LIMD1 (LIM domains containing 1)
2011-03-01 Sybil CK Wong , Gregory D Longmore , Tyson V Sharp AffiliationDNA/RNA
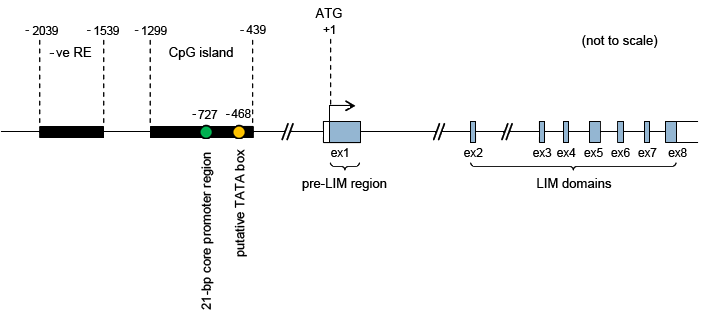
The eight exons of the LIMD1 gene are denoted by open rectangles; the protein coding regions of the exons are coloured in blue. Exons 2 to 8 encode three LIM domains in tandem. Exon 1 encodes the N-terminal pre-LIM region. Regions important for transcriptional regulation upstream of the start codon (ATG, +1) are denoted by black rectangles. Experimental evidence suggests that a negative regulatory element (-ve RE) occurs in the region between -2039 and -1539. The CpG island between -1299 and -439 contains a putative TATA box at -439 (yellow circle) and a 21-bp core promoter region at -727 (green circle).
Description
The gene encompasses 86432 bp and lies in the plus strand, starting at 45636323 and ending at 45722754 bp from pter, within C3CER1 (chromosome 3 common eliminated region 1) at chromosome 3p21.3, a region frequently deleted in solid tumours. The 5-promoter region includes a negative regulatory element and a CpG island that contains a putative TATA box and an experimentally validated core basal promoter region of 21 bp (Sharp et al., 2008). The core promoter region contains four CpG methylation sites (Sharp et al., 2008).
Transcription
The transcript (NCBI ID: NM_014240.2) of 6284 bp comprises eight exons, the first of which encodes the N-terminal pre-LIM region. Exons 2 to 8 encode three LIM domains in tandem. The size of the open reading frame is 5129 bp.
Pseudogene
None known.
Proteins

LIMD1 can be split primarily into the N-terminal pre-LIM region (residues 1 to 471) and the C-terminal triple LIM domains (residues 472 to 676). The pre-LIM region contains a proline/serine-rich region (residues 69 to 471), a LEM-like domain (residues 18 to 68), a nuclear export signal (NES, residues 54 to 134) and the binding site for pRB (residues 404 to 442). The three LIM domains are an inherent nuclear localisation signal (NLS, residues 472 to 676).
Description
LIMD1 (676 amino acids, 72 kDa, NCBI ID: NP_055055.1) belongs to the Ajuba LIM protein subgroup of the Ajuba/Zyxin protein family. This six-member family (see Homology) is characterised by a varied proline-rich pre-LIM region at the N-terminus followed by three homologous LIM domains arranged in tandem at the C-terminus. LIM domains (InterPro ID: IPR001781) coordinate two zinc ions in a tandem zinc-finger topology and mediate protein-protein interactions (Kadrmas and Beckerle, 2004). The three LIM domains of LIMD1 are an inherent nuclear localisation signal (NLS) (Sharp et al., 2004). The pre-LIM region includes the pRB binding region and a nuclear export signal (NES) partially overlapping the N-terminal LEM-like domain (Sharp et al., 2004).
Expression
LIMD1 is ubiquitously expressed, but at a lower level in the brain and testis (Kiss et al., 1999).
Localisation
LIMD1 shuttles between the cytoplasm and nucleus, but is not found in the nucleoli (Sharp et al., 2004). In the cytoplasm, LIMD1 colocalises with focal adhesions (Huggins and Andrulis, 2008; Petit et al., 2005), adherens junctions (Das Thakur et al., 2010) and mRNA processing bodies (P-bodies) (James et al., 2010).
Function
LIMD1 is involved in a range of cellular functions (listed below). A common theme is that LIMD1 acts as a protein-protein scaffold or mediator to facilitate signalling between cytoplasmic components and the nucleus. Functional redundancy between LIMD1 and the other Ajuba LIM proteins has also been proposed in a variety of cases (Das Thakur et al., 2010; Feng and Longmore, 2005; James et al., 2010; Langer et al., 2008).
