DCC (deleted in colorectal carcinoma)
2010-01-01 Sarah Derks , Manon van Engeland AffiliationDepartment of Internal Medicine, Maastricht University Medical Center, PO BOX 616, 6200 MD Maastricht, The Netherlands (SD); Department of Pathology, Maastricht University Medical Center, PO BOX 616, 6200 MD Maastricht, The Netherlands (MvE)
Identity
HGNC
LOCATION
18q21.2
IMAGE

LEGEND
Diagram of chromosomal region 18q21.
LOCUSID
ALIAS
CRC18,CRCR1,HGPPS2,IGDCC1,MRMV1,NTN1R1
FUSION GENES
DNA/RNA

Diagram of the DCC gene. Blue boxes represent exons.
Description
The DCC gene is composed of 29 exons spanning in a region of 1.2 million bp. The promoter contains a CpG island located -72 bp to +217 bp relative to the transcription startsite.
Transcription
The complete transcribed mRNA is 5693 bp long.
Splice variants:
13 splice variants have been documented.
Proteins
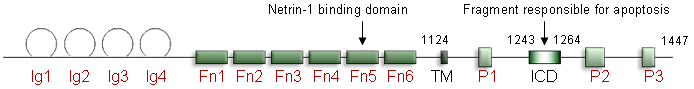
Diagram of the DCC protein. Ig: immunoglobulin, Fn: fibronectin-type III, TM: transmembrane domain, ICD: intracellular domain.
Description
DCC encodes a 158.5 kDa Type I membrane protein of 1447 amino acids with an extracellular (1100 amino acids), transmembrane and cytoplasmic (325 amino acids) domain. The extracellular domain includes four immunoglobulin-like domains and six fibronectin type III-like motifs. The cytoplasmic domain is composed of three conserved domains named P1, P2 and P3.
Expression
Expression is detected in many tissues (testis, lung, colon, esophagus, skeletal muscle) but is highest in normal brain tissue. Most tissues express low levels of transcripts and proteins.
Localisation
Cell surface.
Function
DCC is a member of the immunoglobulin superfamily of cell adhesion molecules and acts as a transmembrane dependence receptor for netrins, key factors in the regulation of axon guidance during development of the central nerve system.
In response to netrin-1, DCC becomes tyrosine phosphorylated, localizes to lipid rafts and selectively interacts with the Src family kinases Fyn and Lck to mediate axon attraction.
Furthermore DCC induces apoptosis when unbound to its ligand netrin-1. The DCC cytoplasmic domain is required for the induction of apoptosis and contains a caspase cleavage site which is recognized by caspase 3 in vitro. Upstream of the caspase cleavage domain a proapoptotic domain named addiction dependence domain (ADD) is located, which interacts with caspase 9 in the absence of Netrin-1 and binds DCC-interacting protein-13a (DIP13-a) to mediate DCC-induced cell death.
DCCs downstream effects involve MAPK activation and subsequent activation of the transcription factor Elk-1 and SRE regulated gene expression.
In response to netrin-1, DCC becomes tyrosine phosphorylated, localizes to lipid rafts and selectively interacts with the Src family kinases Fyn and Lck to mediate axon attraction.
Furthermore DCC induces apoptosis when unbound to its ligand netrin-1. The DCC cytoplasmic domain is required for the induction of apoptosis and contains a caspase cleavage site which is recognized by caspase 3 in vitro. Upstream of the caspase cleavage domain a proapoptotic domain named addiction dependence domain (ADD) is located, which interacts with caspase 9 in the absence of Netrin-1 and binds DCC-interacting protein-13a (DIP13-a) to mediate DCC-induced cell death.
DCCs downstream effects involve MAPK activation and subsequent activation of the transcription factor Elk-1 and SRE regulated gene expression.
Homology
DCC has a homolog in mammals named neogin and is conserved in chimpanzee, dog, cow, mouse, rat, zebrafish, Caenorhabditis elegans (UNC40) and Drosophila (Frazzled).
