ENAH (enabled homolog (Drosophila))
2008-08-01 Paola Nisticò , Francesca Di Modugno AffiliationRegina Elena Cancer Institute, Department of Experimental Oncology, via delle Messi dOro 156, 00158 Rome, Italy
Identity
HGNC
LOCATION
1q42.12
LOCUSID
ALIAS
ENA,MENA,NDPP1
FUSION GENES
DNA/RNA
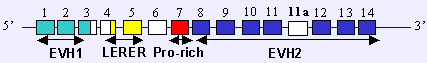
Diagrammatic representation of human ENAH gene trancripts. Exons are enumerated and the relative protein domains are indicated.
Description
The human ENAH gene is located on the minus strand of chromosome 1 and is constituted by 15 exons. Other features of the ENAH gene such as promoter or enhancer have not been fully investigated.
Transcription
Two alternative splice variants isolated in the human ENAH. The size of the longer variant (hMena+11a or ENAH variant 1) is 1776 nt. The shorter variant (hMena or ENAH variant 2) lacks an internal exon (exon 11a) of 63nt.
Proteins
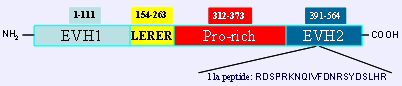
Human ENAH protein, the four conserved domains are indicated with the respective amino acid positions. The 11a peptide is included at position 513 and characterizes the ENAH isoform a (hMena+11a).
Description
Mena is a member of Ena/VASP proteins characterized by the presence of specific domains including the NH2-terminal EVH1 domain, which plays a role in intracellular protein localization (Prehoda et al., 1999) and interacts with proteins bearing FPPPP motifs. Among the Ena/VASP proteins only the EVH1 domain of Mena possesses the ability to bind to the LIM3 domain of the oncosuppressor TES (Boeda et al., 2007). The central proline-rich domain mediates the interaction with proteins containing the SH3 and WW domains and with the small actin monomer binding protein profilin (Gertler et al., 1996). The LERER region is constituted by a long repeat of Arg/Leu/Glu amino acids probably acting as a protein protein binding interface, located between the EVH1 and the prol-rich region. Only Mena among the Ena/VASP proteins possesses this domain. The EVH2 COOH-terminal domain, which forms a right handed alpha helical coiled coil structure, is responsible for tetramerization and for the binding to G- and F-actin (Kuhnel et al., 2004); its interaction with the growing ends of the actin filaments is required for targeting the Ena/VASP to lamellipodia and filopodia (Louriero et al., 2002). Human Mena (hMena or ENAH isoform b) is a 570 amino acid protein. The longer hMena+11a isoform (ENAH isoform a) presents an additional internal peptide of 21aa located in the EVH2 domain of the protein. This isoform undergoes phosphorylation upon treatment of breast cancer cell lines with EGF and NRG1 (Di Modugno et al., 2007).
Mena is alternately spliced to give rise to multiple isoforms, an additional reported isoform is the neuronal specific Mena-140 found in mouse and humans (Gertler et al., 1996; Urbanelli et al., 2006).
Mena is alternately spliced to give rise to multiple isoforms, an additional reported isoform is the neuronal specific Mena-140 found in mouse and humans (Gertler et al., 1996; Urbanelli et al., 2006).
Expression
In normal tissues, hMena expression was confined to isolated epithelia (i.e.pancreas). Mammary epithelium was negative and hMena was overexpressed in about 75% of breast primary tumors tested, with a variable staining intensity.
Localisation
Predominantly in cytoplasm and in some tumor cells with a reinforced juxtamembrane staining.
Function
Mena controls cell shape and movement (Bear et al., 2002; Vasioukhin et al., 2000; Krause et al., 2003) by protecting actin filaments from capping proteins at their barbed ends (Barzik et al., 2005). It controls actin organization on cadherin adhesion contact (Scott et al., 2006).
Homology
The sequence of hMena (ENAH isoform b) displays 87% identity with the murine protein but is longer with the majority of the additional aminoacids located in the Arg/Leu/Glu rich region (LERER). The human hMena sequence conserved the two serine phosphorylation sites of murine Mena, whereas the tyrosine residue, site of phosphorylation in mouse Mena (Tani et al., 2003), is substituted by a glutamine residue in the human sequence.