- LIMD1 inhibits E2F-mediated transcription and thus cell proliferation via interaction with pRB and also via a pRB-independent mechanism that has yet to be clarified (Sharp et al., 2004).
- LIMD1 stimulates stress osteoclastogenesis (Feng et al., 2007) and regulates osteoblast differentiation and function (Luderer et al., 2008). LIMD1 can act as both a positive regulator of osteoclast and osteoblast development, by linking the p62/sequestosome protein and TRAF6 to form a p62/TRAF6/a-PKC complex which can activate NF-kB and AP-1 (Feng and Longmore, 2005; Feng et al., 2007), and a negative regulator of osteoblast differentiation, by repressing canonical Wnt/beta-catenin signalling in osteoblasts (Luderer et al., 2008).
- LIMD1 facilitates miRNA-mediated silencing by (i) binding to eIF4E to prevent recruitment of 4EBP1 and eIF4G to the mRNA 5 m(7)GTP cap, and (ii) creating an inhibitory closed-loop complex by acting as a scaffold to link eIF4E at the 5 cap to Ago1/Ago2 in the miRISC complex bound at the 3 UTR of mRNA (James et al., 2010).
- LIMD1 interacts with LATS/Warts and WW45/Sav to inhibit Hippo (Hpo) signalling pathway-mediated phosphorylation and nuclear-targeting of YAP/Yki (Das Thakur et al., 2010). Implication of the Hpo pathway in cell contact growth inhibition, and the observation that in confluent cells with no further proliferation, Ajuba LIM proteins gather at adherens junctions whilst YAP is phosphorylated and returned to the cytosol has led Das Thakur et al. (2010) to propose that Ajuba LIM proteins "release" LATS/Warts upon localisation at adherens junctions, leading to growth inhibition.
- In the nucleus, LIMD1 interacts with the SNAG domain of Snail/Slug transcriptional repressors and acts as a co-repressor of E-cadherin and other Snail-regulated genes to affect neural crest development and possibly epithelial-mesenchymal transitions (Langer et al., 2008).
- Huggins and Andrulis (2008) suggested that LIMD1 may play a part in cell anchoring as a component of focal adhesions and in the cell cycle (especially during mitosis), based on the respective observations that LIMD1 colocalises with vinculin at focal adhesions and that phosphorylation of LIMD1 occurs during mitosis in HeLa cells.
- LIMD1 inhibits E2F-mediated transcription and thus cell proliferation via interaction with pRB and also via a pRB-independent mechanism that has yet to be clarified (Sharp et al., 2004).
- LIMD1 stimulates stress osteoclastogenesis (Feng et al., 2007) and regulates osteoblast differentiation and function (Luderer et al., 2008). LIMD1 can act as both a positive regulator of osteoclast and osteoblast development, by linking the p62/sequestosome protein and TRAF6 to form a p62/TRAF6/a-PKC complex which can activate NF-kB and AP-1 (Feng and Longmore, 2005; Feng et al., 2007), and a negative regulator of osteoblast differentiation, by repressing canonical Wnt/beta-catenin signalling in osteoblasts (Luderer et al., 2008).
- LIMD1 facilitates miRNA-mediated silencing by (i) binding to eIF4E to prevent recruitment of 4EBP1 and eIF4G to the mRNA 5 m(7)GTP cap, and (ii) creating an inhibitory closed-loop complex by acting as a scaffold to link eIF4E at the 5 cap to Ago1/Ago2 in the miRISC complex bound at the 3 UTR of mRNA (James et al., 2010).
- LIMD1 interacts with LATS/Warts and WW45/Sav to inhibit Hippo (Hpo) signalling pathway-mediated phosphorylation and nuclear-targeting of YAP/Yki (Das Thakur et al., 2010). Implication of the Hpo pathway in cell contact growth inhibition, and the observation that in confluent cells with no further proliferation, Ajuba LIM proteins gather at adherens junctions whilst YAP is phosphorylated and returned to the cytosol has led Das Thakur et al. (2010) to propose that Ajuba LIM proteins "release" LATS/Warts upon localisation at adherens junctions, leading to growth inhibition.
