PRKAB1 (protein kinase, AMP-activated, beta 1 non-catalytic subunit)
2007-09-01 Monserrat Sanchez-Cespedes AffiliationMolecular Pathology Program, Spanish National Cancer Center (cnio), Melchor Fernandez Almagro, 3, 28029 Madrid, Spain
DNA/RNA
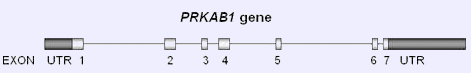
PRKAB1 gene structure. UTR: untranslated region.
Description
The PRKAB1 gene, with a total genomic size of 13639 bp, is composed of 7 coding exons of varying lengths.
Transcription
The human PRKAB1 transcript has approximately 2500-bp and contains an open reading frame of 813 bases, resulting in a predicted protein of 270 amino acid residues. PRKAB mRNA is detected in most tissues.
Proteins
Description
PRKAB1 has a molecular mass of 30382 Da; This protein constitutes a regulatory subunit of the AMP-activated protein kinase (AMPK).
AMPK is an heterotrimer of an alpha catalytic, a beta and a gamma non-catalytic regulatory subunits, each encoded by two or three distinct genes (AMPKalpha1-AMPKalpha2; AMPKbeta1-AMPKbeta2; AMPKgamma1-AMPKgamma2-AMPKgamma3) which have varying tissue and subcellular expression.
Post-translational modifications of PRKAB1 include myristoylation and phosphorylation in vivo at Ser24/25, Ser108 and Ser182. Phosphorylation at Ser108 is required for the activation of the AMPK enzyme, whereas phosphorylation at Ser24/25 and Ser182 affects the localization of the complex.
AMPK is an heterotrimer of an alpha catalytic, a beta and a gamma non-catalytic regulatory subunits, each encoded by two or three distinct genes (AMPKalpha1-AMPKalpha2; AMPKbeta1-AMPKbeta2; AMPKgamma1-AMPKgamma2-AMPKgamma3) which have varying tissue and subcellular expression.
Post-translational modifications of PRKAB1 include myristoylation and phosphorylation in vivo at Ser24/25, Ser108 and Ser182. Phosphorylation at Ser108 is required for the activation of the AMPK enzyme, whereas phosphorylation at Ser24/25 and Ser182 affects the localization of the complex.
Expression
AMPKbeta1 protein expression is highest in the liver, and testis and low in kidney and skeletal muscle. In contrast, expression of PRKAB2, encoded by a different gene, is higher in skeletal muscle. This indicates a tissue specific pattern of expression of these two regulatory beta subunits.
Localisation
AMPKbeta1 localises in the cytoplasm. However nuclear translocation is possible because mutations of Ser24/25 and Ser182 to alanine lead to the redistribution of PRKAB1 to the nucleus.
Function
AMPKbeta1, may act as a positive regulator of AMPK activity and it may serve as an adaptor molecule for the catalytic subunit. It has also been reported that AMPKbeta1 contains a glycogen-binding domain (GBD) that may target the heterotrimer to glycogen storage sites.
The heterotrimeric protein AMPK senses low intracellular energy levels upon increased in the AMP/ATP ratio. Binding of AMP results in allosteric activation, inducing phosphorylation on Thr-172 of the AMPKa regulatory subunit (PRKAA) by LKB1 in complex with STE20-related adapter-alpha (STRAD alpha). AMPK activation leads to the modulation of the activity of multiple downstream targets to normalize ATP levels.
Among these substrates is the tuberin protein, the product of the tuberous sclerosis complex 2 gene (TSC2) that upon activation by AMPK represses the activity of the mammalian target of rapamycin, mTOR.
It has been reported that PRKAB1 and PRKAB2 contain a glycogen binding domain that targets AMPK to glycogen.
Moreover, it has been shown that expression of PRKAB1 and PRKAB2 genes, in human cells, may be mediated by p53.
