inv(7)(p15q34) TRB/HOXA10
t(7;7)(p15;q34) TRB/HOXA10
2005-10-01 Barbara Cauwelier , Nicole Dastugue , Anne Hagemeijer , Frank Speleman
Affiliation
1.Génétique des Hémopathies, Laboratoire dHématologie, Hpital Purpan, 31000, Toulouse Cedex, France (ND)
2.Génétique des Hémopathies, Laboratoire d Hématologie, Hpital Purpan, 31000, Toulouse Cedex, France (ND)
Clinics and Pathology
Disease
T cell acute lymphoblastic leukemia (T-ALL) and non-Hodgkin lymphoma (T-NHL)
Phenotype stem cell origin
T lineage; occurs at early stage of T cell development (CD2-, CD4+, CD8-)
Epidemiology
3.5 % of T-ALL or T-NHL
Clinics
hepato and/or splenomegaly, lymphadenopathy, mediastinal mass, moderate WBC count (15 to 100 X 109/l)
Cytology
FAB L1 or L2
Cytogenetics
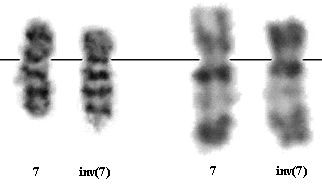
inv (7)(p15q34) G- banding (left) and R- banding (right)
Cytogenetics morphological
This rearrangement remains undetected in poor quality metaphases. In less condensed, well banded metaphases, the abnormality may be suspected as del(7)(p15) or clearly as an inv(7)(p15q34).
Cytogenetics molecular
inv(7)(p15q34) and t(7;7)(p15;q34) can be detected by FISH using either TRB and HOXA flanking probes which gives a split signal of both probes in these cases. A fusion signal will be detected when combining the proximal TCR/distal HOXA flanking or the distal TCR/proximal HOXA flanking FISH probe.
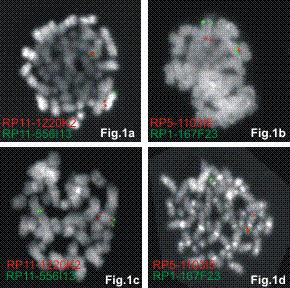
FISH results of inv(7)(Fig.1a and b) and t(7;7)(Fig.1c and d) case using TRB flanking (Fig.1a and 1c) and HOXA (Fig.1b and 1d) flanking probes
Additional anomalies
most patients show no additional karyotypic abnormalities
Genes Involved and Proteins
Note
Chromosomal disruption of the HOXA gene cluster (7p15) following chromosomal rearrangement, leads to upregulation of HOXA gene expression, which are normally weakly expressed in T-ALL. Further studies are needed to determine the various patterns of HOXA gene upregulation but from present data, HOXA10 seems most consistently involved in keeping with the breakpoint position near HOXA9. The upregulation of HOXA10 expression is thought to result from enhancers embedded within the TRB locus, which is translocated upstream from these genes. Upregulation of HOXA genes has also been described for other subgroups of T-ALL i.e. the CALM-AF10 and MLL rearranged T-ALLs indicating a more general role for HOXA genes in T-ALL development.
HOXA,together with HOXB, HOXC and HOXD, belongs to the class I homeobox genes and compromises 11 HOXA cluster genes : HOXA1, HOXA2, HOXA3, HOXA4, HOXA5, HOXA6, HOXA7, HOXA9, HOXA10, HOXA11, HOXA13. Given the breakpoint position 5¹ to HOXA10 and its consistent overexpression in all TRB-HOXA rearranged cases, we currently assume that this gene exerts a specific oncogenic effect in this subgroup of T-ALLs.
HOXA,together with HOXB, HOXC and HOXD, belongs to the class I homeobox genes and compromises 11 HOXA cluster genes : HOXA1, HOXA2, HOXA3, HOXA4, HOXA5, HOXA6, HOXA7, HOXA9, HOXA10, HOXA11, HOXA13. Given the breakpoint position 5¹ to HOXA10 and its consistent overexpression in all TRB-HOXA rearranged cases, we currently assume that this gene exerts a specific oncogenic effect in this subgroup of T-ALLs.
Gene name
HOXA10 (homeobox A10)
Location
7p15.2
Note
DNA-binding transcription factor which regulates gene expression, morphogenesis, and differentiation. More specifically, it may function in fertility, embryo viability, and regulation of hematopoietic lineage commitment. Two transcript variants encoding different isoforms have been found for this gene. HOXA10 expression is normally present in hematopoietic stem cells and developing T-cells with decreasing expression as T-cells mature.
Dna rna description
2 transcripts :transcript variant 1 (isoform a) : 2 exons, transcript 2618 bp, protein 393 amino-acids transcript variant 2 (isoform b) : 2 exons, transcript 2241 bp, protein 94 amino-acids
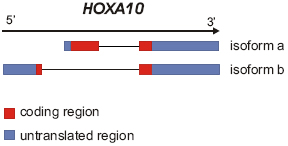
Protein description
DNA binding, transcription factor activity
Gene name
TRB (T cell Receptor Beta)
Location
7q34
Note
The human TRB locus at 7q34 spans 620 kb and consists of 82-85 genes. Enhancer sequences have been characterized 5.5kb 3 from TRBC2.
Protein description
Proteins encoded by the TRB locus are the T-cell receptor beta chains
Result of the Chromosomal Anomaly
Description
no fusion protein, but ectopic expression of HOXA10
Oncogenesis
Little is known about the target genes for HOXA10. Cyclin-dependent kinase inhibitor p21 (alias CDKN1A,CIP1,WAF1) was shown to be a transcriptional target of HOXA10 in differentiating myelomonocytic cells. However, a potential role of p21 in HOXA10 driven oncogenesis has not been proved so far. In vitro transfection experiments with HOXA9 and HOXA10 showed upregulation of several genes of the Wnt pathway (Wnt10b, Frizzled1, Frizzled5) which are essential in hematopoietic stem cell renewal.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 11040212 | 2000 | p21 is a transcriptional target of HOXA10 in differentiating myelomonocytic cells. | Bromleigh VC et al |
| 15849172 | 2005 | Activation of stem-cell specific genes by HOXA9 and HOXA10 homeodomain proteins in CD34+ human cord blood cells. | Ferrell CM et al |
| 15774621 | 2005 | HOXA genes are included in genetic and biologic networks defining human acute T-cell leukemia (T-ALL). | Soulier J et al |
| 15674412 | 2005 | A new recurrent inversion, inv(7)(p15q34), leads to transcriptional activation of HOXA10 and HOXA11 in a subset of T-cell acute lymphoblastic leukemias. | Speleman F et al |
| 12764384 | 2003 | Homeobox gene expression profile in human hematopoietic multipotent stem cells and T-cell progenitors: implications for human T-cell development. | Taghon T et al |
Summary
Fusion gene
TRB/HOXA10 TRB (-) HOXA10 (7p15.2) M inv(7)(p15q34)
Fusion gene
TRB/HOXA10 TRB (-) HOXA10 (7p15.2) M inv(7)(p15q34)
Citation
Barbara Cauwelier ; Nicole Dastugue ; Anne Hagemeijer ; Frank Speleman
inv(7)(p15q34) TRB/HOXA10
t(7;7)(p15;q34) TRB/HOXA10
Atlas Genet Cytogenet Oncol Haematol. 2005-10-01
Online version: http://atlasgeneticsoncology.org/haematological/1384/inv(7)(p15q34)-trb-hoxa10-br-t(7;7)(p15;q34)-trb-hoxa10