Mutations
Somatic
DCC mutations rarely occur in cancer. In colorectal cancer (CRC) the most common somatic mutations are 120 to 300 bp expansions in a dinucleotide repeat tract located in an intron region immediately downstream of exon 7. Expansions are present in 10-15 % of CRCs which are cancers with microsatellite instability.
Implicated in
Entity name
Colorectal cancer
Note
By regulating apoptosis in the absence of netrin-1, DCC is a conditional tumor suppressor. In normal conditions, DCC induced apoptosis limits cellular lifespan in the intestinal crypt and thereby inhibits the initiation of malignant transformation. Transfection of DCC cDNA into a human cell line lacking DCC expression suppresses tumor growth and results in apoptosis and cell cycle arrest.
DCC is located on chromosomal region 18q21-pter which is affected by loss of heterozygosity (LOH) which is associated with reduced DCC expression in approximately 70% of colorectal cancers.
Although DCC mutation is rare, DCC promoter CpG island methylation occurs in about 80% of CRCs.
DCC is located on chromosomal region 18q21-pter which is affected by loss of heterozygosity (LOH) which is associated with reduced DCC expression in approximately 70% of colorectal cancers.
Although DCC mutation is rare, DCC promoter CpG island methylation occurs in about 80% of CRCs.
Prognosis
Absent DCC expression is a strong predictor of poor survival in stage II and stage III CRCs. In patients with stage II disease decreased DCC expression is associated with a five-year survival rate of 61.6% versus 94.3% in DCC expressing stage II CRCs. In patients with stage III disease, the respective survival rates are 59.3 percent and 33.2 percent. LOH of 18q21 alone was also associated with poor prognosis and risk of metastasis in some studies although this association was not observed by others.
Furthermore 18q LOH is associated with decreases responsiveness to fluorouracil-based adjuvant chemotherapy in stage III CRC.
Furthermore 18q LOH is associated with decreases responsiveness to fluorouracil-based adjuvant chemotherapy in stage III CRC.
Oncogenesis
LOH of chromosome 18q21 profoundly occurs in progressed adenomas and colorectal carcinomas and is present in about 100% of hepatic metastasis but rarely occurs in early stage lesions. The same accounts for DCC promoter CpG island methylation which is present in 80% of adenomas and carcinomas and in only 23% of normal colon tissues.
Entity name
Gastric cancer
Note
Reduced DCC mRNA expression is observed in 52% of gastric cancers, being present in 72% of intestinal type gastric cancers and in 17% of infiltrative type gastric cancers.
LOH of 18q21 occurs in 30% of intestinal type gastric cancers and is an infrequent event in early or advanced gastric cancer.
LOH of 18q21 occurs in 30% of intestinal type gastric cancers and is an infrequent event in early or advanced gastric cancer.
Oncogenesis
All liver metastasis of gastric carcinomas showed reduced DCC expression.
Entity name
Head and neck squamous cell carcinoma (HNSCC)
Note
DCC promoter CpG island methylation occurs in 75% of HNSCC. Restoration of DCC expression (by transfection) led to inhibition of cell growth in HNSCC cell lines.
Prognosis
LOH of chromosomal region 18q21 occurs in 40% of HNSCC and is associated with poor patient survival.
Entity name
Esophageal cancer
Note
18q21 LOH occurs in 23% of esophageal squamous cell carcinomas (ESCC). DCC promoter CpG island methylation occurs in 74% of primary ESCCs in a cancer-specific manner.
Sixty-nine percent of esophageal adenocarcinomas show LOH of 18q21.
Sixty-nine percent of esophageal adenocarcinomas show LOH of 18q21.
Oncogenesis
18q21 LOH occurs in 32% of barrets mucosae, 42% of low-grade dysplastic lesions, 73% of high grade dysplastic lesions and 69% of adenocarcinomas.
Entity name
Glioma
Note
DCC expression is reduced in 66% of high-grade astrocytomas, 53% of secondary glioblastomas (progressed for low-grade astrocytomas) and 23% of de novo glioblastoma.
