Identity

Abstract
HMGA2, the High Mobility Group A2 gene, is a non-histone and architectural transcription factor. As an oncofetal protein, HMGA2 plays an important role in development and contributes to the tumorigenesis of many epithelial and mesenchymal tumors. Upregulation of HMGA2 by non-random chromosomal translocations is common in mesenchymal tumors, whereas by the altered transcription regulation is likely the major mechanism in malignant epithelial tumors and it involves much more complex mechanisms. HMGA2 directly and indirectly regulates the multiple biological and oncogenic pathways. Its oncogenic property remains to be fully characterized.
DNA/RNA
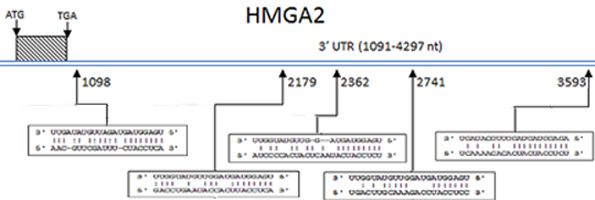
Description
Transcription
Proteins
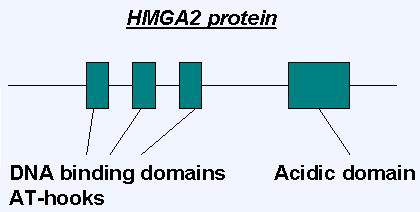
Description
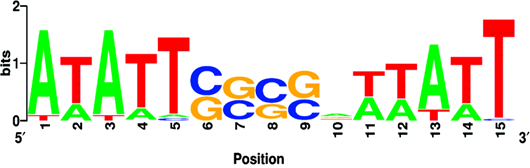
Expression
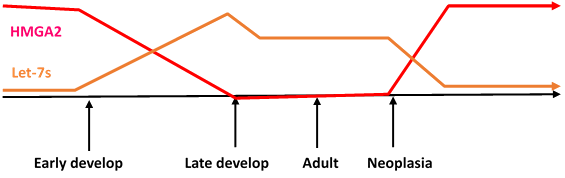
Localisation
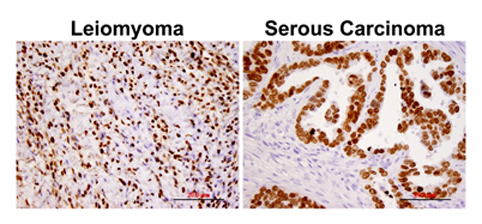
Function
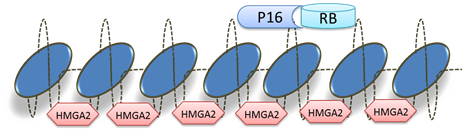
Homology
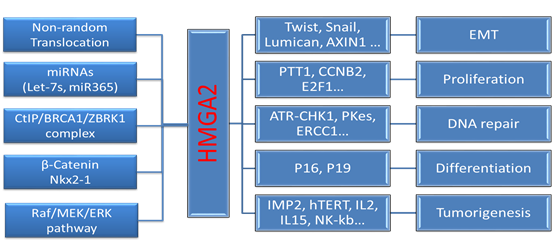
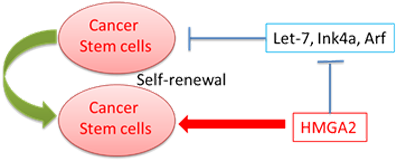
Mutations
Germinal
8-year-old boy had a de novo pericentric inversion of chromosome 12, with breakpoints at p11.22 and q14.3. The phenotype included extreme somatic overgrowth, advanced endochondral bone and dental ages, a cerebellar tumour, and multiple lipomas. His chromosomal inversion was found to truncate HMGA2, which maps to the 12q14.3 breakpoint.
