KMT2A (myeloid/lymphoid or mixed lineage leukemia)
2005-10-01 Jean-Loup Huret AffiliationInstituts für Pharmazeutische Biologie, JWG Universitaet Frankfurt\\\/Main, Biozentrum, N230, 303, Marie Curie Str. 9, D-60439 Frankfurt\\\/Main, Germany
Identity


DNA/RNA
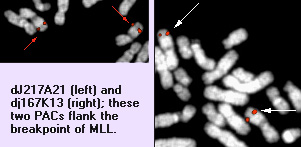
Description
Transcription
Proteins
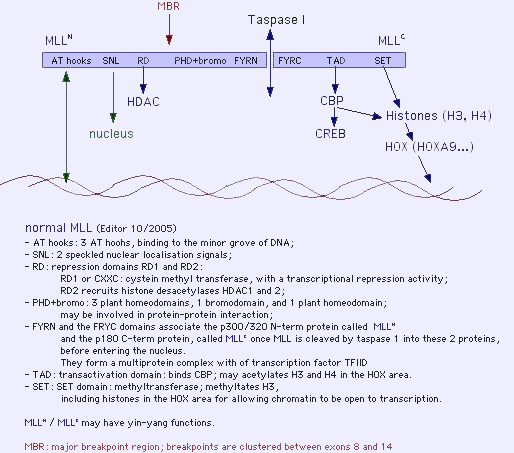
Description

Expression
Localisation
Function
Homology
Mutations
Note
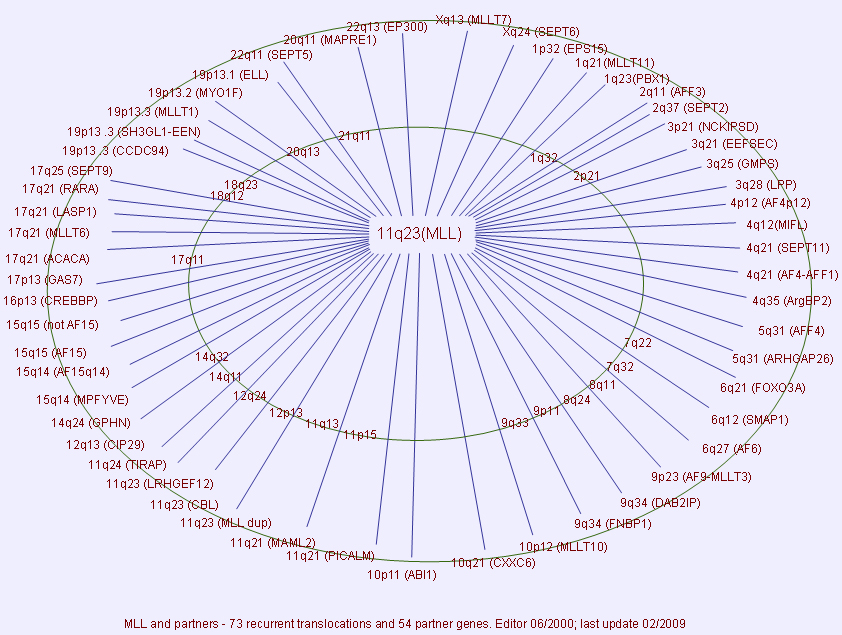
Implicated in
Breakpoints
Note
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 7542910 | 1995 | Molecular basis of 11q23 rearrangements in hematopoietic malignant proliferations. | Bernard OA et al |
| 10468849 | 1999 | Mll rearrangements in haematological malignancies: lessons from clinical and biological studies. | Dimartino JF et al |
| 10637508 | 1999 | MLL2, the second human homolog of the Drosophila trithorax gene, maps to 19q13.1 and is amplified in solid tumor cell lines. | Huntsman DG et al |
| 8703835 | 1996 | Exon/intron structure of the human ALL-1 (MLL) gene involved in translocations to chromosomal region 11q23 and acute leukaemias. | Nilson I et al |
| 9247308 | 1997 | Structure and expression pattern of human ALR, a novel gene with strong homology to ALL-1 involved in acute leukemia and to Drosophila trithorax. | Prasad R et al |
| 8620491 | 1996 | Complete exon structure of the ALL1 gene. | Rasio D et al |
| 8558942 | 1996 | 11q23 rearrangements in acute leukemia. | Rubnitz JE et al |
| 7837391 | 1995 | Self-fusion of the ALL1 gene. A new genetic mechanism for acute leukemia. | Schichman SA et al |
| 9103672 | 1997 | Disruption of a homolog of trithorax by 11q23 translocations: leukemogenic and transcriptional implications. | Waring PM et al |
| 8977038 | 1996 | Chromosome abnormalities in leukaemia: the 11q23 paradigm. | Young BD et al |
Other Information
Locus ID:
NCBI: 4297
MIM: 159555
HGNC: 7132
Ensembl: ENSG00000118058
Variants:
dbSNP: 4297
ClinVar: 4297
TCGA: ENSG00000118058
COSMIC: KMT2A
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38029383 | 2024 | Functional relevance of circRNA aberrant expression in pediatric acute leukemia with KMT2A::AFF1 fusion. | 1 |
| 38317232 | 2024 | Epigenetic drug screening for trophoblast syncytialization reveals a novel role for MLL1 in regulating fetoplacental growth. | 0 |
| 38576284 | 2024 | Impact of MLL::AF9 Gene Rearrangement on Survival of Acute Myeloid Leukaemia Patients: An Insight into Pakistani Population. | 0 |
