inv(16)(p13q22) CBFB/MYH11
t(16;16)(p13;q22) CBFB/MYH11
del(16)(q22) CBFB/MYH11
1999-06-01 Jean-Loup Huret
Affiliation
1.Genetics, Dept Medical Information, University of Poitiers, CHU Poitiers Hospital, F-86021 Poitiers, France
Clinics and Pathology
Disease
acute non lymphoblastic leukaemia (AML); myelodysplastic syndromes (MDS) at times
Phenotype stem cell origin
nearly pathognomonic of M4eo-AML (all M4eo share the 16q22 anomaly -see also below-, but not all 16p13/16q22 are found in the M4eo subtype: i.e. this anomaly, although mainly found in M4-AML with marked eosinophilia, may (rarely) been found in : M2 or M5, M4 without eo, or in MDS; there are also known cases of chronic myelogenous leukaemia in blast crisis (BC-CML) with a M4 eo phenotype and inv(16); found at times in treatment related AML; 3 cases of infant leukaemia so far described; note: CD2 (T-cell marker) may be co-expressed
Epidemiology
5-10% of AML, 20% of M4
Clinics
CNS involvement is frequent, according to some authors, in particular at relapse
Cytology
most often: eosinophils > 5%, with large immature basophilic granules, NASCA+, in the bone marrow (but normal in blood: this M4 do not show the eo characteristic in blood)
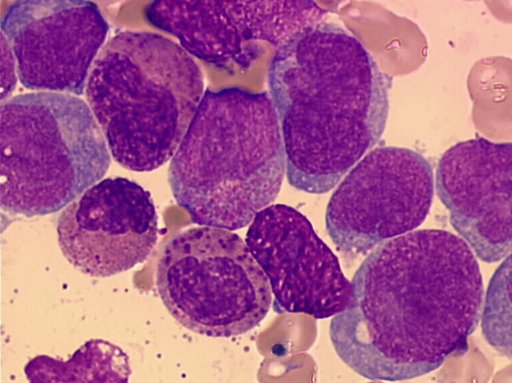
Patients with inv(16) usually correspond to the subclass of AML M4, with a specific abnormal eosinophil component and is considered as a distinct entity in correlation with these specific chromosomal abnormalities. These cases of AML M4 are referred as AML M4EO. In addition to the morphological features of AML M4 excess of monocytes), the bone marrow shows a variable number of eosinophils at all stages of maturation without significant maturation arrest. The most striking abnormalities involve the immature eosinophilic granules. Those are mainly evident at the promyelocyte and myelocyte stages. The abnormalities are not usually evident at later stages of maturation. These eosinophilic granules are often larger than those normally seen in immature eosinophils, purple-violet in color and in some cells are so dense that they obscure the cell morphology - Text and iconography Courtesy Georges Flandrin 2001.
Prognosis
high CR rate; better prognosis than most other AML; median survival may be 5yrs
Cytogenetics
Cytogenetics morphological
may be overlooked, especially with R-banding; best seen without banding procedure (giemsa) for some workers
Cytogenetics molecular
with 16p13 probes : as a deletion within 16p13 often accompany the 16p13/16q22 rearrangement (in 20% of cases), the split signal may be lost
Additional anomalies
none 2/3 of cases; +8, +22 in 15% each, del(7q), +2; apparently without prognostic significance
Variants
are known:
1- t(16;16)(p13;q22);
- del(16)(q22): may be associated with less typical phenotype and preceding MDS, older age, complex karyotype, worse prognosis;
1- t(16;16)(p13;q22);
- del(16)(q22): may be associated with less typical phenotype and preceding MDS, older age, complex karyotype, worse prognosis;
2- but also: translocations of 16q22 with various partners in: t(1;16)(p31-32;q22), t(3;16)(q21;q22), t(5;16)(q33;q22), associated with eosinophils anomalies
Genes Involved and Proteins
Gene name
MYH11 (myosin heavy chain) (incomplete)
Location
16p13.11
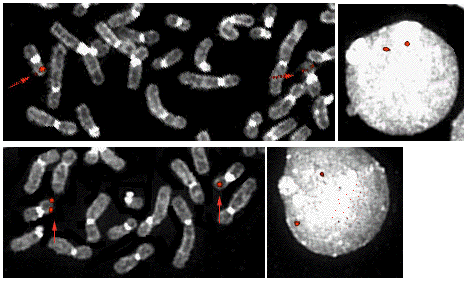
c-MYH11 (16p13) in normal cells: PAC 1032E3 (top) and PAC 1179J13 (below) - Courtesy Mariano Rocchi, Resources for Molecular Cytogenetics.