Implicated in
Entity name
Breast Cancer
Disease
In human tissues, human ENAH (hMena) protein, not expressed in normal breast, is detectable in a small percentage of low-risk proliferative lesions, with a progressive increase of positivity in benign breast lesions at higher risk of transformation and in in situ and invasive cancers. In the latter, a significant direct correlation was found between hMena, tumor size, proliferation index, and HER-2 overexpression whereas an inverse relationship was evidenced with estrogen receptor (ER) and progesterone receptor (PgR) expression (Di Modugno et al., 2006).
These results suggest that hMena may be a marker of breast cancerogenesis and breast cancer progression.
In cancer cell lines of different histological origin, hMena is overexpressed respect to the normal counterparts (i.e. breast, melanoma, colon, cervical cancer). hMena expression while up-regulated by Neuregulin-1 and EGF, is down-regulated by Herceptin treatment in breast cancer cell lines, thus suggesting that hMena couples tyrosine kinase receptor (TKR) signaling to the actin cytoskeleton.
hMena+11a isoform (ENAH isoform a) is epithelial-specific and is phosphorylated after mitogenic stimuli, such as EGF. This phosphorylation is accompanied by an increased proliferation rate and p42 / 44 MAPK activation in breast cancer cell lines (Di Modugno et al., 2007), thus suggesting a functional role of hMena+11a in breast cancer cell proliferation. In a murine model Mena is overexpressed, among a set of genes coding for the minimum motility machine regulating β-actin polymerization, in a subpopulation of invasive breast tumor cells collected using the in vivo invasion assay in response to EGF (Wang et al., 2004). A role of hMena in the invasive behaviour of human tumor cells has not yet been reported.
These results suggest that hMena may be a marker of breast cancerogenesis and breast cancer progression.
In cancer cell lines of different histological origin, hMena is overexpressed respect to the normal counterparts (i.e. breast, melanoma, colon, cervical cancer). hMena expression while up-regulated by Neuregulin-1 and EGF, is down-regulated by Herceptin treatment in breast cancer cell lines, thus suggesting that hMena couples tyrosine kinase receptor (TKR) signaling to the actin cytoskeleton.
hMena+11a isoform (ENAH isoform a) is epithelial-specific and is phosphorylated after mitogenic stimuli, such as EGF. This phosphorylation is accompanied by an increased proliferation rate and p42 / 44 MAPK activation in breast cancer cell lines (Di Modugno et al., 2007), thus suggesting a functional role of hMena+11a in breast cancer cell proliferation. In a murine model Mena is overexpressed, among a set of genes coding for the minimum motility machine regulating β-actin polymerization, in a subpopulation of invasive breast tumor cells collected using the in vivo invasion assay in response to EGF (Wang et al., 2004). A role of hMena in the invasive behaviour of human tumor cells has not yet been reported.
Entity name
Pancreatic Cancer
Disease
hMena is expressed in primary and metastatic pancreatic cancer. The expression of hMena+11a isoform (ENAH isoform a) characterizes pancreatic cancer cell lines with an epithelial phenotype which express the epithelial marker E-Cadherin and lack the expression of mesenchymal markers as N-Cadherin and Vimentin. These cell lines show a constitutive activated EGFR and are sensitive to the treatment with the EGFR inhibitor Erlotinib. ENAH acts as a mediator of the EGFR signaling pathway and significantly modulates the growth of pancreatic cancer cell lines dependent on EGFR signaling. Thus the expression of hMena/hMena+11a is predictive of in vitro response to EGFR inhibitors (Simo et al., 2008).
