Soft Tissues: Alveolar rhabdomyosarcoma
2009-01-01 Frederic G Barr Affiliation1.Department of Pathology, Laboratory Medicine, University of Pennsylvania School of Medicine, Philadelphia, PA 19104, USA
Summary
Note
Alveolar rhabdomyosarcoma (ARMS) refers to one subtype of the rhabdomyosarcoma family of soft tissue tumors. These are mesenchymal tumours related to the skeletal muscle lineage.
Classification
Classification
ARMS is one of two subtypes of rhabdomyosarcoma. The other major subype is embryonal rhabdosarcoma (ERMS). Within the ARMS subtype, there is no further histopathologic subtyping of clinical significance. Genetic subtyping has identified PAX3-FKHR, PAX7-FKHR, and fusion-negative subsets of ARMS; the clinical significance of these subsets is still being actively investigated.
Clinics and Pathology
Embryonic origin
ARMS tends to occur within skeletal muscle and is postulated to derive from precursor cells within the skeletal muscle. Therefore, the embryonic derivation of these tumors is presumed to be mesoderm, the same as that of skeletal muscle.
Epidemiology
ARMS accounts for 20-30% of all RMS tumors. Therefore ARMS represents ~1% of all malignancies among children and adolescents, and has an annual incidence of ~1 per million. Many ARMS tumors occur in older children and young adults - this age association characterizes the more prevalent PAX3-FKHR-positive subset of ARMS. In contrast, PAX7-FKHR-positive ARMS as well as fusion-negative tumors tend to occur in younger children.
Clinics
ARMS often occurs within skeletal muscle of the extremities but can also occur in other sites including the trunk and head and neck. This tumor often presents as a painless mass, but in other cases, may be discovered from symptoms produced by compression of structures at the primary site. A substantial fraction of patients with ARMS (25-30%) will have metastatic disease at the time of diagnosis. ARMS most frequently spreads to bone marrow, distal nodes, and bone. The standard treatment for ARMS is a combination of surgery, radiation, and intensive chemotherapy.
Pathology
Tumor cells in ARMS are relatively small with scant cytoplasm. They have round regular nuclei with a monotonous chromatin pattern. The cells form aggregates interrupted by fibrovascular septae, and within these aggregates, areas of discohesion often form, resulting in spaces that resemble alveoli of the lung. In some ARMS cases, there are few fibrovascular septae, no alveoli-like spaces, and a predominant cellular small round cell population; the term solid variant applies to this situation.
In addition to general immunohistochemical markers to identify RMS, certain markers aid in the identification of ARMS. Immunostaining for myogenin and MyoD show different patterns between ARMS and ERMS, such that most cells within an ARMS tumor stain positive whereas fewer cells within an ERMS tumor stain positive. In addition, based on microarray studies that distinguish fusion-postive ARMS from fusion-negative ERMS, AP2β and p-cadherin were found to be specific markers for the fusion-positive ARMS subtype.
In addition to general immunohistochemical markers to identify RMS, certain markers aid in the identification of ARMS. Immunostaining for myogenin and MyoD show different patterns between ARMS and ERMS, such that most cells within an ARMS tumor stain positive whereas fewer cells within an ERMS tumor stain positive. In addition, based on microarray studies that distinguish fusion-postive ARMS from fusion-negative ERMS, AP2β and p-cadherin were found to be specific markers for the fusion-positive ARMS subtype.
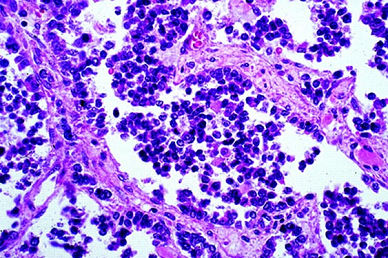
Histopathology of alveolar rhabdomyosarcoma (hematoxylin-eosin, original magnification: 100X; courtesy of Dr. Linda Ernst).
Prognosis
Patients with ARMS tumors have a poorer outcome than patients with ERMS tumors. The 4-year failure free survival rates for patients with localized and metastastic ARMS are 65% and 15%, respectively. Other risk factors that influence outcome of ARMS include primary site, size of primary tumor, extent of local spread, and the presence of nodal and distal metastases.