- In the nucleus, LIMD1 interacts with the SNAG domain of Snail/Slug transcriptional repressors and acts as a co-repressor of E-cadherin and other Snail-regulated genes to affect neural crest development and possibly epithelial-mesenchymal transitions (Langer et al., 2008).
- Huggins and Andrulis (2008) suggested that LIMD1 may play a part in cell anchoring as a component of focal adhesions and in the cell cycle (especially during mitosis), based on the respective observations that LIMD1 colocalises with vinculin at focal adhesions and that phosphorylation of LIMD1 occurs during mitosis in HeLa cells.
Homology
The phylogenetic tree below shows the evolutionary distance (as inferred from amino acid sequence) between LIMD1 and the five other members of the Ajuba/Zyxin protein family (James et al., 2010).
LIMD1 is most closely related to the other Ajuba LIM proteins, Ajuba and WTIP (respectively 41% and 39% identity overall and 64% and 72% identity in the LIM domains). In contrast, LIMD1 shares only 32%, 33% and 34% identity overall and 43%, 46% and 41% identity in the LIM domains with the Zyxin LIM proteins, Zyxin, LPP and TRIP6 respectively.
The orthologue of LIMD1 in mus musculus shares 77% identity overall and 96% identity in the LIM domains with human LIMD1.
LIMD1 is most closely related to the other Ajuba LIM proteins, Ajuba and WTIP (respectively 41% and 39% identity overall and 64% and 72% identity in the LIM domains). In contrast, LIMD1 shares only 32%, 33% and 34% identity overall and 43%, 46% and 41% identity in the LIM domains with the Zyxin LIM proteins, Zyxin, LPP and TRIP6 respectively.
The orthologue of LIMD1 in mus musculus shares 77% identity overall and 96% identity in the LIM domains with human LIMD1.

Implicated in
Entity name
Lung cancer
Disease
Lung cancer is the primary cause of cancer-related deaths worldwide. Although smoking is the major trigger for lung cancer, another important risk factor involves exposure to harmful air-borne particles, such as asbestos or industrial pollutants.
Cytogenetics
Loss of coding sequence from chromosome 3p is one of the most common mutations found in both small cell (almost 100%) and non-small cell (almost 90%) lung cancers. The C3CER1 region (containing LIMD1) of chromosome 3p21.3 is more frequently deleted in solid malignancies than any of the other 3p regions studied (Petursdottir et al., 2004). Particularly in lung tumours, loss of heterozygosity (LOH) at C3CER1 (over 90%) exceeds that of tumour suppressor gene (TSG) VHL (72%) and putative TSG FHIT (65%) (Petursdottir et al., 2004). Homozygous gene deletion, LOH and epigenetic silencing via promoter hypermethylation - but not coding sequence mutations - contribute towards reduced expression of LIMD1 in human lung tumours (Sharp et al., 2008).
Oncogenesis
The expression of LIMD1, a bona fide lung tumour suppressor, was down-regulated in all lung cancers analysed compared with normal tissue (Sharp et al., 2008). From a genetic perspective, increased lung tumour incidence and volume was observed in the offspring of LIMD1 knockout (Limd1-/-) mice cross bred with K-RasG12D mice, an established K-Ras-induced lung cancer model (Sharp et al., 2008). LIMD1, which interacts with pRB to repress E2F-responsive transcription, reduces cell proliferation in vitro and incidence of lung metastases in vivo (Sharp et al., 2004). Loss of LIMD1 expression leading to misregulation of pRB and the cell cycle may therefore be an early critical step in lung tumour development (Sharp et al., 2004).
Entity name
Breast cancer
Disease
Breast cancer is the leading cause of cancer-related deaths of women in the western world. Increasing age, non-vegetarian diets, social class, nulliparity and familial history of breast cancer have been observed to increase the probability of breast cancer development.
Prognosis
LIMD1 has been proposed to be a prognostic marker for breast cancer, as absence of nuclear LIMD1 staining correlates strongly to poor tumour prognosis and patient mortality (Spendlove et al., 2008).
Cytogenetics
Although alteration or deletion of chromosome 3p is common in breast cancers, LIMD1 was not mutated or deleted in most of the breast tumours analysed (Huggins et al., 2007) and LIMD1 mRNA levels were also found to be unchanged in breast cancers compared to normal tissues (Huggins and Andrulis, 2008).
