RSF1 (remodeling and spacing factor 1)
2008-07-01 Tian-Li Wang , Joon T Park AffiliationDepartments of Gynecology\\\/Obstetrics, Oncology Johns Hopkins Medical Institutions CRBII, Rm: 306 1550 Orleans Street Baltimore, MD 21231, USA
Identity
HGNC
LOCATION
11q14.1
LOCUSID
ALIAS
HBXAP,RSF-1,XAP8,p325
FUSION GENES
DNA/RNA
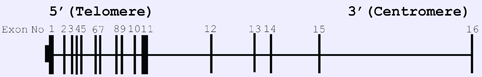
Genomic organization of human RSF1.
Description
Rsf1 gene is encoded by 16 exons spanning 154.607 Mb that are located on chromosome 11p14.1.
Transcription
5.131 Kb mRNA, the coding sequence is from 121 bp-4446 bp.
Proteins
Note
1441 amino acids, highly acidic protein (pI 4.94), with calculated molecular mass of 164 kD. SDS-PAGE detected native RSF1 at molecular mass of 200 to 300 kD
Contain a DDT domain, Zinc finger PHD-type domain which was found in nuclear proteins thought to be involved in chromatin-mediated transcriptional regulation.
RSF1 also contains 3 nuclear localization signal near C-terminus.
Contain a DDT domain, Zinc finger PHD-type domain which was found in nuclear proteins thought to be involved in chromatin-mediated transcriptional regulation.
RSF1 also contains 3 nuclear localization signal near C-terminus.
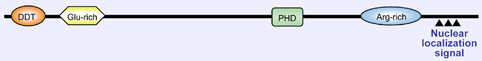
Diagram of the RSF1 protein.
Description
Using a variation of the yeast 2-hybrid screen aiming to identify proteins interacting with hepatitis B virus X protein (HBX), Shamay et al (2002a) cloned RSF1 from a spleen cDNA library. Therefore, the initial name of RSF1 was called Hepatitis B virus X-Associated Protein (HBXAP). Using 5RACE, 3 splice variants were cloned and named HBXAP-alpha, -beta, -gamma, which contains 1431, 1400, and 1189 amino acids, respectively. By characterizing a protein remodeling complex RSF, Loyola A et al has identified two interacting subunits: RSF1 and SNF2H. They used peptide sequence information of RSF1 to clone the full-length cDNA. The deduced sequence contains 1441 amino acids which includes 252 additional amino acids at N-terminus as compared to HBXAP-gamma. Reconstitute experiment by isolating protein complex from coinfection of viruses caring each subunit can recapitulate the chromatin assembly ability and ATPase activity of native complex. They also showed that SNF2H binds to DNA independently of histones. However, RSF1 couldnt bind to DNA unless histones are present.
Expression
Highly expressed in heart, skeletal muscle, kidney, and placenta, and expressed weakly in brain and colon.
Localisation
Mainly located at cell nucleus. During mitosis, the expression in the nucleus is decreased.
Function
RSF1 functions as transcription coactivator when associated with hepatitis B virus X protein (HBX). Shamay et al (2002b) observed the direct interaction between the RSF1 variant, HBXAP-gamma, and HBX. HBXAP-gamma increased hepatitis B viral transcription in an HBX-dependent manner. Furthermore, in the presence of both HBX and HBXAP, the transcription of a nuclear factor kappa-B ( NFKB ) was significantly increased. However, in the presence of HBXAP alone, the transcription of a NFKB was decreased in a dose-dependent manner. Examination of HBXAP-gamma deletion mutants showed that the interaction between HBX and HBXAP-gamma was mediated by the PHD domain in HBXAP-gamma.
