to normal ovarian tissue. Phosphorylated ACLY was also increased and the expression correlated well with tumor grade, FIGO stage, and poorer prognosis. Also knockdown of ACLY in A2780 cells inhibited their proliferation and induced cell cycle arrest.
Entity name
Pancreatic cancer (ductal adenocarcinoma)
Note
An 80 year old woman was treated with a formulation containing hydroxycitrate to inhibit ACLY in addition to gemcitabine with favorable temporary results.
Entity name
Prostate carcinoma
Note
Aberrant expression of ACLY has been found in prostatic cancer with levels inversely correlating with tumor stage and differentiaion. The expression of ACLY has predicted a reduced citrate level which is characteristic of prostatic cancer. Normal prostatic tissue has very high levels of citrate. Benign prostatic hypertrophy also has high levels of citrate. The change to oxidation of citrate in prostatic cancer rather than production of citrate has been viewed as a type of metabolic transformation that may provide a bioenergetic theory for prostatic malignancy.
Entity name
Hepatitis B Virus (HBV) infection
Note
In HBV transgenic mice that replicate HBV in the liver without producing gross liver pathology, the largest functional category for upregulated genes was lipid biosynthesis, including ACLY.
Entity name
Obesity/fatty liver
Note
Inhibition of ACLY is a strategy to counteract weight gain that has led to the development of commercially available formulations of hydroxycitrate. Inhibition of ACLY has been suggested as being helpfull for fatty liver.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 21655919 | 2012 | Screening of well-established drugs targeting cancer metabolism: reproducibility of the efficacy of a highly effective drug combination in mice. | Abolhassani M et al |
| 19795461 | 2010 | Identification of ATP citrate lyase as a positive regulator of glycolytic function in glioblastomas. | Beckner ME et al |
| 14662765 | 2004 | ATP-citrate lyase deficiency in the mouse. | Beigneux AP et al |
| 22510345 | 2012 | Changes in gene transcription underlying the aberrant citrate and choline metabolism in human prostate cancer samples. | Bertilsson H et al |
| 7520580 | 1994 | Bioenergetic theory of prostate malignancy. | Costello LC et al |
| 8088842 | 1994 | Localization of the gene for ATP citrate lyase (ACLY) distal to gastrin(GAS) and proximal to D17S856 on chromosome 17q12-q21. | Couch FJ et al |
| 19968906 | 2010 | Plenary Lecture 2: Transcription factors, regulatory elements and nutrient-gene communication. | Cousins RJ et al |
| 1371749 | 1992 | Cloning and expression of a human ATP-citrate lyase cDNA. | Elshourbagy NA et al |
| 1458840 | 1992 | Epidermal growth factor (EGF) stimulation of ATP citrate lyase activity in isolated rat hepatocytes is age dependent. | Emmerson K et al |
| 15608338 | 2005 | Reverse genetic characterization of cytosolic acetyl-CoA generation by ATP-citrate lyase in Arabidopsis. | Fatland BL et al |
| 8603738 | 1996 | Insulin- and polyunsaturated fatty acid-responsive region(s) of rat ATP citrate lyase gene promoter. | Fukuda H et al |
| 20931262 | 2012 | Adding a combination of hydroxycitrate and lipoic acid (METABLOC™) to chemotherapy improves effectiveness against tumor development: experimental results and case report. | Guais A et al |
| 16032730 | 2005 | cDNA microarray analysis of HBV transgenic mouse liver identifies genes in lipid biosynthetic and growth control pathways affected by HBV. | Hajjou M et al |
| 21688263 | 2012 | Inhibition of lung cancer growth: ATP citrate lyase knockdown and statin treatment leads to dual blockade of mitogen-activated protein kinase (MAPK) and phosphatidylinositol-3-kinase (PI3K)/AKT pathways. | Hanai J et al |
| 16226706 | 2005 | ATP citrate lyase inhibition can suppress tumor cell growth. | Hatzivassiliou G et al |
| 7417478 | 1980 | Properties and organ distribution of ATP citrate (pro-3S)-lyase. | Hoffmann GE et al |
| 2857659 | 1985 | Both insulin and epidermal growth factor stimulate fatty acid synthesis and increase phosphorylation of acetyl-CoA carboxylase and ATP-citrate lyase in isolated hepatocytes. | Holland R et al |
| 3871637 | 1985 | Effects of phosphorylation on the kinetic properties of rat liver ATP-citrate lyase. | Houston B et al |
| 1332698 | 1992 | Identification of multifunctional ATP-citrate lyase kinase as the alpha-isoform of glycogen synthase kinase-3. | Hughes K et al |
| 6301829 | 1983 | The protein phosphatases involved in cellular regulation. 6. Measurement of type-1 and type-2 protein phosphatases in extracts of mammalian tissues; an assessment of their physiological roles. | Ingebritsen TS et al |
| 8941657 | 1996 | Adenosine triphosphate citrate lyase mediates hypocitraturia in rats. | Melnick JZ et al |
| 22101433 | 2011 | Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. | Metallo CM et al |
| 18922930 | 2008 | ATP citrate lyase: activation and therapeutic implications in non-small cell lung cancer. | Migita T et al |
| 22101431 | 2011 | Reductive carboxylation supports growth in tumour cells with defective mitochondria. | Mullen AR et al |