The heterotrimeric protein AMPK senses low intracellular energy levels upon increased in the AMP/ATP ratio. Binding of AMP results in allosteric activation, inducing phosphorylation on Thr-172 of the AMPKa regulatory subunit (PRKAA) by LKB1 in complex with STE20-related adapter-alpha (STRAD alpha). AMPK activation leads to the modulation of the activity of multiple downstream targets to normalize ATP levels.
Among these substrates is the tuberin protein, the product of the tuberous sclerosis complex 2 gene (TSC2) that upon activation by AMPK represses the activity of the mammalian target of rapamycin, mTOR.
It has been reported that PRKAB1 and PRKAB2 contain a glycogen binding domain that targets AMPK to glycogen.
Moreover, it has been shown that expression of PRKAB1 and PRKAB2 genes, in human cells, may be mediated by p53.
Homology
There is a AMPKbeta1 isoform, designated AMPKbeta2, encoded by the PRKAB2 gene. The N-terminal region of beta2 differs significantly from that AMPKbeta1 isoform, suggesting that this region could play a role in isoform-specific AMPK activity.
The AMPKbeta subunit is the mammalian homolog of the S. cerevisiae Sip1p/Sip2p/Gal83p family of proteins that interact with the AMPKa homolog, Snf1p, and are involved in glucose regulation of gene expression.
The AMPKbeta subunit is the mammalian homolog of the S. cerevisiae Sip1p/Sip2p/Gal83p family of proteins that interact with the AMPKa homolog, Snf1p, and are involved in glucose regulation of gene expression.
Mutations
Note
No germ-line or somatic mutations have been reported in the PRKAB1 gene.
Implicated in
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 17409411 | 2007 | The regulation of AMPK beta1, TSC2, and PTEN expression by p53: stress, cell and tissue specificity, and the role of these gene products in modulating the IGF-1-AKT-mTOR pathways. | Feng Z et al |
| 8621499 | 1996 | Non-catalytic beta- and gamma-subunit isoforms of the 5'-AMP-activated protein kinase. | Gao G et al |
| 12960015 | 2003 | Minireview: the AMP-activated protein kinase cascade: the key sensor of cellular energy status. | Hardie DG et al |
| 14511394 | 2003 | Complexes between the LKB1 tumor suppressor, STRAD alpha/beta and MO25 alpha/beta are upstream kinases in the AMP-activated protein kinase cascade. | Hawley SA et al |
| 14651849 | 2003 | TSC2 mediates cellular energy response to control cell growth and survival. | Inoki K et al |
| 12771025 | 2003 | AMPK-beta1 subunit is a p53-independent stress responsive protein that inhibits tumor cell growth upon forced expression. | Li J et al |
| 9305909 | 1997 | Posttranslational modifications of the 5'-AMP-activated protein kinase beta1 subunit. | Mitchelhill KI et al |
| 12747837 | 2003 | AMPK beta subunit targets metabolic stress sensing to glycogen. | Polekhina G et al |
| 17599048 | 2007 | A role for LKB1 gene in human cancer beyond the Peutz-Jeghers syndrome. | Sanchez-Cespedes M et al |
| 9224708 | 1997 | AMP-activated protein kinase isoenzyme family: subunit structure and chromosomal location. | Stapleton D et al |
| 9575201 | 1998 | Identification of a novel AMP-activated protein kinase beta subunit isoform that is highly expressed in skeletal muscle. | Thornton C et al |
| 11171104 | 2001 | Post-translational modifications of the beta-1 subunit of AMP-activated protein kinase affect enzyme activity and cellular localization. | Warden SM et al |
| 14614828 | 2003 | LKB1 is the upstream kinase in the AMP-activated protein kinase cascade. | Woods A et al |