DCC expression is reduced in 88% of glioblastoma multiforme.
DCC expression is reduced in 88% of glioblastoma multiforme.
Oncogenesis
Only a minority (6%) of low-grade astrocytomas show reduced DCC expression, whereas high-grade tumors show reduces DCC expression in 66% of cases.
Entity name
Neuroblastoma
Note
Reduced DCC expression and 18q21 LOH occurs in 25-40% and 31% of primary neuroblastomas respectively.
Oncogenesis
In neuroblastomas decreased DCC expression increases from 25% in stage 1-3 to 72% in stage 4 disease to 81% in metastatic disease.
Entity name
Hematologic malignancies/Lymphoma
Note
DCC is inactivated in 30% of acute leukemias in 25% of chronic myelogenous leukemias (CML) and in 53% of non Hodgkins lymphoma.
In 18% of follicle centre cell lymphoma LOH of DCC is observed.
In 18% of follicle centre cell lymphoma LOH of DCC is observed.
Entity name
Bladder cancer
Note
LOH of 18q21 occurs in 36% of bladder carcinomas and 33% of human bladder transitional cell carcinomas (TCCs).
Entity name
Breast cancer
Note
DCC expression is reduced in 40-55.6% of primary breast cancer.
Prognosis
DCC expression is associated with longer relapse free and overall survival.
Entity name
Prostate cancer
Note
Eighty-five percent of prostate cancers exhibit decrease DCC expression compared to normal tissue. 18q21 LOH occurs in 26-31% prostate cancers.
Entity name
Renal cancer
Note
DCC protein expression is reduced in 40% of clear cell renal cell carcinomas (cRCC) and LOH of 18q21 occurs in 19% of cRCC.
18q21 LOH is observed in 20% of nephroblastomas.
18q21 LOH is observed in 20% of nephroblastomas.
Prognosis
Decreased DCC protein expression occurred more frequently in patients who died from the disease (63%) compared to patients who did not (36%). 18q21 LOH in nephroblastomas is associated with poor prognosis.
Entity name
Ovarian cancer
Note
DCC mRNA expression is reduced in 60% of epithelial ovarian cancer and in 50% of serous ovarian cancers.
Prognosis
Reduced DCC expression is associated with poor patient outcome in epithelial ovarian cancer which is also observed in a cohort treated with combined chemotherapy platinum-paclitaxel.
Oncogenesis
Reduces DCC mRNA expression is reported in only 10% of ovarian adenomas and 6% of borderline tumors compared with 60% in ovarian carcinomas.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 16971669 | 2006 | Automated quantitative analysis of DCC tumor suppressor protein in ovarian cancer tissue microarray shows association with beta-catenin levels and outcome in patients with epithelial ovarian cancer. | Bamias A et al |
| 12901278 | 2003 | Prognostic significance of the deleted in colorectal cancer gene protein expression in high-risk resected gastric carcinoma. | Bamias AT et al |
| 7510345 | 1994 | Somatic allelic loss at the DCC, APC, nm23-H1 and p53 tumor suppressor gene loci in human prostatic carcinoma. | Brewster SF et al |
| 11304690 | 2001 | Analysis of human meningiomas for aberrations of the MADH2, MADH4, APM-1 and DCC tumor suppressor genes on the long arm of chromosome 18. | Büschges R et al |
| 9609755 | 1998 | Prognostic significance of allelic lost at chromosome 18q21 for stage II colorectal cancer. | Carethers JM et al |
| 17018594 | 2006 | Deleted in colorectal cancer is a putative conditional tumor-suppressor gene inactivated by promoter hypermethylation in head and neck squamous cell carcinoma. | Carvalho AL et al |
| 8188295 | 1994 | The DCC gene: structural analysis and mutations in colorectal carcinomas. | Cho KR et al |
| 15050007 | 2004 | DCC protein expression in clear cell renal cell carcinoma. | Dekel Y et al |
