CD200 (CD200 molecule)
2008-12-01 Lesley Mathews , William Farrar AffiliationCancer Stem Cell Section, Laboratory of Cancer Prevention, Center for Cancer Research, National Cancer Institute at Frederick, Frederick, MD 21702, USA
DNA/RNA
Description
29744 bp
Transcription
Proteins
Description
The CD200 molecule is a single-pass type I membrane protein spanning the membrane once, with its N-terminus on the extracellular side of the membrane and subsequent processing results in removal of its signal sequence. Variant 1 produces a protein that is 269 amino acids in length, variant 2 yields a protein that is 294 amino acids, and variant 3 has yet to be experimentally validated for its product. CD200 is a highly conserved member of the immunoglobulin superfamily containing both an Ig-like C2-type (immunoglobulin-like) domain as well as Ig-like V-type (immunoglobulin-like) domain.
Expression
CD200 is commonly expressed on cells originating from the hematopoietic cells, specifically those of the myeloid lineage such as macrophages, dendritic cells, neutrophils, mast cells, and eosinophils, yet it can also be expressed on B cells, activated T cells, endothelial neuronal cells and cells of the reproductive organs (ovaries and placental trophoblasts). This molecule exerts its effect by binding to its cognate receptor CD200R (also on chromosome 3 and containing two Ig-like domains) which is expressed on cells specifically derived from the myeloid lineage and certain populations of T cells. This interaction can only occur when the cell carrying the receptor and the cell expressing CD200 are in close contact to one another. Unlike the CD200 molecule itself, the CD200R has 67 amino acids in its cytoplasmic domain compared to 19 for CD200, and it contains tyrosine residues that can be phosphorylated to carry on further signal transduction. A recent review has further summarized the tissue distribution of CD200 and its receptor in rat, mouse and human samples.
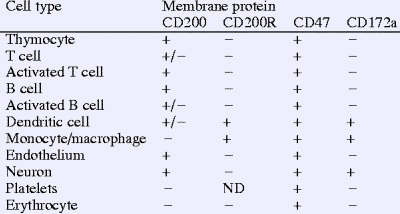

CD200 and membrane protein interactions in the control of myeloid cells.
Neil Barclay, Gavin J. Wrightb, Gary Brooke and Marion H. Brown. Trends in Immunology. Volume 23, Issue 6, 1 June 2002, Pages 285-290.
Localisation
Single-pass type I plasma membrane protein or plasma membrane associated protein.
Function
The function of CD200 was orginally identified using mice lacking the CD200 gene where the mice developed an earlier onset of disease in experimental models of both autoimmune encephalomyelitis (EAE), collagen-induced arthritis (CIA), and autoimmune alopecia. The mice have increased numbers of macrophages in the spleen and after immunological stimulation, these macrophages demonstate a more active phenoype in vivo. These studies demonstrate the importance of CD200 in maintaining self-tolerance and further studies have demonstrated that CD200 acts as an important molecule in controlling autoimmunity, inflammation and adaptive immune responses implicating it as a key immunosuppressive molecule. The CD200-CD200R interaction exerts its inhibitory signal to myeloid cells that suppresses INF (Interferon)-gamma-induced proliferation, activation and secretion of nitrous oxide (NO), IL5, IL13 and IL6. Interestingly, a few phylogenetically distinct viruses including poxviruses and KSHV, the agent that causes Kaposi sarcoma and other human cancers, have been shown to attenuate the host antiviral immune response by expressing a viral CD200 homolog.

