MAML2 (mastermind-like 2)
2007-10-01 Kazumi Suzukawa , Jean-Loup Huret AffiliationDepartment of Hematology, Institute of Clinical Medicine, University of Tsukuba (KS); Genetics, Dept Medical Information, University of Poitiers, CHU Poitiers Hospital, F-86021 Poitiers, France (JLH)
Identity
HGNC
LOCATION
11q21
LOCUSID
ALIAS
MAM-3,MAM2,MAM3,MLL-MAML2
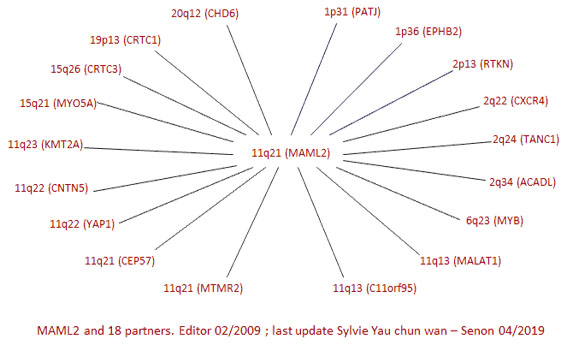
FUSION GENES
DNA/RNA
Description
Spans 365 kb; 5 exons.
Transcription
A major transcript of 7.5 kb.
Proteins
Description
1153 aa, 125 kDa; conserved N-terminal basic domain (aa 29-92) which binds to the ankyrin repeat domain of Notch receptors; two acidic domains (aa 263-360 and 1124-1153) and a C-terminal transcriptional activation domain.
Expression
Widely expressed.
Localisation
Nuclear granules.
Function
Mastermind-like coactivator for all four Notch receptors; forms a complex with the Notch intracellular domain (Notch ICD) and the CSL family of transcription factors (CSL: CBF1/RBP-jk, Suppressor of Hairless, LAG1), resulting in activation of the Notch target genes HES1 and HES5; functions as a CSL-dependent transcriptional coactivator for ligand-stimulated Notch.
Homology
MAML1 and MAML3.
Implicated in
Entity name
mucoepidermoid carcinoma with t(11;19)(q21-22;p13).
Disease
- Most common type of malignant salivary gland tumor;
The t(11;19) was found in samples from the three different sites. - Rare tumour in the thyroid.
The t(11;19) was found in samples from the three different sites. - Rare tumour in the thyroid.
Prognosis
- Mucoepidermoid carcinomas have an unpredictable behaviour.
- The CRTC1-MAML2 fusion transcript was found equally in low, intermediate and high grade tumours; however, tumours lacking the fusion transcript were significantly associated with metastases; they may represent a subset of aggressive tumours.
- In another study, the median survival for fusion-positive patients was greater than 10 years compared to 1.6 years for fusion-negative patients.
- The CRTC1-MAML2 fusion transcript was found equally in low, intermediate and high grade tumours; however, tumours lacking the fusion transcript were significantly associated with metastases; they may represent a subset of aggressive tumours.
- In another study, the median survival for fusion-positive patients was greater than 10 years compared to 1.6 years for fusion-negative patients.
Hybrid gene
CRTC1-MAML2; exon 1 of CRTC1 fused to exons 2-5 of MAML2. Note: CRTC1 is also known as MECT1, or WAMTP1.
Fusion protein
CRTC1-MAML2. In the fusion protein, the first 171 aa including the basic domain of MAML2 are replaced by 42 aa of CRTC1; there are no sequence similarities in the N-terminal domains of MAML2 and CRTC1. The fusion protein activates transcription of the Notch target gene HES1 independently of both Notch ligand and CSL.
Transforming activity of CRTC1-MAML2 fusion oncoprotein is mediated by mimicking constitutive activation of cAMP signaling, by activating CREB directly.
Transforming activity of CRTC1-MAML2 fusion oncoprotein is mediated by mimicking constitutive activation of cAMP signaling, by activating CREB directly.
Entity name
Warthins tumor with t(11;19)(q21-22;p13).
