SEPT9 (MLL septin-like fusion)
2001-08-01 Linda M. Kalikin , Elizabeth M. Petty AffiliationResearch Investigator, Department of Internal Medicine, The University of Michigan, USA
Identity
HGNC
LOCATION
17q25.2
LOCUSID
ALIAS
AF17q25,MSF,MSF1,NAPB,PNUTL4,SEPT9,SINT1,SeptD1
FUSION GENES
DNA/RNA
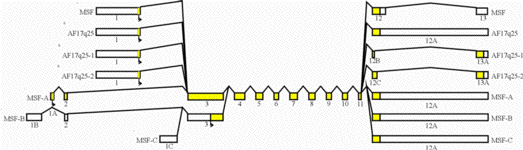
Genomic structure of published MSF alternatively spliced transcripts. Boxes indicate exons with coding regions colored in yellow and are drawn to scale. Exons are tentatively positioned in relative genomic order with overlapping exons indicating identical sequences. Translation start sites are indicated by an arrows and proceeds centromere to telomere on 17q25.
Transcription
MSF exhibits 5 and 3 alternative splicing. The variable exons encode different translational start and stop sequences and are spliced on to a core of 8 coding exons. It is unclear the splicing mechanism between exon 12 and exon 13 subsets as splicing does not occur at traditional GT/AG conserved sequences. Multiple reports are consistent in observing expression of an approximately 4.0 kb transcript in all fetal and adult tissues. Additional transcripts at approximately 3.0 kb and 1.7 kb are variably reported and may reflect differences in probe sequence and experimental conditions between laboratories. MSF spans approximately 260 kb of DNA based on NotI PFGE mapping.
Proteins
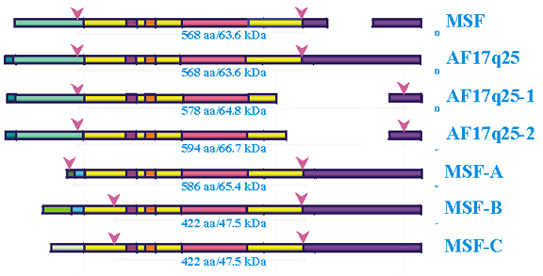
Structure of MSF protein isoforms. Coding regions are indicated between red arrows. Like colors denote identical sequences. Domains are marked by light purple boxes for xylose isomerase, orange for polybasic, and red for GTPase. Gaps indicate missing sequences in that variant. References for a-c match those in the exon/intron figure. Figure is not drawn to scale.
Description
Alternative transcript splicing results in translation of multiple MSF isoforms with distinct amino-and carboxy-termini. MSF-B and MSF-C proteins are identical and are sub-sequences within the other larger isoforms. All isoforms contain a GTPase domain, a xylose isomerase domain of unknown mammalian function but previously identified for sugar interconversion in some microorganisms, and a semi-conserved polybasic domain shown in the septin H5 to be necessary for membrane phospholipid binding.
Expression
The MSF protein is believed to be widely expressed based on ubiquitous adult and fetal transcript expression, although individual isoforms may have tissue specific expression.
Localisation
The MSF protein, like other septin family members, contains no subcellular localization signals and is thought to be largely cytoplasmic. Despite this, mammalian septins have been found in the mitochondria, in the nucleus, and associated with the plasma membrane and brain synaptic vesicles.
Function
Although little information is currently available for MSF itself, research on other members of this highly conserved septin GTPase subfamily provides insight into potential functional roles of MSF. The original family members, Saccharomyces cerevisiae CDC3, CDC10, CDC11, and CDC12, were identified by rescue of temperature sensitive mutants exhibiting the near identical phenotypes of cell-cycle arrest, elongated bud growth, and impaired cytokinesis. They localize to the cleavage furrow, co-immunoprecipitate, and polymerize into filaments. Septins have since been broadly identified in most eukaryotic organisms except for plants. Despite the distinct mechanistic differences in cell division between yeast and animal cells, animal septins similarly localize to the contractile ring and polymerize into filaments. These filaments are composed of homo- and heteromultimers of septins, require GTP hydrolysis to assemble, and interact with anillin, an actin binding protein found at the contractile ring during cytokinesis. Like yeast, multinucleated cells are produced when septins are mutated. Isolation of the yeast septins SPR3 and SPR28 with roles in sporulation, however, provided the initial evidence for other septin functions in addition to cytokinesis. Binding of Cdc12p to the mating hormone induced Afr1p and of Cdc10p to the chitin associated Bni4p, suggests roles for septins in determining the site of fusion in yeast in chitin deposition, respectively. Identification of interacting proteins implicates yeast septins in cell cycle regulation based on binding to the mitosis-inducing protein kinase Gin4p (Cdc3p, Cdc10p, Cdc11p, Cdc12p, Sep7p), to the mitotic checkpoint Bub2p (Cdc3p), and to the cyclin Clb2p degrading polo kinase Cdc5p (Cdc11p and Cdc12p). In addition, yeast septins Cdc3p and Cdc11p are cell-cycle specific substrates for conjugation to the ubiquitin-related SUMOp. Of the mammalian septins, the ARTS protein translocates from the mitochondria to the nucleus and enhances cell death in response to TGF-b. CDC10, NEDD5, H5, E-septin and Septin 6 associate with the rat SEC6/SEC8 multimer, a key conserved complex in targeting exocytosis at the plasma membrane. H5 binds in a GDP-associated form to membrane phospholipids through a polybasic domain, and CDCREL-1 and NEDD5 co-purify with brain synaptic vesicles and interact with syntaxin, a key protein for vesicle-membrane fusion. Given that cytokinesis is one highly conserved role of septins, this later observation is somewhat unexpected given that brain typically exhibits minimal cell division activity. However, further evidence for septin roles in neuronal development are suggested by the degradation of CDCREL-1 after ubiquitination by the Parkinsons disease gene parkin and by the association of NEDD5, H5, and DIFF6 septins with neurofibrillary tangles in Alzheimers disease. Thus, these data present a myriad of functions for members of the septin protein family that can be more broadly grouped into integral roles in cell cycle regulation, signal transduction, and protein/vesicle trafficking through cytoskeletal scaffolding. These functions are probably partially determined by tissue- and temporal-specific expression levels of individual septins and their isoforms and provide a framework for further characterization of MSF.