Implicated in
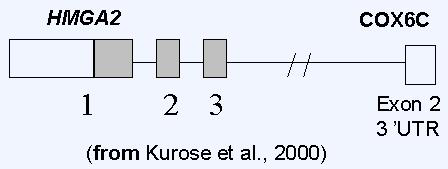
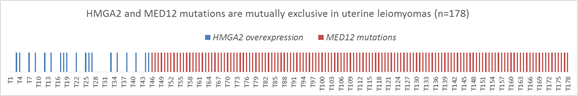
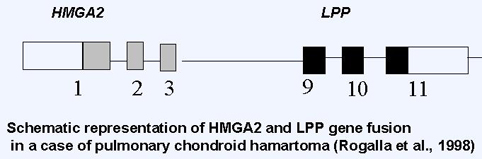
Overexpression of HMGA2 is associated with early tumorigenesis, tumor cell proliferation, invasion and worse outcome through regulation of cell cycle, epithelial to mesenchymal transition (Wu et al. 2011).
Inhibition of PAR1 signaling suppresses HMGA2-driven invasion in breast cancer cells (Yang et al. 2015).
Overexpression of HMGA2 is associated with metastasis and unequivocally occurred in parallel with reduced survival rates of patients with colorectal carcinoma (Wang et al. 2011).
HMGA2 promotes the transformation of lung cancer cells independent of protein-coding function. Tgfbr3 expression is regulated by the Hmga2 ceRNA through differential recruitment to Argonaute 2 (AGO2), and TGF-β signalling driven by Tgfbr3 is important for Hmga2 to promote lung cancer progression (Kumar et al. 2014).
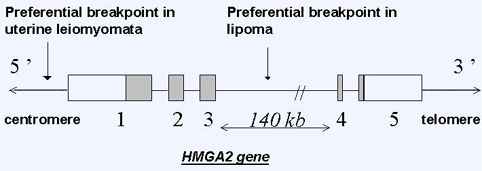
Breakpoints
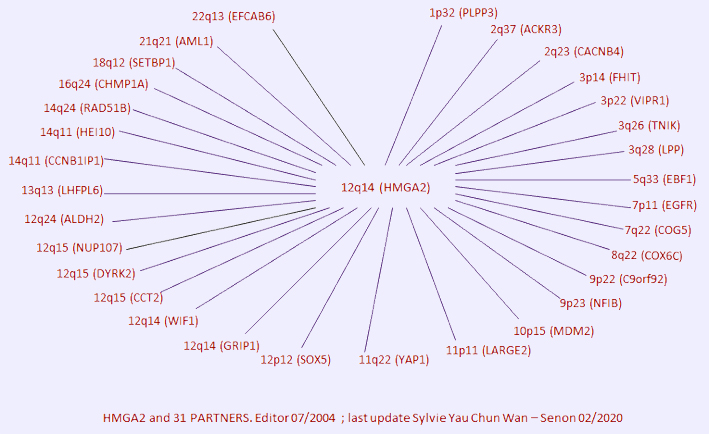
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 14647145 | 2003 | An increased high-mobility group A2 expression level is associated with malignant phenotype in pancreatic exocrine tissue. | Abe N et al |
| 15497774 | 2004 | High mobility group I-C protein in astrocytoma and glioblastoma. | Akai T et al |
| 10742101 | 2000 | In vivo modulation of Hmgic reduces obesity. | Anand A et al |
| 14603445 | 2004 | Dysregulation and overexpression of HMGA2 in myelofibrosis with myeloid metaplasia. | Andrieux J et al |
| 10747931 | 2000 | Transgenic mice expressing a truncated form of the high mobility group I-C protein develop adiposity and an abnormally high prevalence of lipomas. | Arlotta P et al |
| 7606786 | 1995 | Disruption of the architectural factor HMGI-C: DNA-binding AT hook motifs fused in lipomas to distinct transcriptional regulatory domains. | Ashar HR et al |
| 4081525 | 1985 | Prevention from naturally acquired viral respiratory infection by interferon nasal spray. | Saito H et al |
| 12569368 | 2003 | Human HMGA2 promoter is coregulated by a polymorphic dinucleotide (TC)-repeat. | Borrmann L et al |
| 12118328 | 2002 | Fusion of RDC1 with HMGA2 in lipomas as the result of chromosome aberrations involving 2q35-37 and 12q13-15. | Broberg K et al |
| 15650238 | 2004 | HMGA molecules in neuroblastic tumors. | Cerignoli F et al |
| 12032866 | 2002 | HMGA1 and HMGA2 protein expression in mouse spermatogenesis. | Chieffi P et al |
| 16125217 | 2005 | Array comparative genomic hybridization analysis of uterine leiomyosarcoma. | Cho YL et al |
| 11979097 | 2002 | Hyaline vascular Castleman's disease with HMGIC rearrangement in follicular dendritic cells: molecular evidence of mesenchymal tumorigenesis. | Cokelaere K et al |
| 15755872 | 2005 | Transactivation functions of the tumor-specific HMGA2/LPP fusion protein are augmented by wild-type HMGA2. | Crombez KR et al |
| 17956125 | 2007 | Specific recognition of AT-rich DNA sequences by the mammalian high mobility group protein AT-hook 2: a SELEX study. | Cui T et al |