Protein description
contains a N-term ATPase head responsible for actin binding and mechanical movement, and a C-term long repeat of coil-coil domain to facilitate filament aggregates; member of the myosin II family
Gene name
CBFB (subunit b of core binding factor)
Location
16q22.1
Protein description
subunit of the transcription factor complex CBF; CBFb by itself does not contain any DNA binding motif or transcriptional activation domain, but forms a dimer with CBFa: --> transcription factor
Result of the Chromosomal Anomaly
Description
5 CBFb - 3 MYH11; breakpoint in CBFB intron n° 5 and in MYH11 intron A (i.e. : 5)
Transcript
at least 8 different CBFb-MYH11 fusion transcripts have been described, transcript type A (with positions at nucleotides 495 and 1921 respectively) being found in about 90% of the patients;most braekpoints in MYH11 are also clustered; no reciprocal MYH11-CBFB transcript
Description
N-term - the first 165 (or 133 in a few cases) amino acids of CBFb, removing only 17 or 22 amino acids fused to the tail of MYH11 C-term with its multimerization domain; also variable breakpoint in MYH11; identical fusion protein in the cases of RAEBT and BC-CML
Expression localisation
nuclear localisation
Oncogenesis
the fusion protein seems both to diminuish the quantity of active CBF and to compete with it, there is accumulation of CBFb-MYH11/CBFa multimeres in the nucleus
Highly cited references
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 31817911 | 2019 | Core Binding Factor Leukemia: Chromatin Remodeling Moves Towards Oncogenic Transcription. | 150 |
| 34440157 | 2021 | How to Improve Prognostication in Acute Myeloid Leukemia with CBFB-MYH11 Fusion Transcript: Focus on the Role of Molecular Measurable Residual Disease (MRD) Monitoring. | 123 |
| 38061017 | 2023 | Proteogenomic analysis reveals cytoplasmic sequestration of RUNX1 by the acute myeloid leukemia-initiating CBFB::MYH11 oncofusion protein. | 112 |
| 32831296 | 2020 | CBFB-MYH11 fusion neoantigen enables T cell recognition and killing of acute myeloid leukemia. | 93 |
| 36874378 | 2023 | Therapy-related core binding factor acute myeloid leukemia. | 86 |
| 29755956 | 2018 | Preleukemia and Leukemia-Initiating Cell Activity in inv(16) Acute Myeloid Leukemia. | 81 |
| 34804441 | 2021 | Gastrointestinal Myeloid Sarcoma a Case Presentation and Review of the Literature. | 80 |
| 27798625 | 2016 | The genomic landscape of core-binding factor acute myeloid leukemias. | 79 |
| 37019972 | 2023 | Etiology of oncogenic fusions in 5,190 childhood cancers and its clinical and therapeutic implication. | 71 |
| 34686664 | 2021 | Targeting miR-126 in inv(16) acute myeloid leukemia inhibits leukemia development and leukemia stem cell maintenance. | 70 |
| 34336831 | 2021 | CBFB-MYH11 Fusion Sequesters RUNX1 in Cytoplasm to Prevent DNMT3A Recruitment to Target Genes in AML. | 67 |
| 20589720 | 2010 | RUNX1 repression-independent mechanisms of leukemogenesis by fusion genes CBFB-MYH11 and AML1-ETO (RUNX1-RUNX1T1). | 57 |
| 35973983 | 2022 | Phenotypically-defined stages of leukemia arrest predict main driver mutations subgroups, and outcome in acute myeloid leukemia. | 52 |