Oncogenesis
Delocalisation of LIMD1 from the nucleus may contribute to breast cancer progression, as nuclear LIMD1 acts as a Snail co-repressor of E-cadherin expression and Snail-mediated repression of E-cadherin has been implicated in tumour metastasis (Langer et al., 2008; Spendlove et al., 2008).
Entity name
Head and neck squamous cell carcinoma (HNSCC)
Disease
HNSCCs account for 90% of all head and neck cancers and 30-40% of cancer types in Indian patients. Globally, over 0,5 million new cases are diagnosed every year. Risk factors include smoking, alcohol, betel nut leaf liquid, HPV-16/18 infection and inadequate oral hygiene.
Prognosis
Ghosh et al. (2010) have suggested LIMD1 as a "predictive clinical marker" for HNSCC from their multivariate analysis showing significant correlation of LIMD1 alteration (deletion/methylation/mutation) to poor prognosis in HPV-negative patients addicted to tobacco.
Cytogenetics
Loss of heterozygosity in chromosome 3p21.3 was found to be highly frequent in Indian HNSCC patients (Ghosh et al., 2008). The same group later reported alteration of LIMD1 in the majority of HNSCC samples studied, and increasing mutation, but not deletion or methylation, with tumour progression (Ghosh et al., 2010). Six mutations in exon 1, an intron 4/exon 5 splice-junction mutation in LIMD1 and a (CA)20 polymorphism of the hmlimd1 microsatellite marker were reported, some of which lead to non-functional LIMD1 or reduced expression of LIMD1 (Ghosh et al., 2010).
Oncogenesis
Ghosh et al. (2010) have proposed that repression of LIMD1, rather than pRB, is crucial for HNSCC development, due to the higher frequency of LIMD1 alterations compared with pRB alterations in the HNSCC samples analysed and better survival of patients with LIMD1+/pRB- compared with LIMD1-/pRB+.
Entity name
Pagets disease of the bone (pagets disease)
Disease
Pagets disease involves pathological bone resorption triggering compensatory osteoblast activity, which results in the formation of hypercellular, deformed bones. This chronic disease can be familial or sporadic.
Prognosis
In the regulation of osteoclast and osteoblast development, LIMD1 interacts with cytosolic protein p62, which is sometimes mutated in Pagets disease (Feng et al., 2007; Luderer et al., 2008). As none of the reported Pagetic mutations of p62 are predicted to affect its binding to LIMD1, and coinciding phenotypes were observed between LIMD1-null mice and p62-null mice, it was suggested that LIMD1 may act as a complex with mutated p62 in the development of Pagets disease (Feng et al., 2007; Luderer et al., 2008).
Breakpoints
Note
N/A.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 20303269 | 2010 | Ajuba LIM proteins are negative regulators of the Hippo signaling pathway. | Das Thakur M et al |
| 15870274 | 2005 | The LIM protein Ajuba influences interleukin-1-induced NF-kappaB activation by affecting the assembly and activity of the protein kinase Czeta/p62/TRAF6 signaling complex. | Feng Y et al |
| 17092936 | 2007 | The LIM protein, Limd1, regulates AP-1 activation through an interaction with Traf6 to influence osteoclast development. | Feng Y et al |
| 20226061 | 2010 | LIMD1 is more frequently altered than RB1 in head and neck squamous cell carcinoma: clinical and prognostic implications. | Ghosh S et al |
| 18439753 | 2008 | Cell cycle regulated phosphorylation of LIMD1 in cell lines and expression in human breast cancers. | Huggins CJ et al |
| 20616046 | 2010 | LIM-domain proteins, LIMD1, Ajuba, and WTIP are required for microRNA-mediated gene silencing. | James V et al |
| 15520811 | 2004 | The LIM domain: from the cytoskeleton to the nucleus. | Kadrmas JL et al |
| 11156420 | 2000 | Inactivation of the human fragile histidine triad gene at 3p14.2 in monochromosomal human/mouse microcell hybrid-derived severe combined immunodeficient mouse tumors. | Kholodnyuk ID et al |
| 10647888 | 1999 | A novel gene containing LIM domains (LIMD1) is located within the common eliminated region 1 (C3CER1) in 3p21.3. | Kiss H et al |