| 19329758 | 2009 | Promoter CpG island hypermethylation- and H3K9me3 and H3K27me3-mediated epigenetic silencing targets the deleted in colon cancer (DCC) gene in colorectal carcinogenesis without affecting neighboring genes on chromosomal region 18q21. | Derks S et al |
| 11986622 | 2002 | Netrin-1-mediated axon outgrowth requires deleted in colorectal cancer-dependent MAPK activation. | Forcet C et al |
| 12770737 | 2003 | Genetic alterations in ovarian carcinoma: with specific reference to histological subtypes. | Fujita M et al |
| 8504411 | 1993 | Frequent loss of expression and loss of heterozygosity of the putative tumor suppressor gene DCC in prostatic carcinomas. | Gao X et al |
| 16337851 | 2005 | Deletion mapping of 18q in conventional renal cell carcinoma. | Hirata H et al |
| 10469217 | 1999 | Reduced expression of APC and DCC gene protein in breast cancer. | Ho KY et al |
| 1423299 | 1992 | Loss of heterozygosity involves multiple tumor suppressor genes in human esophageal cancers. | Huang Y et al |
| 8015568 | 1994 | Allelic loss of chromosome 18q and prognosis in colorectal cancer. | Jen J et al |
| 15491747 | 2004 | Expression of DCC and netrin-1 in normal human endometrium and its implication in endometrial carcinogenesis. | Kato HD et al |
| 7731713 | 1995 | The DCC gene suppresses the malignant phenotype of transformed human epithelial cells. | Klingelhutz AJ et al |
| 12908825 | 2003 | The expression of DCC protein in female breast cancer. | Koren R et al |
| 12471613 | 2003 | Quantification of expression of netrins, slits and their receptors in human prostate tumors. | Latil A et al |
| 12011067 | 2002 | Mediation of the DCC apoptotic signal by DIP13 alpha. | Liu J et al |
| 11387206 | 2001 | Netrin-1 acts as a survival factor via its receptors UNC5H and DCC. | Llambi F et al |
| 7674315 | 1995 | Hereditary nonpolyposis colorectal cancer: the syndrome, the genes, and historical perspectives. | Marra G et al |
| 15343335 | 2004 | Netrin-1 controls colorectal tumorigenesis by regulating apoptosis. | Mazelin L et al |
| 9796814 | 1998 | The DCC gene product induces apoptosis by a mechanism requiring receptor proteolysis. | Mehlen P et al |
| 8187090 | 1994 | Point mutations and allelic deletion of tumor suppressor gene DCC in human esophageal squamous cell carcinomas and their relation to metastasis. | Miyake S et al |
| 8632608 | 1996 | Loss of heterozygosity at the p53, RB, DCC and APC tumor suppressor gene loci in human bladder cancer. | Miyamoto H et al |
| 18302152 | 2008 | DCC promoter hypermethylation in esophageal squamous cell carcinoma. | Park HL et al |
| 9559345 | 1998 | Loss of 18q predicts poor survival of patients with squamous cell carcinoma of the head and neck. | Pearlstein RP et al |
| 8490178 | 1993 | DCC tumor suppressor gene is inactivated in hematologic malignancies showing monosomy 18. | Porfiri E et al |
| 8580831 | 1995 | DCC (deleted in colorectal cancer) inactivation in hematological malignancies. | Porfiri E et al |
| 14631365 | 2004 | Microsatellite analysis of the DCC gene in nephroblastomas: pathologic correlations and prognostic implications. | Ramburan A et al |
| 8724537 | 1996 | Allele imbalance at tumour suppressor loci during the indolent phase of follicle centre cell lymphoma. | Randerson J et al |
| 9816273 | 1996 | Loss of DCC expression in neuroblastoma is associated with disease dissemination. | Reale MA et al |
| 18253061 | 2008 | Tyrosine phosphorylation of netrin receptors in netrin-1 signaling. | Ren XR et al |