| 17185462 | 2007 | CBFB-MYH11 hinders early T-cell development and induces massive cell death in the thymus. | 50 |
| 22160378 | 2012 | KIT with D816 mutations cooperates with CBFB-MYH11 for leukemogenesis in mice. | 50 |
| 36011278 | 2022 | PPP1R7 Is a Novel Translocation Partner of CBFB via t(2;16)(q37;q22) in Acute Myeloid Leukemia. | 49 |
| 20007544 | 2010 | Cbfb/Runx1 repression-independent blockage of differentiation and accumulation of Csf2rb-expressing cells by Cbfb-MYH11. | 48 |
| 36442087 | 2022 | Transcriptome-based molecular subtypes and differentiation hierarchies improve the classification framework of acute myeloid leukemia. | 47 |
| 32929473 | 2020 | RUNX1 and CBFβ-SMMHC transactivate target genes together in abnormal myeloid progenitors for leukemia development. | 47 |
| 31624376 | 2020 | Gata2 deficiency delays leukemogenesis while contributing to aggressive leukemia phenotype in Cbfb-MYH11 knockin mice. | 46 |
| 15044690 | 2004 | Identification of genes that synergize with Cbfb-MYH11 in the pathogenesis of acute myeloid leukemia. | 44 |
| 30742053 | 2019 | IL1RL1 is dynamically expressed on Cbfb-MYH11(+) leukemia stem cells and promotes cell survival. | 43 |
| 34771519 | 2021 | CBFB Break-Apart FISH Testing: An Analysis of 1629 AML Cases with a Focus on Atypical Findings and Their Implications in Clinical Diagnosis and Management. | 42 |
| 29018080 | 2017 | Chd7 deficiency delays leukemogenesis in mice induced by Cbfb-MYH11. | 41 |
| 31896782 | 2020 | The clinical mutatome of core binding factor leukemia. | 40 |
| 28030795 | 2017 | AML associated oncofusion proteins PML-RARA, AML1-ETO and CBFB-MYH11 target RUNX/ETS-factor binding sites to modulate H3ac levels and drive leukemogenesis. | 40 |
| 36452349 | 2022 | Analytical study of RUNX1-RUNXT1, PML-RARA, CBFB-MYH11, BCR-ABL1(p210) , and KMT2-MLLT3 in Mexican children with acute myeloid leukemia: A multicenter study of the Mexican interinstitutional group for the identification of the causes of childhood leukemia (MIGICCL). | 39 |
| 21638519 | 2011 | CBFB and MYH11 in inv(16)(p13q22) of acute myeloid leukemia displaying close spatial proximity in interphase nuclei of human hematopoietic stem cells. | 39 |
| 38427924 | 2024 | Targeting Molecular Measurable Residual Disease and Low-Blast Relapse in AML With Venetoclax and Low-Dose Cytarabine: A Prospective Phase II Study (VALDAC). | 37 |
| 35794567 | 2022 | Molecular dissection of a hyper-aggressive CBFB-MYH11/FLT3-ITD-positive acute myeloid leukemia. | 37 |
| 34547772 | 2021 | CBFB-MYH11 fusion transcripts distinguish acute myeloid leukemias with distinct molecular landscapes and outcomes. | 35 |
| 25742748 | 2015 | Runx1 is required for hematopoietic defects and leukemogenesis in Cbfb-MYH11 knock-in mice. | 34 |
| 38493200 | 2024 | Application of droplet digital PCR in minimal residual disease monitoring of rare fusion transcripts and mutations in haematological malignancies. | 34 |
| 31899799 | 2020 | Prospective evaluation of prognostic impact of KIT mutations on acute myeloid leukemia with RUNX1-RUNX1T1 and CBFB-MYH11. | 33 |
| 25266220 | 2014 | CBFB-MYH11 hypomethylation signature and PBX3 differential methylation revealed by targeted bisulfite sequencing in patients with acute myeloid leukemia. | 33 |