| 17957144 | 2007 | Let-7 prevents early cancer progression by suppressing expression of the embryonic gene HMGA2. | Park SM et al |
| 14614053 | 2003 | Fusion, disruption, and expression of HMGA2 in bone and soft tissue chondromas. | Dahlén A et al |
| 1855228 | 1991 | The Mary Lasker Conference, on growth factors in hormone-related tumors. | Mueller GC et al |
| 12082634 | 2002 | Overexpression of the HMGA2 gene in transgenic mice leads to the onset of pituitary adenomas. | Fedele M et al |
| 11956103 | 2002 | The High Mobility Group A2 gene is amplified and overexpressed in human prolactinomas. | Finelli P et al |
| 10398424 | 1999 | HMGIC expression in human adult and fetal tissues and in uterine leiomyomata. | Gattas GJ et al |
| 11921278 | 2002 | Sequencing of intron 3 of HMGA2 uncovers the existence of a novel exon. | Hauke S et al |
| 11170289 | 2001 | Chromosomal rearrangements leading to abnormal splicing within intron 4 of HMGIC? | Hauke S et al |
| 9495195 | 1998 | Chromosomal translocations in benign tumors: the HMGI proteins. | Hess JL et al |
| 2769577 | 1989 | Predictors of nutrition supplement use in the elderly. Part I: A review of the literature. | Cotugna N et al |
| 16056249 | 2005 | HMGA proteins in malignant peripheral nerve sheath tumor and synovial sarcoma: preferential expression of HMGA2 in malignant peripheral nerve sheath tumor. | Hui P et al |
| 10686944 | 1999 | Inflammatory myofibroblastic tumor with HMGIC rearrangement. | Kazmierczak B et al |
| 8908160 | 1996 | Amplification of a rearranged form of the high-mobility group protein gene HMGIC in OsA-CI osteosarcoma cells. | Kools PF et al |
| 3886898 | 1985 | Nutritional requirements of Plasmodium falciparum in culture. I. Exogenously supplied dialyzable components necessary for continuous growth. | Divo AA et al |
| 11135440 | 2001 | Three aberrant splicing variants of the HMGIC gene transcribed in uterine leiomyomas. | Kurose K et al |
| 12505264 | 2002 | Expression of the HMGA2-LPP fusion transcript in only 1 of 61 karyotypically normal pulmonary chondroid hamartomas. | Lemke I et al |
| 15593017 | 2005 | Constitutional rearrangement of the architectural factor HMGA2: a novel human phenotype including overgrowth and lipomas. | Ligon AH et al |
| 22322863 | 2012 | MiR-182 overexpression in tumourigenesis of high-grade serous ovarian carcinoma. | Liu Z et al |
| 11391797 | 2001 | Ectopic sequences from truncated HMGIC in liposarcomas are derived from various amplified chromosomal regions. | Meza-Zepeda LA et al |
| 11223542 | 2001 | Fusion of a sequence from HEI10 (14q11) to the HMGIC gene at 12q15 in a uterine leiomyoma. | Mine N et al |
| 15026339 | 2004 | Expression of mesenchyme-specific gene HMGA2 in squamous cell carcinomas of the oral cavity. | Miyazawa J et al |
| 3249932 | 1988 | [The effect of atropine, quinuclidinyl benzilate and their antidote tetrahydroaminoacridine on adhesion and spreading in cells cultured in vitro]. | Kolárová J et al |
| 1690178 | 1990 | Expression of HRF20, a regulatory molecule of complement activation, on peripheral blood mononuclear cells. | Hideshima T et al |