RSF1 functions as chromatin remodeling and spacing when associated with SNF2H. Loyola et al reconstituted the RSF complex by overexpressing two subunits, RSF1 and SNF2H. RSF1 assembled nucleosome randomly as a histone chaperone in the nuclei. The resulting nucleosomes were then redistributed into a regularly spaced nucleosome array by the ATP-utilizing nucleosome mobilization factor SNF2H. At the cellular level, Rsf1/SNF2H complex participated in chromatin remodeling by mobilizing nucleosomes in response to a variety of growth modifying signals and environmental cues. Sheu JJ et al found that the induction of RSF1 expression affected the molecular partnership of SNF2H and translocated SNF2H into nuclei where it colocalized with RSF1. To determine which domain in the RSF1 is involved in the binding to SNF2H, a series of RSF1-deletion mutants were generated. Only the fragment that contains DDT, Glu-rich, and PHD motifs could be immunoprecipitated with SNF2H. Ectopic expression of this RSF1 fragment disrupted RSF1/SNF2H complex and resulted in remarkable growth inhibition in ovarian cancer cells with RSF1 gene amplification and overexpression, but not in the cells without detectable RSF1 expression. This finding suggests that interaction between RSF1 and SNF2H may define a survival signal in the tumors overexpressing RSF1.
RSF1 functions as chromatin remodeling and spacing when associated with SNF2H. Loyola et al reconstituted the RSF complex by overexpressing two subunits, RSF1 and SNF2H. RSF1 assembled nucleosome randomly as a histone chaperone in the nuclei. The resulting nucleosomes were then redistributed into a regularly spaced nucleosome array by the ATP-utilizing nucleosome mobilization factor SNF2H. At the cellular level, Rsf1/SNF2H complex participated in chromatin remodeling by mobilizing nucleosomes in response to a variety of growth modifying signals and environmental cues. Sheu JJ et al found that the induction of RSF1 expression affected the molecular partnership of SNF2H and translocated SNF2H into nuclei where it colocalized with RSF1. To determine which domain in the RSF1 is involved in the binding to SNF2H, a series of RSF1-deletion mutants were generated. Only the fragment that contains DDT, Glu-rich, and PHD motifs could be immunoprecipitated with SNF2H. Ectopic expression of this RSF1 fragment disrupted RSF1/SNF2H complex and resulted in remarkable growth inhibition in ovarian cancer cells with RSF1 gene amplification and overexpression, but not in the cells without detectable RSF1 expression. This finding suggests that interaction between RSF1 and SNF2H may define a survival signal in the tumors overexpressing RSF1.
Mutations
Somatic
Chromosome 11q is one of the most common targets for allelic imbalance alteration in human cancers. Several candidate tumor-promoting genes were postulated previously. In ovarian carcinoma, Shih et al, has pinpointed a minimal amplicon spanning from 76.6 to 78.4 Mb on the chromosome 11q, which harbor 13 genes. Rsf1 was identified as the gene with most consistent correlation between gene amplification and transcription up-regulation.
Implicated in
Entity name
Ovarian cancer
Prognosis
Shih et al found that amplification of Rsf1 locus was correlated with shorter overall survival of ovarian cancer patients.
Cytogenetics
Amplification of 11q13 detected an ovarian cancer cell line, OVCAR3.
Oncogenesis
Shih et al found that the amplification of chr11q13 is associated with RSF1 protein over-expression. siRNA knockdown experiment in OVCAR3 cell, which harbor Rsf-1 amplification, demonstrated it is essential for tumor cell survival.
Mao TL et al found that RSF1 overexpression was observed in 25% of high-grade ovarian serous carcinomas and in less than 7% of low-grade ovarian serous carcinoma and ovarian endometrioid. RSF1 does not express in any of the ovarian serous borderline tumors, ovarian clear cell carcinomas, ovarian mucinous carcinomas, and normal ovaries. Overexpression of RSF1 was significantly associated with high-grade ovarian serous carcinoma (P < 0.05) as compared with other types of ovarian tumors, demonstrating that RSF1 expression is primarily confined to high-grade serous carcinoma, the most aggressive ovarian cancer.