| 35445594 | 2022 | Comprehensive Mutation Profile in Acute Myeloid Leukemia Patients with RUNX1-RUNX1T1 or CBFB-MYH11 Fusions. | 31 |
| 35497938 | 2022 | Therapy-related acute myeloid leukemia: A case series. | 30 |
| 33204999 | 2020 | Identification of Fusion Gene Breakpoints is Feasible and Facilitates Accurate Sensitive Minimal Residual Disease Monitoring on Genomic Level in Patients With PML-RARA, CBFB-MYH11, and RUNX1-RUNX1T1. | 15 |
| 37225965 | 2023 | Characterization of leukemia progression in the Cbfb-MYH11 knockin mice by single cell RNA sequencing. | 14 |
| 33816105 | 2021 | Primary peritoneal myeloid sarcoma in association with CBFB/MYH11 fusion. | 13 |
| 23236551 | 2013 | Acute myeloid leukemia with cryptic CBFB-MYH11 type D. | 10 |
| 34512985 | 2021 | Abnormal eosinophils with immature eosinophilic granules in chronic myeloid leukemia in accelerated phase. | 2 |
| 38153768 | 2023 | CBFB::MYH11 fusion in a nonmonocytic acute myeloid leukemia. | 0 |
| 11561156 | 2001 | Function of the inv(16) fusion gene CBFB-MYH11. | 0 |
| 31671434 | 2020 | Myeloid Sarcoma With CBFB-MYH11 Fusion (inv(16) or t(16;16)) Prevails in the Abdomen. | 0 |
| 34183780 | 2021 | Prognostic values of D816V KIT mutation and peri-transplant CBFB-MYH11 MRD monitoring on acute myeloid leukemia with CBFB-MYH11. | 0 |
| 28906004 | 2017 | Acute myeloid leukaemia (FAB AML-M4Eo) with cryptic insertion of cbfb resulting in cbfb-Myh11 fusion. | 0 |
| 38388794 | 2024 | Clinical implications of additional chromosomal abnormalities in adult acute myeloid leukemia with inv (16)/t(16;16)/CBFB::MYH11. | 0 |
| 12239155 | 2002 | Role of Cbfb in hematopoiesis and perturbations resulting from expression of the leukemogenic fusion gene Cbfb-MYH11. | 0 |
| 36653696 | 2023 | Comprehensive molecular understanding of pediatric acute myeloid leukemia. | 0 |
| 32202250 | 2021 | Detection of del(16q) using the CBFB-MYH11 translocation dual fusion probe. | 0 |
| 31353165 | 2020 | Detection of a novel CBFB-MYH11 fusion transcript in acute myeloid leukemia M1 with inv(16)(p13q22). | 0 |
| 32588434 | 2021 | The loss or absence of minimal residual disease of <0·1% at any time after two cycles of consolidation chemotherapy in CBFB-MYH11-positive acute myeloid leukaemia indicates poor prognosis. | 0 |
| 11241787 | 2001 | Detection and quantification of CBFB/MYH11 fusion transcripts in patients with inv(16)-positive acute myeloblastic leukemia by real-time RT-PCR. | 0 |
| 35128634 | 2022 | TET2 deficiency cooperates with CBFB-MYH11 to induce acute myeloid leukaemia and represents an early leukaemogenic event. | 0 |
| 36039018 | 2022 | Mutual exclusivity between the fusion gene CBFB::MYH11 and somatic mitochondrial mutations in acute myeloid leukaemia. | 0 |
| 15156186 | 2004 | Mechanism of leukemogenesis by the inv(16) chimeric gene CBFB/PEBP2B-MHY11. | 0 |
| 20513535 | 2010 | Acute myeloid leukemia with inv(16) with CBFB-MYH11, 3'CBFB deletion, variant t(9;22) with BCR-ABL1, and del(7)(q22q32) in a pediatric patient: case report and literature review. | 0 |
| 22032582 | 2011 | Core-binding factor acute myeloid leukemia. | 0 |