Homology
| % Identity | ||||
| Homo sapiens | Protein | DNA | ||
| CD200 | vs. Pan troglodytes (Common Chimpanzee) | CD200 | 99.3 | 99.4 |
| vs. Canis lupus familiaris (Domestic Dog) | CD200 | 77.1 | 82.7 | |
| vs. Bos taurus (Domestic Cow) | CD200 | 80.2 | 84.8 | |
| vs. Mus musculus (The House Mouse) | CD200 | 77.6 | 81.7 | |
| vs. Rattus norvegicus (Norway Rat) | CD200 | 77.2 | 80.7 | |
| vs. Gallus gallus (Domesticated Fowl) | CD200 | 36.8 | 51.0 | |
Mutations
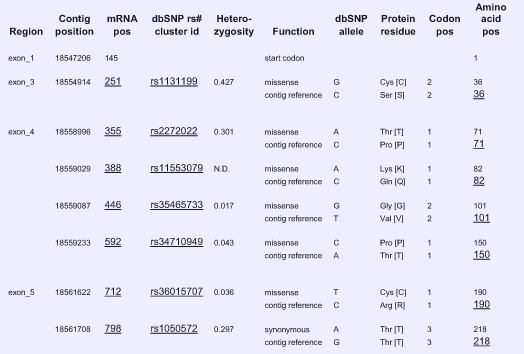
Germinal
None noted.
Somatic
Restricted CD200R isoform expression at the feto-maternal interface suggests CD200-CD200R interactions may serve important function(s) determining the successful outcome of pregnancy.
Implicated in
Entity name
Cancer
Note
The expression of CD200 has been found in expressed several cancer cell lines or tissues, including ovarian, melanoma, head and neck carcinoma, testicular, malignant mesothelioma, neuroblastoma, renal cell carcinoma, chronic lymphocytic leukemia, prostate, breast and colon cancer. In multiple myeloma (MM) and acute myeloid leukemia (AML) CD200 has been implicated as a prognostic factor. In MM patients, CD200 was expressed in 78% of samples analyzed and for AML patients, in 43% of the patient samples. Patients who demonstrated expression of CD200 on their leukemic cells also demonstrated a decrease in survival overall. With regards to cancer progression, expression of CD200 is associated with metastasis or a relapse in bladder, lung, CML (chronic myeloid leukemia), breast, melanoma and prostate cancers. The expression of CD200 on cancer cells has been linked to the theory of tumor immunity since mice treated wtih a CD200-Fc fusion protein (which blocks function of CD200) were able to clear injected leukemic, cells unlike mice who did not receive the antibody.
Entity name
Cancer Stem Cells
Note
A relatively new hypothesis about cancer progression has suggested that cancer stem cells (CSCs) mediate solid tumor initiation and progression. The theory suggests that CSCs are only a minor subpopulation of cells (generally
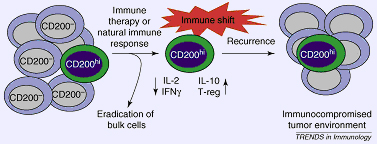
A hypothetical model of how CD200 might be involved in cancer stem cell (CSC) immunoevasion. CD200-expressing CSCs survive immune therapy or a natural immune response and induce a shift away from a Th1 response. These immunosuppressive effects of CD200+ CSCs could be mediated by events such as deviations in cytokine production, increased proliferation and/or induction of regulatory T cells (Treg). The altered immune response to the tumor might indefinitely dampen future immune reactions to the tumor, thus resulting in tumor cells that are resistant and free to proliferate in an immunocompromised tumor microenvironment.
Kawasaki, B.T. et al. (2008) Cancer stem cells, CD200 and immunoevasion. Trends in Iummunology Vol. 29 No. 10 (464-468).
Entity name
Experimental autoimmune encephalomyelitis (EAE)
Note
EAE is a model of the human disease multiple sclerosis, which results from activation of peripheral T lymphocytes, macrophages, and granulocytes. These activated cells migrate to the Central Nervous System and can lead to microglial activation, tissue damage, and neurological deficits including paralysis. In these EAE mice, loss of CD200 resulted in an increase of disease onset and progression compared to mice expressing the gene. Using cultured rat hippocampal neurons, it was found that if the CD200-CD200R signalling pathway is blocked using an anti-CD200R-Fc antibody, the INF-gamma induced IL6 response of macrophages is increased and macrophage-mediated cell death of the neurons occured.