| 38719857 | 2024 | Regulation of cytokine and chemokine expression by histone lysine methyltransferase MLL1 in rheumatoid arthritis synovial fibroblasts. | 0 |
| 38892207 | 2024 | Distinct Responses to Menin Inhibition and Synergy with DOT1L Inhibition in KMT2A-Rearranged Acute Lymphoblastic and Myeloid Leukemia. | 0 |
| 38963485 | 2024 | The Core Complex of Yeast COMPASS and Human Mixed-Lineage Leukemia (MLL), Structure, Function, and Recognition of the Nucleosome. | 0 |
| 38029383 | 2024 | Functional relevance of circRNA aberrant expression in pediatric acute leukemia with KMT2A::AFF1 fusion. | 1 |
| 38317232 | 2024 | Epigenetic drug screening for trophoblast syncytialization reveals a novel role for MLL1 in regulating fetoplacental growth. | 0 |
| 38576284 | 2024 | Impact of MLL::AF9 Gene Rearrangement on Survival of Acute Myeloid Leukaemia Patients: An Insight into Pakistani Population. | 0 |
| 38719857 | 2024 | Regulation of cytokine and chemokine expression by histone lysine methyltransferase MLL1 in rheumatoid arthritis synovial fibroblasts. | 0 |
| 38892207 | 2024 | Distinct Responses to Menin Inhibition and Synergy with DOT1L Inhibition in KMT2A-Rearranged Acute Lymphoblastic and Myeloid Leukemia. | 0 |
| 38963485 | 2024 | The Core Complex of Yeast COMPASS and Human Mixed-Lineage Leukemia (MLL), Structure, Function, and Recognition of the Nucleosome. | 0 |
| 34970734 | 2023 | Comparison of minimal residual disease measurement by multicolour flow cytometry and PCR for fusion gene transcripts in infants with acute lymphoblastic leukaemia with KMT2A gene rearrangements. | 1 |
| 35833755 | 2023 | Mutational landscape and clinical outcome of pediatric acute myeloid leukemia with 11q23/KMT2A rearrangements. | 4 |
| 36094360 | 2023 | MLL1:EZH2 Ratio in Uterine Secretions and Endometrial Receptivity in Patients with Endometriosis. | 0 |
Citation
Jean-Loup Huret
KMT2A (myeloid/lymphoid or mixed lineage leukemia)
Atlas Genet Cytogenet Oncol Haematol. 2005-10-01
Online version: http://atlasgeneticsoncology.org/gene/13/kmt2a-(myeloid-lymphoid-or-mixed-lineage-leukemia)
Historical Card
2002-11-01 KMT2A (myeloid/lymphoid or mixed lineage leukemia) by Rolf Marschalek Affiliation
2000-12-01 KMT2A (myeloid/lymphoid or mixed lineage leukemia) by Jay L Hess,Jean-Loup Huret Affiliation
1997-12-01 KMT2A (myeloid/lymphoid or mixed lineage leukemia) by Jean-Loup Huret Affiliation