| 9645757 | 1998 | Status of deleted in colorectal cancer gene expression correlates with neuroblastoma metastasis. | Reyes-Mugica M et al |
| 10682668 | 2000 | Loss of DCC gene expression during ovarian tumorigenesis: relation to tumour differentiation and progression. | Saegusa M et al |
| 8242611 | 1993 | Expression of the tumor suppressor gene DCC in human gliomas. | Scheck AC et al |
| 8929264 | 1996 | The DCC protein and prognosis in colorectal cancer. | Shibata D et al |
| 7478611 | 1995 | Allelotype of neuroblastoma. | Takita J et al |
| 11309634 | 2001 | Molecular predictors of survival after adjuvant chemotherapy for colon cancer. | Watanabe T et al |
| 9136822 | 1997 | Genetic alterations in gastric cancer: relation to histological subtypes, tumor stage, and Helicobacter pylori infection. | Wu MS et al |
| 9665490 | 1998 | Genetic alterations in Barrett esophagus and adenocarcinomas of the esophagus and esophagogastric junction region. | Wu TT et al |
| 7742542 | 1995 | Decreased expression of the deleted in colorectal carcinoma gene in non-Hodgkin's lymphoma. | Younes A et al |
Other Information
Locus ID:
NCBI: 1630
MIM: 120470
HGNC: 2701
Ensembl: ENSG00000187323
Variants:
dbSNP: 1630
ClinVar: 1630
TCGA: ENSG00000187323
COSMIC: DCC
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38419228 | 2024 | Identification of specific codon 201 mutation of the DCC Gene in the colonoscopic specimen of colorectal cancer. | 0 |
| 38556604 | 2024 | NME1 and DCC variants are associated with susceptibility and tumor characteristics in Mexican patients with colorectal cancer. | 0 |
| 38419228 | 2024 | Identification of specific codon 201 mutation of the DCC Gene in the colonoscopic specimen of colorectal cancer. | 0 |
| 38556604 | 2024 | NME1 and DCC variants are associated with susceptibility and tumor characteristics in Mexican patients with colorectal cancer. | 0 |
| 36852451 | 2023 | An imbalance of netrin-1 and DCC during nigral degeneration in experimental models and patients with Parkinson's disease. | 2 |
| 36889039 | 2023 | Mirror movements and callosal dysgenesis in a family with a DCC mutation: Neuropsychological and neuroimaging outcomes. | 2 |
| 36852451 | 2023 | An imbalance of netrin-1 and DCC during nigral degeneration in experimental models and patients with Parkinson's disease. | 2 |
| 36889039 | 2023 | Mirror movements and callosal dysgenesis in a family with a DCC mutation: Neuropsychological and neuroimaging outcomes. | 2 |
| 34259111 | 2022 | Genetic Susceptibility of DCC Gene in Gallbladder Cancer in Kashmir and Meta-Analysis. | 2 |
| 34875255 | 2022 | Association between serum netrin-1, netrin-4 and risk of the acute coronary syndrome in patients with type 2 diabetes mellitus-A pilot study. | 0 |
| 35388180 | 2022 | Corticolimbic DCC gene co-expression networks as predictors of impulsivity in children. | 5 |
| 34259111 | 2022 | Genetic Susceptibility of DCC Gene in Gallbladder Cancer in Kashmir and Meta-Analysis. | 2 |
| 34875255 | 2022 | Association between serum netrin-1, netrin-4 and risk of the acute coronary syndrome in patients with type 2 diabetes mellitus-A pilot study. | 0 |
| 35388180 | 2022 | Corticolimbic DCC gene co-expression networks as predictors of impulsivity in children. | 5 |
| 33141514 | 2021 | A novel homozygous frameshift mutation in the DCC gene in a Pakistani family with autosomal recessive horizontal gaze palsy with progressive scoliosis-2 with impaired intellectual development. | 2 |
Citation
Sarah Derks ; Manon van Engeland
DCC (deleted in colorectal carcinoma)
Atlas Genet Cytogenet Oncol Haematol. 2010-01-01
Online version: http://atlasgeneticsoncology.org/gene/331/dcc