Entity name
Collagen-induced arthritis (CIA)
Note
CIA is a model of human rheumatoid arthritis which is an inflammatory autoimmune disease of the joints, and like EAE involves tissue-specific influx of T cells, macrophages, and granulocytes. Disruption of CD200-CD200 receptor interaction using cd200 KO mice increased susceptibility to mice normally resistant to this disease. The animals developed moderate to severe arthritis with synovial inflammation and formation of invasive pannus which results in degradation of cartilage and bone.
Entity name
Skin Graft Rejection/Transplantation
Note
Experiments from mice have demonstrated that when wild-type tail skin from a male mouse was grafted onto the flank of a strain matched female, the skin was rejected at 59 days. However, if the male skin was CD200-deficient, it was only rejected after only 25 days.
Entity name
Alopecia
Note
When expression of CD200 is lost from follicular kertainocytes the CD200R-positive cells (Langerhans cells and dendritic cells) switch to having an activated phenotype. Any additional trauma to the skin can initate a full activation of CD200 positive cells and cause folliculocentric inflammation. The chronic inflammation that ensues leads to recruitment and activation of autoreactive T cells which are specific for hair follicles and lead to their destruction.
Entity name
Autoimmune uveoretinitis (EAU)
Note
EAU is an organ-specific autoimmunity in the eye, a tissue with extensive neuronal and endothelial CD200 expression.One experiment demonstrated that mice lacking CD200 expression had increased numbers of retina-infiltrating macrophages. In addition, these mice displayed an increase in nitric oxide responses. When DX109 (a monoclonal antibody to CD200) was administered, it suppressed EAU despite maintained T-cell proliferation and IFNgamma production and resulted in an earlier resolution of disease. Thus promoting CD200R-mediated signaling can successfully prevent full expression of IFNgamma-mediated macrophage activation and protect against tissue damage during autoimmune responses.
Entity name
Recurrent Abortion/Pre-Eclampsia
Note
A number of immunoregulatory cells have been shown to help prevent abortions and/or play a role in pre-eclampsia. For CD200, when male lymyphocytes from aborting mice were immunized (inactivated), this stimulated female gdT CD4 cells to secrete Th2 cytokines via a CD200-dependent mechanism and reversed the abortive affects.
Entity name
Microglia activation and Parkinsons Disease (PD)
Note
A number of studies have demonstrated that CD200 down-regulates microglia activation through binding to CD200R. Microglia are the primary immune cells in the brain and play an important role in the progression of PD. Similar to true myleoid cells, microglia in CD200-deficient mice exhibited more activated characterisitics, such as shorter glial processes and less of a disordered arrangement, as well as an increase in inducible nitric oxide synthetase (specific to immune cells). To further support this, Frank et al. in 2006 demonstrated that expression of CD200 mRNA (measured by PCR) in the hippocampus decreased with increase in aging. Thus, the CD200-CD200R signaling pathway should be considered as target for developing therapies against PD.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 12072366 | 2002 | CD200 and membrane protein interactions in the control of myeloid cells. | Barclay AN et al |
| 15890435 | 2006 | mRNA up-regulation of MHC II and pivotal pro-inflammatory genes in normal brain aging. | Frank MG et al |
| 15274657 | 2004 | Structural and functional heterogeneity in the CD200R family of immunoregulatory molecules and their expression at the feto-maternal interface. | Gorczynski RM et al |
| 11099416 | 2000 | Down-regulation of the macrophage lineage through interaction with OX2 (CD200). | Hoek RM et al |