Mao TL et al found that RSF1 overexpression was observed in 25% of high-grade ovarian serous carcinomas and in less than 7% of low-grade ovarian serous carcinoma and ovarian endometrioid. RSF1 does not express in any of the ovarian serous borderline tumors, ovarian clear cell carcinomas, ovarian mucinous carcinomas, and normal ovaries. Overexpression of RSF1 was significantly associated with high-grade ovarian serous carcinoma (P < 0.05) as compared with other types of ovarian tumors, demonstrating that RSF1 expression is primarily confined to high-grade serous carcinoma, the most aggressive ovarian cancer.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 12972596 | 2003 | Functional analysis of the subunits of the chromatin assembly factor RSF. | Loyola A et al |
| 16938522 | 2006 | Expression of Rsf-1, a chromatin-remodeling gene, in ovarian and breast carcinoma. | Mao TL et al |
| 11944984 | 2002 | HBXAP, a novel PHD-finger protein, possesses transcription repression activity. | Shamay M et al |
| 18519663 | 2008 | The roles of human sucrose nonfermenting protein 2 homologue in the tumor-promoting functions of Rsf-1. | Sheu JJ et al |
| 16172393 | 2005 | Amplification of a chromatin remodeling gene, Rsf-1/HBXAP, in ovarian carcinoma. | Shih IeM et al |
Other Information
Locus ID:
NCBI: 51773
MIM: 608522
HGNC: 18118
Ensembl: ENSG00000048649
Variants:
dbSNP: 51773
ClinVar: 51773
TCGA: ENSG00000048649
COSMIC: RSF1
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36856209 | 2023 | [Circular RNA circRSF1 binds to HuR to promote radiation-induced inflammatory phenotype in hepatic stellate cells]. | 0 |
| 37108169 | 2023 | Genome-Wide Meta-Analysis Identifies Multiple Novel Rare Variants to Predict Common Human Infectious Diseases Risk. | 2 |
| 36856209 | 2023 | [Circular RNA circRSF1 binds to HuR to promote radiation-induced inflammatory phenotype in hepatic stellate cells]. | 0 |
| 37108169 | 2023 | Genome-Wide Meta-Analysis Identifies Multiple Novel Rare Variants to Predict Common Human Infectious Diseases Risk. | 2 |
| 35818253 | 2022 | miR-129-5p Plays an Anticancer Role in Colon Cancer by Targeting RSF1. | 1 |
| 36056431 | 2022 | linc00958/miR-185-5p/RSF-1 modulates cisplatin resistance and angiogenesis through AKT1/GSK3β/VEGFA pathway in cervical cancer. | 6 |
| 35818253 | 2022 | miR-129-5p Plays an Anticancer Role in Colon Cancer by Targeting RSF1. | 1 |
| 36056431 | 2022 | linc00958/miR-185-5p/RSF-1 modulates cisplatin resistance and angiogenesis through AKT1/GSK3β/VEGFA pathway in cervical cancer. | 6 |
| 34635673 | 2021 | Spatiotemporal coordination of the RSF1-PLK1-Aurora B cascade establishes mitotic signaling platforms. | 9 |
| 34850117 | 2021 | The chromatin remodeler RSF1 coordinates epigenetic marks for transcriptional repression and DSB repair. | 3 |
| 34635673 | 2021 | Spatiotemporal coordination of the RSF1-PLK1-Aurora B cascade establishes mitotic signaling platforms. | 9 |
| 34850117 | 2021 | The chromatin remodeler RSF1 coordinates epigenetic marks for transcriptional repression and DSB repair. | 3 |
| 31960423 | 2020 | Circular RNA RSF1 promotes inflammatory and fibrotic phenotypes of irradiated hepatic stellate cell by modulating miR-146a-5p. | 25 |
| 32202941 | 2020 | Long Noncoding RNA SNHG6 Promotes Proliferation and Inhibits Apoptosis in Non-small Cell Lung Cancer Cells by Regulating miR-490-3p/RSF1 Axis. | 14 |
| 32791147 | 2020 | CircRSF1 contributes to endothelial cell growth, migration and tube formation under ox-LDL stress through regulating miR-758/CCND2 axis. | 11 |
Citation
Tian-Li Wang ; Joon T Park
RSF1 (remodeling and spacing factor 1)
Atlas Genet Cytogenet Oncol Haematol. 2008-07-01
Online version: http://atlasgeneticsoncology.org/gene/44107/rsf1-(remodeling-and-spacing-factor-1)