In an analysis of patients from the IRS-IV study, patients with localized PAX3-FKHR and PAX7-FKHR-positive ARMS had comparable outcomes. In contrast, among patients presenting with metastatic disease, those with PAX3-FKHR-positive tumors had a significantly poorer outcome than those with PAX7-FKHR-positive tumors (4-year overall survival rate of 8% compared to 75%, p=0.0015).
In an analysis of patients from the IRS-IV study, patients with localized PAX3-FKHR and PAX7-FKHR-positive ARMS had comparable outcomes. In contrast, among patients presenting with metastatic disease, those with PAX3-FKHR-positive tumors had a significantly poorer outcome than those with PAX7-FKHR-positive tumors (4-year overall survival rate of 8% compared to 75%, p=0.0015).
Cytogenetics
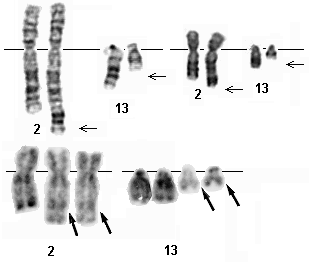
Examples of 2;13 translocation from ARMS tumors. G-banding (above) - Courtesy G. Reza Hafez, Eric B.Johnson, and Sara Morrison-Delap, Cytogenetics at the Waisman Center ; R- banding (below) - Courtesy Jean-Luc Lai.
Cytogenetics morphological
Most ARMS cases contain one of two recurrent chromosomal translocations: t(2;13)(q35;q14) or t(1;13)(p36;q14).Reciprocal balanced translocations are often present for the 2;13 translocation. The 1;13 is sometimes visible as a balanced translocation, and other times is associated with a subsequent amplification event.
Additional cytogenetic changes in ARMS include:
Amplification events involving 12q13-15, 2p24, 1p36, 13q14, 2q34-qter, 13q31 (as determined by comparative genomic hybridization studies).
Numerical chromosome gains, such as chromosomes 2, 12, and 20.
Additional cytogenetic changes in ARMS include:
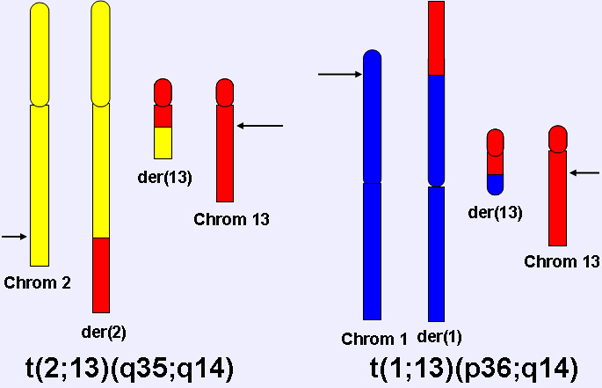
Diagram of t(2;13)(q35;q14) and t(1;13)(p36;q14) chromosomal translocations.
Genetics
Note
Most cases of ARMS occur sporadically without an apparent genetic predisposition.
Genes Involved and Proteins
Gene name
FOXO1 (Forkhead box O1)
Location
13q14.11
Protein description
Transcription factor.
Gene name
PAX3 (paired box gene 3 (Waardenburg syndrome 1))
Location
2q36.1
Protein description
Transcription factor.
Gene name
PAX7 (paired box 7)
Location
1p36.13
Protein description
Transcription factor.
Result of the chromosomal anomaly
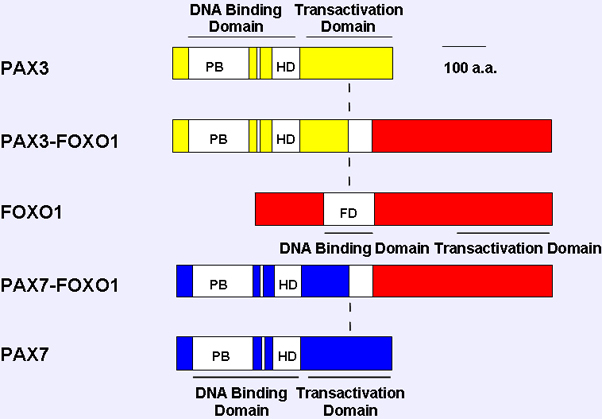
Comparison of wild-type and fusion products associated with the 2;13 and 1;13 translocations. The paired box, octapeptide, homeobox and fork head domain are indicated as open boxes, and transcriptional domains (DNA binding domain, DBD; transcriptional activiation domain) are shown as solid bars. The vertical dash line indicates the translocation fusion point.