Note
In rare instances mucoepidermoid carcinoma may arise from or coexist with Warthins tumors.
Disease
Warthins tumor is a salivary gland neoplasm consisting of benign epithelial and lymphoid components; malignant transformation is extremely rare.
Hybrid gene
CRTC1-MAML2
Entity name
Clear Cell Hidradenomas of the skin with t(11;19)(q21-22;p13)
Disease
Clear Cell Hidradenomas of the skin are benign sweat gland tumors of eccrine duct origin.
Hybrid gene
CRTC1-MAML2; exon 1 of CRTC1 fused to exons 2 of MAML2.
Entity name
Disease
therapy-related acute leukemia and MDS.
Hybrid gene
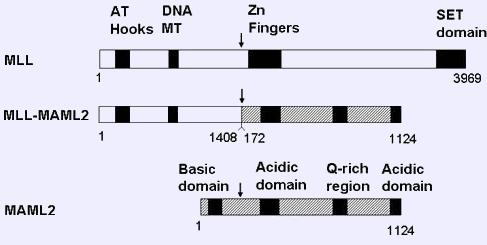
MLL-MAML2; exon 1-7 of MLL fused to exons 2-5 of MAML2.
Fusion protein
Hybrid transcript MLL/MAML2 contains the following domains: from MLL: AT-hook, DNA-Methyltransferase; from MAML2: Q rich domain, acidic domain.
Breakpoints
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 16444749 | 2006 | Molecular classification of mucoepidermoid carcinomas-prognostic significance of the MECT1-MAML2 fusion oncogene. | Behboudi A et al |
| 3180001 | 1988 | Translocation t(11;19)(q21;p13.1) as the sole chromosome abnormality in a cystadenolymphoma (Warthin's tumor) of the parotid gland. | Bullerdiek J et al |
| 14720503 | 2004 | Altered Notch signaling resulting from expression of a WAMTP1-MAML2 gene fusion in mucoepidermoid carcinomas and benign Warthin's tumors. | Enlund F et al |
| 1372276 | 1992 | Expression of an activated Notch-related int-3 transgene interferes with cell differentiation and induces neoplastic transformation in mammary and salivary glands. | Jhappan C et al |
| 12386158 | 2002 | Identification of new human mastermind proteins defines a family that consists of positive regulators for notch signaling. | Lin SE et al |
| 2331681 | 1990 | Chromosomal patterns in Warthin's tumor. A second type of human benign salivary gland neoplasm. | Mark J et al |
| 15269296 | 2004 | A study of MECT1-MAML2 in mucoepidermoid carcinoma and Warthin's tumor of salivary glands. | Martins C et al |
| 17551948 | 2007 | Identification of a novel fusion gene MLL-MAML2 in secondary acute myelogenous leukemia and myelodysplastic syndrome with inv(11)(q21q23). | Nemoto N et al |
| 8019965 | 1994 | Recurrent rearrangements of 11q14-22 in mucoepidermoid carcinoma. | Nordkvist A et al |
| 9870694 | 1998 | A child with a t(11;19)(q14-21;p12) in a pulmonary mucoepidermoid carcinoma. | Stenman G et al |
| 17437281 | 2007 | CRTC1/MAML2 fusion transcript in high grade mucoepidermoid carcinomas of salivary and thyroid glands and Warthin's tumors: implications for histogenesis and biologic behavior. | Tirado Y et al |
| 12539049 | 2003 | t(11;19)(q21;p13) translocation in mucoepidermoid carcinoma creates a novel fusion product that disrupts a Notch signaling pathway. | Tonon G et al |
| 17334997 | 2007 | Frequent fusion of the CRTC1 and MAML2 genes in clear cell variants of cutaneous hidradenomas. | Winnes M et al |
| 11101851 | 2000 | MAML1, a human homologue of Drosophila mastermind, is a transcriptional co-activator for NOTCH receptors. | Wu L et al |
| 15961999 | 2005 | Transforming activity of MECT1-MAML2 fusion oncoprotein is mediated by constitutive CREB activation. | Wu L et al |
| 12370315 | 2002 | Identification of a family of mastermind-like transcriptional coactivators for mammalian notch receptors. | Wu L et al |