| 12764152 | 2003 | Identification of phosphorylation sites in AMP-activated protein kinase (AMPK) for upstream AMPK kinases and study of their roles by site-directed mutagenesis. | Woods A et al |
Other Information
Locus ID:
NCBI: 5564
MIM: 602740
HGNC: 9378
Ensembl: ENSG00000111725
Variants:
dbSNP: 5564
ClinVar: 5564
TCGA: ENSG00000111725
COSMIC: PRKAB1
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
PharmGKB
| Entity ID | Name | Type | Evidence | Association | PK | PD | PMIDs |
|---|---|---|---|---|---|---|---|
| PA134983031 | GPAM | Gene | Pathway | associated | 22722338 | ||
| PA142672073 | CRTC2 | Gene | Pathway | associated | 22722338 | ||
| PA189 | HMGCR | Gene | Pathway | associated | 22722338 | ||
| PA24421 | ACACA | Gene | Pathway | associated | 22722338 | ||
| PA24422 | ACACB | Gene | Pathway | associated | 22722338 | ||
| PA30861 | MLYCD | Gene | Pathway | associated | 22722338 | ||
| PA335 | SREBF1 | Gene | Pathway | associated | 22722338 | ||
| PA35879 | SLC2A4 | Gene | Pathway | associated | 22722338 | ||
| PA36198 | STK11 | Gene | Pathway | associated | 22722338 | ||
| PA37353 | MLXIPL | Gene | Pathway | associated | 22722338 | ||
| PA37935 | SIRT1 | Gene | Pathway | associated | 22722338 | ||
| PA450395 | metformin | Chemical | Pathway | associated | 22722338 | ||
| PA61 | ATM | Gene | Pathway | associated | 22722338 |
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38167670 | 2024 | Direct effects of adipocyte lipolysis on AMPK through intracellular long-chain acyl-CoA signaling. | 1 |
| 38233384 | 2024 | Phosphorylation of INF2 by AMPK promotes mitochondrial fission and oncogenic function in endometrial cancer. | 1 |
| 38266576 | 2024 | Endothelial H(2)S-AMPK dysfunction upregulates the angiocrine factor PAI-1 and contributes to lung fibrosis. | 0 |
| 38322520 | 2024 | [Extracellular Matrix Stiffness Induces Mitochondrial Morphological Heterogeneity via AMPK Activation]. | 0 |
| 38167670 | 2024 | Direct effects of adipocyte lipolysis on AMPK through intracellular long-chain acyl-CoA signaling. | 1 |
| 38233384 | 2024 | Phosphorylation of INF2 by AMPK promotes mitochondrial fission and oncogenic function in endometrial cancer. | 1 |
| 38266576 | 2024 | Endothelial H(2)S-AMPK dysfunction upregulates the angiocrine factor PAI-1 and contributes to lung fibrosis. | 0 |
| 38322520 | 2024 | [Extracellular Matrix Stiffness Induces Mitochondrial Morphological Heterogeneity via AMPK Activation]. | 0 |
| 37142272 | 2023 | Exosomal miR-663b from "M1" macrophages promotes pulmonary artery vascular smooth muscle cell dysfunction through inhibiting the AMPK/Sirt1 axis. | 1 |
| 37143083 | 2023 | Promoting AMPK/SR-A1-mediated clearance of HMGB1 attenuates chemotherapy-induced peripheral neuropathy. | 1 |
| 37217519 | 2023 | AMPK is a mechano-metabolic sensor linking cell adhesion and mitochondrial dynamics to Myosin-dependent cell migration. | 8 |
| 37142272 | 2023 | Exosomal miR-663b from "M1" macrophages promotes pulmonary artery vascular smooth muscle cell dysfunction through inhibiting the AMPK/Sirt1 axis. | 1 |
| 37143083 | 2023 | Promoting AMPK/SR-A1-mediated clearance of HMGB1 attenuates chemotherapy-induced peripheral neuropathy. | 1 |
| 37217519 | 2023 | AMPK is a mechano-metabolic sensor linking cell adhesion and mitochondrial dynamics to Myosin-dependent cell migration. | 8 |
| 32020222 | 2020 | Inhibitory mechanism of muscone in liver cancer involves the induction of apoptosis and autophagy. | 9 |
Citation
Monserrat Sanchez-Cespedes
PRKAB1 (protein kinase, AMP-activated, beta 1 non-catalytic subunit)
Atlas Genet Cytogenet Oncol Haematol. 2007-09-01
Online version: http://atlasgeneticsoncology.org/gene/44100/prkab1