Entity name
Tumor Immunity
Disease
Human ENAH (hMena) protein is able to induce a cancer-restricted antibody response. Twenty percent of cancer patient sera, showed anti-hMena-specific IgG, while no antibodies were present in healthy donors (Di Modugno et al., 2004).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15939738 | 2005 | Ena/VASP proteins enhance actin polymerization in the presence of barbed end capping proteins. | Barzik M et al |
| 12086607 | 2002 | Antagonism between Ena/VASP proteins and actin filament capping regulates fibroblast motility. | Bear JE et al |
| 18158903 | 2007 | Tes, a specific Mena interacting partner, breaks the rules for EVH1 binding. | Boëda B et al |
| 17363586 | 2007 | Molecular cloning of hMena (ENAH) and its splice variant hMena+11a: epidermal growth factor increases their expression and stimulates hMena+11a phosphorylation in breast cancer cell lines. | Di Modugno F et al |
| 8861907 | 1996 | Mena, a relative of VASP and Drosophila Enabled, is implicated in the control of microfilament dynamics. | Gertler FB et al |
| 14570581 | 2003 | Ena/VASP proteins: regulators of the actin cytoskeleton and cell migration. | Krause M et al |
| 15569942 | 2004 | The VASP tetramerization domain is a right-handed coiled coil based on a 15-residue repeat. | Kühnel K et al |
| 12134088 | 2002 | Critical roles of phosphorylation and actin binding motifs, but not the central proline-rich region, for Ena/vasodilator-stimulated phosphoprotein (VASP) function during cell migration. | Loureiro JJ et al |
| 18676769 | 2008 | Human Mena+11a isoform serves as a marker of epithelial phenotype and sensitivity to epidermal growth factor receptor inhibition in human pancreatic cancer cell lines. | Pino MS et al |
| 10338211 | 1999 | Structure of the enabled/VASP homology 1 domain-peptide complex: a key component in the spatial control of actin assembly. | Prehoda KE et al |
| 16371509 | 2006 | Ena/VASP proteins can regulate distinct modes of actin organization at cadherin-adhesive contacts. | Scott JA et al |
| 12672821 | 2003 | Abl interactor 1 promotes tyrosine 296 phosphorylation of mammalian enabled (Mena) by c-Abl kinase. | Tani K et al |
| 16494957 | 2006 | Characterization of human Enah gene. | Urbanelli L et al |
| 10660044 | 2000 | Directed actin polymerization is the driving force for epithelial cell-cell adhesion. | Vasioukhin V et al |
| 15574765 | 2004 | Identification and testing of a gene expression signature of invasive carcinoma cells within primary mammary tumors. | Wang W et al |
Other Information
Locus ID:
NCBI: 55740
MIM: 609061
HGNC: 18271
Ensembl: ENSG00000154380
Variants:
dbSNP: 55740
ClinVar: 55740
TCGA: ENSG00000154380
COSMIC: ENAH
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38459605 | 2024 | Mammalian enabled protein enhances tamoxifen sensitivity of the hormone receptor-positive breast cancer patients by suppressing the AKT signaling pathway. | 0 |
| 38477606 | 2024 | TRIM67 interacts with ENAH to regulate the apoptosis and autophagy of lung cancer cells. | 0 |
| 38459605 | 2024 | Mammalian enabled protein enhances tamoxifen sensitivity of the hormone receptor-positive breast cancer patients by suppressing the AKT signaling pathway. | 0 |
| 38477606 | 2024 | TRIM67 interacts with ENAH to regulate the apoptosis and autophagy of lung cancer cells. | 0 |
| 36959177 | 2023 | Mena regulates nesprin-2 to control actin-nuclear lamina associations, trans-nuclear membrane signalling and gene expression. | 7 |
| 37027295 | 2023 | Matrix stiffness regulates tumor cell intravasation through expression and ESRP1-mediated alternative splicing of MENA. | 8 |
| 36959177 | 2023 | Mena regulates nesprin-2 to control actin-nuclear lamina associations, trans-nuclear membrane signalling and gene expression. | 7 |
| 37027295 | 2023 | Matrix stiffness regulates tumor cell intravasation through expression and ESRP1-mediated alternative splicing of MENA. | 8 |
| 35030977 | 2022 | Enabled homolog (ENAH) regulated by RNA binding protein splicing factor 3b subunit 4 (SF3B4) exacerbates the proliferation, invasion and migration of hepatocellular carcinoma cells via Notch signaling pathway. | 5 |
| 35254597 | 2022 | MiR-139-5p/ENAH Affects Progression of Hepatocellular Carcinoma Cells. | 1 |
| 35790431 | 2022 | [Screening and identification of key genes ATP1B3 and ENAH in the progression of hepatocellular carcinoma: based on data mining and clinical validation]. | 0 |
| 35030977 | 2022 | Enabled homolog (ENAH) regulated by RNA binding protein splicing factor 3b subunit 4 (SF3B4) exacerbates the proliferation, invasion and migration of hepatocellular carcinoma cells via Notch signaling pathway. | 5 |
| 35254597 | 2022 | MiR-139-5p/ENAH Affects Progression of Hepatocellular Carcinoma Cells. | 1 |
| 35790431 | 2022 | [Screening and identification of key genes ATP1B3 and ENAH in the progression of hepatocellular carcinoma: based on data mining and clinical validation]. | 0 |
| 31323325 | 2020 | Scaffold stiffness influences breast cancer cell invasion via EGFR-linked Mena upregulation and matrix remodeling. | 31 |
Citation
Paola Nisticò ; Francesca Di Modugno
ENAH (enabled homolog (Drosophila))
Atlas Genet Cytogenet Oncol Haematol. 2008-08-01
Online version: http://atlasgeneticsoncology.org/gene/44148/enah