| 16133369 | 2005 | Fusion of the HMGA2 and NFIB genes in lipoma. | Nilsson M et al |
| 2582221 | 1985 | Fc gamma-receptor blocking antibodies in hyperimmune and normal pooled gammaglobulin. | Templeton JG et al |
| 11550285 | 2001 | Chromosomal translocation t(8;12) induces aberrant HMGIC expression in aggressive angiomyxoma of the vulva. | Nucci MR et al |
| 15495192 | 2005 | Frequent deletions at 12q14.3 chromosomal locus in adult acute lymphoblastic leukemia. | Patel HS et al |
| 18403645 | 2008 | Antiproliferative effects by Let-7 repression of high-mobility group A2 in uterine leiomyoma. | Peng Y et al |
| 8812423 | 1996 | LPP, the preferred fusion partner gene of HMGIC in lipomas, is a novel member of the LIM protein gene family. | Petit MM et al |
| 12851685 | 2003 | HMGA2 locus rearrangement in a case of acute lymphoblastic leukemia. | Pierantoni GM et al |
| 12649198 | 2003 | Fusion transcripts involving HMGA2 are not a common molecular mechanism in uterine leiomyomata with rearrangements in 12q15. | Quade BJ et al |
| 9598796 | 1998 | The t(3;12)(q27;q14-q15) with underlying HMGIC-LPP fusion is not determining an adipocytic phenotype. | Rogalla P et al |
| 11890997 | 2002 | Absence of HMGIC-LHFP fusion in pulmonary chondroid hamartomas with aberrations involving chromosomal regions 12q13 through 15 and 13q12 through q14. | Rogalla P et al |
| 8889500 | 1996 | Translocation breakpoints upstream of the HMGIC gene in uterine leiomyomata suggest dysregulation of this gene by a mechanism different from that in lipomas. | Schoenberg Fejzo M et al |
| 9892177 | 1999 | Allelic knockout of novel splice variants of human recombination repair gene RAD51B in t(12;14) uterine leiomyomas. | Schoenmakers EF et al |
| 2040910 | 1991 | [Development of acute endolymphatic hydrops following secondary endolymphatic sac immune response. I: Short-term observation]. | Tomiyama S et al |
| 11135437 | 2001 | Evidence for RAD51L1/HMGIC fusion in the pathogenesis of uterine leiomyoma. | Takahashi T et al |
| 12021922 | 2002 | Translocation of the HMGI-C ( HMGA2) gene in a benign mesenchymoma (chondrolipoangioma). | Van Dorpe J et al |
| 17243163 | 2007 | A micro-RNA signature associated with race, tumor size, and target gene activity in human uterine leiomyomas. | Wang T et al |
| 3079060 | 1987 | A controlled prospective trial of triple therapy with low-dose azathioprine, cyclosporine, and methylprednisolone in renal transplantation. | De Vecchi A et al |
| 9087575 | 1997 | Hidden paracentric inversions of chromosome arm 12q affecting the HMGIC gene. | Wanschura S et al |
| 2665746 | 1989 | Perturbations in nitrogen metabolism of brain and liver of rat following repeated benthiocarb administration. | Babu GR et al |
| 23686260 | 2013 | HMGA2 and high-grade serous ovarian carcinoma. | Wu J et al |
| 21224353 | 2011 | HMGA2 overexpression-induced ovarian surface epithelial transformation is mediated through regulation of EMT genes. | Wu J et al |
| 26165842 | 2016 | Dysregulated protease activated receptor 1 (PAR1) promotes metastatic phenotype in breast cancer through HMGA2. | Yang E et al |