Homology
MSF exhibits protein homology (% identity/% similarity) to the following orthologous septin family members: S. cerevisiae CDC10 47%/67%, D. melanogaster Pnut 39%/57%, R. norvegicus E-septin short 96%/97%, M. musculus SINT1 96%/98%, and to the following human septins also found as fusion proteins with MLL in leukemia patients: hCDCRel-1 in 22q11 45%/66%, Septin 6/KIAA0128 in Xq24 43%/66%.
Implicated in
Entity name
Disease
de novo and treatment related leukemia
Prognosis
poor
Hybrid gene
in-frame transcript joining the 5 of MLL through exon 5 to MSF at the start of exon 3 through the 3 terminus. A reciprocal transcript was amplified in one patient joining the 5 of MSF through exon 1 to exon 7 of MLL but was out of frame. No 5 MSF-3 MLL transcript was amplified in another patient.
Fusion protein
fusion protein of an amino protein terminus MLL, including the nuclear localization signal, the A-T hook DNA binding domain, and the DNA methyltransferase-like DNA binding domain, and a carboxy terminus MSF, including the xylose isomerase, the polybasic and GTPase domains. Lost from the carboxy terminus of MLL is the PHD zinc finger protein-protein interaction domain and the SET domain thought to regulate gene expression through chromatin remodeling. It has been suggested that MLL is the sole clinical culprit in leukemias with 11q23 rearrangements as it is fused to a wide variety of other genes. However, these translocations produce an in-frame fusion protein, suggesting selection for a translatable protein. In addition, the general observations of phenotypic variability with different translocations, including patient age, leukemia types, and prognostic outcomes, provide further evidence that proteins at the varying reciprocally translocated chromosomes are essential contributors to the pathogenesis of leukemia. Thus, speculation on the contributions of MLL-MSF fusion protein expression to haematopoetic cellular transformation would include potential mislocalization of MSF from the cytoplasm to the nucleus, aberrant expression of MLL target proteins and altered activation of MSF GTPase signaling pathways.
Oncogenesis
In addition to the involvement of various septins in leukemia patients, the MSF mouse ortholog SINT1 was identified by virtue of its presence at a provirus insertion site in SL3-3 MLV-induced lymphomas suggesting a subgroup of septins may play a more specific role in leukemogenesis.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 10673329 | 2000 | Genomic and expression analyses of alternatively spliced transcripts of the MLL septin-like fusion gene (MSF) that map to a 17q25 region of loss in breast and ovarian tumors. | Kalikin LM et al |
| 10339604 | 1999 | MSF (MLL septin-like fusion), a fusion partner gene of MLL, in a therapy-related acute myeloid leukemia with a t(11;17)(q23;q25). | Osaka M et al |
| 10987277 | 2000 | Isolation and mapping of a human septin gene to a region on chromosome 17q, commonly deleted in sporadic epithelial ovarian tumors. | Russell SE et al |
| 10485469 | 1999 | AF17q25, a putative septin family gene, fuses the MLL gene in acute myeloid leukemia with t(11;17)(q23;q25). | Taki T et al |
Citation
Linda M. Kalikin ; Elizabeth M. Petty
SEPT9 (MLL septin-like fusion)
Atlas Genet Cytogenet Oncol Haematol. 2001-08-01
Online version: http://atlasgeneticsoncology.org/gene/208/sept9
Historical Card
2000-04-01 SEPT9 (MLL septin-like fusion) by Jean-Loup Huret Affiliation
Genetics, Dept Medical Information, University of Poitiers, CHU Poitiers Hospital, F-86021 Poitiers, France