Description
The PAX7-FKHR fusion is often amplified in tumors (70% of PAX7-FKHR-positive cases) whereas the PAX3-FKHR gene fusion is much less frequently amplified in tumors (5% of PAX3-FKHR-positive cases). Gene amplification appears to be one mechanism to increase the expression level of the gene fusion in ARMS tumor cells.
Description
These fusion genes encode fusion transcription factors with a PAX3 or PAX7 DNA binding domain and FOXO1 transactivation domain.
Expression localisation
Nuclear.
Oncogenesis
Transcription dysregulation.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 8098985 | 1993 | Rearrangement of the PAX3 paired box gene in the paediatric solid tumour alveolar rhabdomyosarcoma. | Barr FG et al |
| 8789435 | 1996 | In vivo amplification of the PAX3-FKHR and PAX7-FKHR fusion genes in alveolar rhabdomyosarcoma. | Barr FG et al |
| 11607823 | 2001 | Gene fusions involving PAX and FOX family members in alveolar rhabdomyosarcoma. | Barr FG et al |
| 12506174 | 2003 | Prognostic factors and clinical outcomes in children and adolescents with metastatic rhabdomyosarcoma--a report from the Intergroup Rhabdomyosarcoma Study IV. | Breneman JC et al |
| 11807989 | 2002 | Genomic gains and losses are similar in genetic and histologic subsets of rhabdomyosarcoma, whereas amplification predominates in embryonal with anaplasia and alveolar subtypes. | Bridge JA et al |
| 11408506 | 2001 | Intergroup rhabdomyosarcoma study-IV: results for patients with nonmetastatic disease. | Crist WM et al |
| 9223312 | 1997 | Fusion genes resulting from alternative chromosomal translocations are overexpressed by gene-specific mechanisms in alveolar rhabdomyosarcoma. | Davis RJ et al |
| 8187070 | 1994 | Fusion of PAX7 to FKHR by the variant t(1;13)(p36;q14) translocation in alveolar rhabdomyosarcoma. | Davis RJ et al |
| 8275086 | 1993 | Fusion of a fork head domain gene to PAX3 in the solid tumour alveolar rhabdomyosarcoma. | Galili N et al |
| 10825007 | 2000 | A novel and consistent amplicon at 13q31 associated with alveolar rhabdomyosarcoma. | Gordon AT et al |
| 9164192 | 1997 | Common and variant gene fusions predict distinct clinical phenotypes in rhabdomyosarcoma. | Kelly KM et al |
| 17090187 | 2006 | Rhabdomyosarcomas in adults and children: an update. | Parham DM et al |
| 12783965 | 2003 | Myogenin and MyoD1 expression in paediatric rhabdomyosarcomas. | Sebire NJ et al |
| 12039929 | 2002 | PAX3-FKHR and PAX7-FKHR gene fusions are prognostic indicators in alveolar rhabdomyosarcoma: a report from the children's oncology group. | Sorensen PH et al |
| 3943053 | 1986 | Consistent chromosomal translocation in alveolar rhabdomyosarcoma. | Turc-Carel C et al |
| 16391296 | 2006 | Subtype and prognostic classification of rhabdomyosarcoma by immunohistochemistry. | Wachtel M et al |
| 3338090 | 1988 | Chromosomal analysis of sixteen human rhabdomyosarcomas. | Wang-Wuu S et al |
| 8764111 | 1996 | Gains, losses, and amplification of genomic material in rhabdomyosarcoma analyzed by comparative genomic hybridization. | Weber-Hall S et al |
Citation
Frederic G Barr
Soft Tissues: Alveolar rhabdomyosarcoma
Atlas Genet Cytogenet Oncol Haematol. 2009-01-01
Online version: http://atlasgeneticsoncology.org/solid-tumor/5194/soft-tissues-alveolar-rhabdomyosarcoma