| 9102301 | 1997 | Conservation of the Notch signalling pathway in mammalian neurogenesis. | de la Pompa JL et al |
Other Information
Locus ID:
NCBI: 84441
MIM: 607537
HGNC: 16259
Ensembl: ENSG00000184384
Variants:
dbSNP: 84441
ClinVar: 84441
TCGA: ENSG00000184384
COSMIC: MAML2
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000184384 | ENST00000524717 | Q8IZL2 |
| ENSG00000184384 | ENST00000618849 | A0A087X0G5 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36208053 | 2023 | Mucoepidermoid carcinoma may be devoid of squamoid cells by immunohistochemistry: expanding the histologic and immunohistochemical spectrum of MAML2- rearranged salivary gland tumours. | 6 |
| 36344258 | 2023 | Malignant undifferentiated epithelioid neoplasms with MAML2 rearrangements: A clinicopathologic study of seven cases demonstrating a heterogenous entity. | 3 |
| 36675234 | 2023 | Establishment of Mucoepidermoid Carcinoma Cell Lines from Surgical and Recurrence Biopsy Specimens. | 0 |
| 36898841 | 2023 | Pathogenic variations in MAML2 and MAMLD1 contribute to congenital hypothyroidism due to dyshormonogenesis by regulating the Notch signalling pathway. | 0 |
| 37489594 | 2023 | Significance of YAP1-MAML2 rearrangement and GTF2I mutation in the diagnosis and differential diagnosis of metaplastic thymoma. | 2 |
| 36208053 | 2023 | Mucoepidermoid carcinoma may be devoid of squamoid cells by immunohistochemistry: expanding the histologic and immunohistochemical spectrum of MAML2- rearranged salivary gland tumours. | 6 |
| 36344258 | 2023 | Malignant undifferentiated epithelioid neoplasms with MAML2 rearrangements: A clinicopathologic study of seven cases demonstrating a heterogenous entity. | 3 |
| 36675234 | 2023 | Establishment of Mucoepidermoid Carcinoma Cell Lines from Surgical and Recurrence Biopsy Specimens. | 0 |
| 36898841 | 2023 | Pathogenic variations in MAML2 and MAMLD1 contribute to congenital hypothyroidism due to dyshormonogenesis by regulating the Notch signalling pathway. | 0 |
| 37489594 | 2023 | Significance of YAP1-MAML2 rearrangement and GTF2I mutation in the diagnosis and differential diagnosis of metaplastic thymoma. | 2 |
| 34550633 | 2022 | KMT2A-MAML2 rearrangement emerged and regressed during neuroblastoma therapy without leukemia after 12.8-year follow-up. | 0 |
| 34657306 | 2022 | Salivary mucoepidermoid carcinoma: histological variants, grading systems, CRTC1/3-MAML2 fusions, and clinicopathological features. | 8 |
| 35457138 | 2022 | Diagnostic Value of MAML2 Rearrangements in Mucoepidermoid Carcinoma. | 1 |
| 35546636 | 2022 | Recurrent YAP1::MAML2 fusions in "nodular necrotizing" variants of myxoinflammatory fibroblastic sarcoma: a comprehensive study of 7 cases. | 6 |
| 35925563 | 2022 | MAML2 Gene Rearrangement Occurs in Nearly All Hidradenomas: A Reappraisal in a Series of 20 Cases. | 1 |
Citation
Kazumi Suzukawa ; Jean-Loup Huret
MAML2 (mastermind-like 2)
Atlas Genet Cytogenet Oncol Haematol. 2007-10-01
Online version: http://atlasgeneticsoncology.org/gene/472/maml2
Historical Card
2003-07-01 MAML2 (mastermind-like 2) by Goran Stenman Affiliation
Lundberg Laboratory for Cancer Research, Department of Pathology, Goteborg University, Sahlgrenska University Hospital, SE-413 45 Goteborg, Sweden