| 9693110 | 1998 | The role of ATP citrate-lyase in the metabolic regulation of plasma lipids. Hypolipidaemic effects of SB-204990, a lactone prodrug of the potent ATP citrate-lyase inhibitor SB-201076. | Pearce NJ et al |
| 10653665 | 2000 | Phosphorylation of recombinant human ATP:citrate lyase by cAMP-dependent protein kinase abolishes homotropic allosteric regulation of the enzyme by citrate and increases the enzyme activity. Allosteric activation of ATP:citrate lyase by phosphorylated sugars. | Potapova IA et al |
| 2176822 | 1990 | Sequence of sites on ATP-citrate lyase and phosphatase inhibitor 2 phosphorylated by multifunctional protein kinase (a glycogen synthase kinase 3 like kinase). | Ramakrishna S et al |
| 7054176 | 1982 | Phosphorylation of dephospho-ATP citrate lyase by the catalytic subunit of cAMP-dependent protein kinase. | Ranganathan NS et al |
| 15200237 | 2004 | Body weight and abdominal fat gene expression profile in response to a novel hydroxycitric acid-based dietary supplement. | Roy S et al |
| 16388597 | 2006 | A new strategy for studying protein kinase B and its three isoforms. Role of protein kinase B in phosphorylating glycogen synthase kinase-3, tuberin, WNK1, and ATP citrate lyase. | Sale EM et al |
| 22797854 | 2013 | Tumor regression with a combination of drugs interfering with the tumor metabolism: efficacy of hydroxycitrate, lipoic acid and capsaicin. | Schwartz L et al |
| 15234082 | 2004 | Safety assessment of (-)-hydroxycitric acid and Super CitriMax, a novel calcium/potassium salt. | Soni MG et al |
| 20946014 | 2010 | Safety assessment of a calcium-potassium salt of (-)-hydroxycitric acid. | Stohs SJ et al |
| 22102020 | 2011 | ADP-Mg2+ bound to the ATP-grasp domain of ATP-citrate lyase. | Sun T et al |
| 7062046 | 1982 | The contribution of citrate to the synthesis of acetyl units in synaptosomes of developing rat brain. | Szutowicz A et al |
| 10698688 | 2000 | Histidine to aspartate phosphotransferase activity of nm23 proteins: phosphorylation of aldolase C on Asp-319. | Wagner PD et al |
| 22266777 | 2012 | Prognostic and therapeutic implications of increased ATP citrate lyase expression in human epithelial ovarian cancer. | Wang Y et al |
| 19461003 | 2009 | ATP-citrate lyase links cellular metabolism to histone acetylation. | Wellen KE et al |
| 22106302 | 2011 | Hypoxia promotes isocitrate dehydrogenase-dependent carboxylation of α-ketoglutarate to citrate to support cell growth and viability. | Wise DR et al |
| 15869874 | 2005 | Co-ordinate activation of lipogenic enzymes in hepatocellular carcinoma. | Yahagi N et al |
| 17472751 | 2007 | Metastatic progression and gene expression between breast cancer cell lines from African American and Caucasian women. | Yancy HF et al |
Other Information
Locus ID:
NCBI: 47
MIM: 108728
HGNC: 115
Ensembl: ENSG00000131473
Variants:
dbSNP: 47
ClinVar: 47
TCGA: ENSG00000131473
COSMIC: ACLY
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38038390 | 2024 | Cutting Edge: STING Induces ACLY Activation and Metabolic Adaptations in Human Macrophages through TBK1. | 2 |
| 38176897 | 2024 | ATP citrate lyase (ACLY)-dependent immunometabolism in mucosal T cells drives experimental colitis in vivo. | 0 |
| 38494608 | 2024 | Enhanced lipid biosynthesis in oral squamous cell carcinoma cancer-associated fibroblasts contributes to tumor progression: Role of IL8/AKT/p-ACLY axis. | 1 |
| 38670440 | 2024 | CILP2 promotes hypertrophic scar through Snail acetylation by interaction with ACLY. | 0 |
| 38038390 | 2024 | Cutting Edge: STING Induces ACLY Activation and Metabolic Adaptations in Human Macrophages through TBK1. | 2 |
| 38176897 | 2024 | ATP citrate lyase (ACLY)-dependent immunometabolism in mucosal T cells drives experimental colitis in vivo. | 0 |
| 38494608 | 2024 | Enhanced lipid biosynthesis in oral squamous cell carcinoma cancer-associated fibroblasts contributes to tumor progression: Role of IL8/AKT/p-ACLY axis. | 1 |
| 38670440 | 2024 | CILP2 promotes hypertrophic scar through Snail acetylation by interaction with ACLY. | 0 |
| 35404187 | 2023 | Selective autophagic degradation of ACLY (ATP citrate lyase) maintains citrate homeostasis and promotes oocyte maturation. | 7 |
| 37076498 | 2023 | Allosteric role of the citrate synthase homology domain of ATP citrate lyase. | 0 |
| 37122003 | 2023 | Activation of ACLY by SEC63 deploys metabolic reprogramming to facilitate hepatocellular carcinoma metastasis upon endoplasmic reticulum stress. | 7 |
| 37140234 | 2023 | ACLY-induced reprogramming of glycolytic metabolism plays an important role in the progression of breast cancer. | 0 |
| 37620891 | 2023 | ACLY as a modulator of liver cell functions and its role in Metabolic Dysfunction-Associated Steatohepatitis. | 1 |
| 37801083 | 2023 | BCAT2 promotes melanoma progression by activating lipogenesis via the epigenetic regulation of FASN and ACLY expressions. | 0 |
| 37927015 | 2023 | [Expression and Clinical Significance of ATP Citrate Lyase in Hepatocellular Carcinoma]. | 0 |
Citation
Marie E Beckner
ACLY (ATP citrate lyase)
Atlas Genet Cytogenet Oncol Haematol. 2012-10-01
Online version: http://atlasgeneticsoncology.org/gene/50486/acly