| 18775673 | 2008 | Cancer stem cells, CD200 and immunoevasion. | Kawasaki BT et al |
| 17645413 | 2007 | The biology of cancer stem cells. | Lobo NA et al |
| 18164423 | 2008 | CNS inflammation and neuronal degeneration is aggravated by impaired CD200-CD200R-mediated macrophage silencing. | Meuth SG et al |
| 17984999 | 2008 | Prostate cell cultures as in vitro models for the study of normal stem cells and cancer stem cells. | Miki J et al |
| 16946299 | 2006 | CD200 is a new prognostic factor in multiple myeloma. | Moreaux J et al |
| 18060862 | 2008 | CD200: a putative therapeutic target in cancer. | Moreaux J et al |
| 16386879 | 2006 | CD200, a "no danger" signal for hair follicles. | Rosenblum MD et al |
| 18060403 | 2008 | Immune modulation by melanoma and ovarian tumor cells through expression of the immunosuppressive molecule CD200. | Siva A et al |
| 17252007 | 2007 | CD200 as a prognostic factor in acute myeloid leukaemia. | Tonks A et al |
| 18040859 | 2007 | CD200-CD200R regulation of microglia activation in the pathogenesis of Parkinson's disease. | Wang XJ et al |
| 16829304 | 2006 | Immunological analogy between allograft rejection, recurrent abortion and pre-eclampsia - the same basic mechanism? | Wilczyński JR et al |
Other Information
Locus ID:
NCBI: 4345
MIM: 155970
HGNC: 7203
Ensembl: ENSG00000091972
Variants:
dbSNP: 4345
ClinVar: 4345
TCGA: ENSG00000091972
COSMIC: CD200
RNA/Proteins
Expression (GTEx)
Pathways
| Pathway | Source | External ID |
|---|---|---|
| Immune System | REACTOME | R-HSA-168256 |
| Adaptive Immune System | REACTOME | R-HSA-1280218 |
| Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell | REACTOME | R-HSA-198933 |
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37616575 | 2024 | Exploiting the CD200-CD200R immune checkpoint axis in multiple myeloma to enhance CAR T-cell therapy. | 1 |
| 37801139 | 2024 | Integrating TSPO PET imaging and transcriptomics to unveil the role of neuroinflammation and amyloid-β deposition in Alzheimer's disease. | 3 |
| 38455039 | 2024 | CD200 genotype is associated with clinical outcome of patients with multiple myeloma. | 0 |
| 38619621 | 2024 | CD200 is overexpressed in the pancreatic tumor microenvironment and predictive of overall survival. | 0 |
| 37616575 | 2024 | Exploiting the CD200-CD200R immune checkpoint axis in multiple myeloma to enhance CAR T-cell therapy. | 1 |
| 37801139 | 2024 | Integrating TSPO PET imaging and transcriptomics to unveil the role of neuroinflammation and amyloid-β deposition in Alzheimer's disease. | 3 |
| 38455039 | 2024 | CD200 genotype is associated with clinical outcome of patients with multiple myeloma. | 0 |
| 38619621 | 2024 | CD200 is overexpressed in the pancreatic tumor microenvironment and predictive of overall survival. | 0 |
| 36738455 | 2023 | The immunoregulatory protein CD200 as a potentially lucrative yet elusive target for cancer therapy. | 5 |
| 37894750 | 2023 | Emerging Immune Checkpoint Molecules on Cancer Cells: CD24 and CD200. | 4 |
| 36738455 | 2023 | The immunoregulatory protein CD200 as a potentially lucrative yet elusive target for cancer therapy. | 5 |
| 37894750 | 2023 | Emerging Immune Checkpoint Molecules on Cancer Cells: CD24 and CD200. | 4 |
| 35326506 | 2022 | The Role of CD200-CD200 Receptor in Human Blood and Lymphatic Endothelial Cells in the Regulation of Skin Tissue Inflammation. | 3 |
| 35326506 | 2022 | The Role of CD200-CD200 Receptor in Human Blood and Lymphatic Endothelial Cells in the Regulation of Skin Tissue Inflammation. | 3 |
| 33015886 | 2021 | Association of CD200 expression in paternal lymphocytes with female Th1/Th2 balance and pregnancy establishment at immunotherapy of recurrent spontaneous abortion. | 3 |
Citation
Lesley Mathews ; William Farrar
CD200 (CD200 molecule)
Atlas Genet Cytogenet Oncol Haematol. 2008-12-01
Online version: http://atlasgeneticsoncology.org/gene/44381/cd200