| 18331720 | 2008 | Ajuba LIM proteins are snail/slug corepressors required for neural crest development in Xenopus. | Langer EM et al |
| 18657804 | 2008 | The LIM protein LIMD1 influences osteoblast differentiation and function. | Luderer HF et al |
| 16137684 | 2005 | The tumor suppressor Scrib selectively interacts with specific members of the zyxin family of proteins. | Petit MM et al |
| 15334546 | 2004 | Interstitial deletions including chromosome 3 common eliminated region 1 (C3CER1) prevail in human solid tumors from 10 different tissues. | Petursdottir TE et al |
| 19060205 | 2008 | The chromosome 3p21.3-encoded gene, LIMD1, is a critical tumor suppressor involved in human lung cancer development. | Sharp TV et al |
| 15542589 | 2004 | LIM domains-containing protein 1 (LIMD1), a tumor suppressor encoded at chromosome 3p21.3, binds pRB and represses E2F-driven transcription. | Sharp TV et al |
| 18712738 | 2008 | Differential subcellular localisation of the tumour suppressor protein LIMD1 in breast cancer correlates with patient survival. | Spendlove I et al |
| 17374122 | 2007 | Genetic abnormalities involved in t(12;21) TEL-AML1 acute lymphoblastic leukemia: analysis by means of array-based comparative genomic hybridization. | Tsuzuki S et al |
Other Information
Locus ID:
NCBI: 8994
MIM: 604543
HGNC: 6612
Ensembl: ENSG00000144791
Variants:
dbSNP: 8994
ClinVar: 8994
TCGA: ENSG00000144791
COSMIC: LIMD1
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000144791 | ENST00000273317 | Q9UGP4 |
| ENSG00000144791 | ENST00000440097 | C9JRJ5 |
Expression (GTEx)
Pathways
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 33891898 | 2021 | LIMD1 phase separation contributes to cellular mechanics and durotaxis by regulating focal adhesion dynamics in response to force. | 27 |
| 34080067 | 2021 | High nuclear expression of HIF1α, synergizing with inactivation of LIMD1 and VHL, portray worst prognosis among the bladder cancer patients: association with arsenic prevalence. | 2 |
| 33891898 | 2021 | LIMD1 phase separation contributes to cellular mechanics and durotaxis by regulating focal adhesion dynamics in response to force. | 27 |
| 34080067 | 2021 | High nuclear expression of HIF1α, synergizing with inactivation of LIMD1 and VHL, portray worst prognosis among the bladder cancer patients: association with arsenic prevalence. | 2 |
| 30600590 | 2019 | LIMD1 phosphorylation in mitosis is required for mitotic progression and its tumor-suppressing activity. | 7 |
| 30600590 | 2019 | LIMD1 phosphorylation in mitosis is required for mitotic progression and its tumor-suppressing activity. | 7 |
| 29440237 | 2018 | Tension-dependent regulation of mammalian Hippo signaling through LIMD1. | 53 |
| 29654110 | 2018 | Deregulation of LIMD1-VHL-HIF-1α-VEGF pathway is associated with different stages of cervical cancer. | 11 |
| 29672635 | 2018 | Differential transmission of the molecular signature of RBSP3, LIMD1 and CDC25A in basal/ parabasal versus spinous of normal epithelium during head and neck tumorigenesis: A mechanistic study. | 2 |
| 29930174 | 2018 | A HIF-LIMD1 negative feedback mechanism mitigates the pro-tumorigenic effects of hypoxia. | 13 |
| 30321376 | 2018 | Site specific target binding controls RNA cleavage efficiency by the Kaposi's sarcoma-associated herpesvirus endonuclease SOX. | 18 |
| 29440237 | 2018 | Tension-dependent regulation of mammalian Hippo signaling through LIMD1. | 53 |
| 29654110 | 2018 | Deregulation of LIMD1-VHL-HIF-1α-VEGF pathway is associated with different stages of cervical cancer. | 11 |
| 29672635 | 2018 | Differential transmission of the molecular signature of RBSP3, LIMD1 and CDC25A in basal/ parabasal versus spinous of normal epithelium during head and neck tumorigenesis: A mechanistic study. | 2 |
| 29930174 | 2018 | A HIF-LIMD1 negative feedback mechanism mitigates the pro-tumorigenic effects of hypoxia. | 13 |
Citation
Sybil CK Wong ; Gregory D Longmore ; Tyson V Sharp
LIMD1 (LIM domains containing 1)
Atlas Genet Cytogenet Oncol Haematol. 2011-03-01
Online version: http://atlasgeneticsoncology.org/gene/41158/limd1