| 18083101 | 2007 | let-7 regulates self renewal and tumorigenicity of breast cancer cells. | Yu F et al |
| 7651535 | 1995 | Mutation responsible for the mouse pygmy phenotype in the developmentally regulated factor HMGI-C. | Zhou X et al |
Other Information
Locus ID:
NCBI: 8091
MIM: 600698
HGNC: 5009
Ensembl: ENSG00000149948
Variants:
dbSNP: 8091
ClinVar: 8091
TCGA: ENSG00000149948
COSMIC: HMGA2
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37461232 | 2024 | Keratin-Positive Giant Cell-Rich Tumor of Bone Harboring an HMGA2::NCOR2 Fusion: Two Cases, Including a Patient With Metastatic Disease, and Review of the Literature. | 2 |
| 37822284 | 2024 | CircDLGAP4 overexpression ameliorates neuronal injury in Parkinson's disease by binding to EIF4A3 and increasing HMGA2 expression. | 0 |
| 37849332 | 2024 | Expanding the histological spectrum of salivary gland neoplasms with HMGA2::WIF1 fusion emphasising their malignant potential: a report of eight cases. | 0 |
| 38151824 | 2024 | Exosomal circFAM63Bsuppresses bone regeneration of postmenopausal osteoporosis via regulating miR-578/HMGA2 axis. | 1 |
| 38266920 | 2024 | SOX10-Internal Tandem Duplications and PLAG1 or HMGA2 Fusions Segregate Eccrine-Type and Apocrine-Type Cutaneous Mixed Tumors. | 0 |
| 38378354 | 2024 | Effects of HMGA2 on the biological characteristics and stemness acquisition of gastric cancer cells. | 0 |
| 38506034 | 2024 | The role of HMGA2 in activating the IGFBP2 expression to promote angiogenesis and LUAD metastasis via the PI3K/AKT/VEGFA signaling pathway. | 0 |
| 38516887 | 2024 | Characterization of HMGA2 variants expands the spectrum of Silver-Russell syndrome. | 0 |
| 38802807 | 2024 | Association of deletion polymorphism rs10573247 in the HMGA2 gene with the risk of breast cancer: bioinformatic and experimental analyses. | 0 |
| 38811851 | 2024 | Chromatin modifier Hmga2 promotes adult hematopoietic stem cell function and blood regeneration in stress conditions. | 0 |
| 37461232 | 2024 | Keratin-Positive Giant Cell-Rich Tumor of Bone Harboring an HMGA2::NCOR2 Fusion: Two Cases, Including a Patient With Metastatic Disease, and Review of the Literature. | 2 |
| 37822284 | 2024 | CircDLGAP4 overexpression ameliorates neuronal injury in Parkinson's disease by binding to EIF4A3 and increasing HMGA2 expression. | 0 |
| 37849332 | 2024 | Expanding the histological spectrum of salivary gland neoplasms with HMGA2::WIF1 fusion emphasising their malignant potential: a report of eight cases. | 0 |
| 38151824 | 2024 | Exosomal circFAM63Bsuppresses bone regeneration of postmenopausal osteoporosis via regulating miR-578/HMGA2 axis. | 1 |
| 38266920 | 2024 | SOX10-Internal Tandem Duplications and PLAG1 or HMGA2 Fusions Segregate Eccrine-Type and Apocrine-Type Cutaneous Mixed Tumors. | 0 |
Citation
Jian-Jun Wei
HMGA2 (high mobility group AT-hook 2)
Atlas Genet Cytogenet Oncol Haematol. 2015-10-01
Online version: http://atlasgeneticsoncology.org/gene/82/hmga2-(high-mobility-group-at-hook-2)
Historical Card
2005-12-01 HMGA2 (high mobility group AT-hook 2) by Karin Broberg Affiliation
2000-05-01 HMGA2 (high mobility group AT-hook 2) by Florence Pedeutour Affiliation