| 28299658 | 2017 | Clinical Relevance of RUNX1 and CBFB Alterations in Acute Myeloid Leukemia and Other Hematological Disorders. | 0 |
| 8751466 | 1996 | The gene product of CBFB-MYH11. | 0 |
| 30919505 | 2019 | Measurable residual disease testing for personalized treatment of acute myeloid leukemia. | 0 |
| 28299661 | 2017 | Molecular Basis and Targeted Inhibition of CBFβ-SMMHC Acute Myeloid Leukemia. | 0 |
| 8929537 | 1996 | Failure of embryonic hematopoiesis and lethal hemorrhages in mouse embryos heterozygous for a knocked-in leukemia gene CBFB-MYH11. | 0 |
| 8585960 | 1996 | Transforming properties of the leukemic inv(16) fusion gene CBFB-MYH11. | 0 |
| 24002588 | 2014 | CBFB-MYH11/RUNX1 together with a compendium of hematopoietic regulators, chromatin modifiers and basal transcription factors occupies self-renewal genes in inv(16) acute myeloid leukemia. | 0 |
| 31721036 | 2020 | Mature neutrophils with Auer rod bundles in CBFB-MYH11-positive acute myeloid leukemia. | 0 |
| 7795233 | 1995 | RT-PCR diagnosis of patients with acute nonlymphocytic leukemia and inv(16)(p13q22) and identification of new alternative splicing in CBFB-MYH11 transcripts. | 0 |
| 17287858 | 2007 | Rare CBFB-MYH11 fusion transcripts in AML with inv(16)/t(16;16) are associated with therapy-related AML M4eo, atypical cytomorphology, atypical immunophenotype, atypical additional chromosomal rearrangements and low white blood cell count: a study on 162 patients. | 0 |
| 30006258 | 2018 | Molecular Minimal Residual Disease Testing in Acute Myeloid Leukemia: A Review for the Practicing Clinician. | 0 |
| 19215788 | 2009 | Detection of a novel CBFB/MYH11 variant fusion transcript (K-type) showing partial insertion of exon 6 of CBFB gene using two commercially available multiplex RT-PCR kits. | 0 |
| 38801226 | 2024 | Integrative immunophenotypic and genetic characterization of acute myeloid leukemia with CBFB rearrangement. | 0 |
| 25348871 | 2015 | Prognosis and monitoring of core-binding factor acute myeloid leukemia: current and emerging factors. | 0 |
| 27650511 | 2018 | Monitoring of post-transplant CBFB-MYH11 as minimal residual disease, rather than KIT mutations, can predict relapse after allogeneic haematopoietic cell transplantation in adults with inv(16) acute myeloid leukaemia. | 0 |
| 25635758 | 2015 | Emerging diagnostic and therapeutic approaches in core binding factor acute myeloid leukaemia. | 0 |
| 21281226 | 2011 | De novo acute myeloid leukemia with Philadelphia chromosome (BCR-ABL) and inversion 16 (CBFB-MYH11): report of two cases and review of the literature. | 0 |
| 27468869 | 2016 | Incidence of preleukemic fusion genes in healthy subjects. | 0 |
| 7577615 | 1995 | Detection of CBFB/MYH11 transcripts in patients with inversion and other abnormalities of chromosome 16 at presentation and remission. | 0 |
| 17052753 | 2007 | Detection of the CBFB/MYH11 fusion gene in de novo acute myeloid leukemia (AML): a single-institution study of 224 Japanese AML patients. | 0 |
| 39803562 | 2024 | Loss of the Inv(16) Oncogene CBFB::MYH11 Eliminates Leukemia from the Blood and Spleen, but not the Bone Marrow. | 0 |
| 37102302 | 2023 | TP53 and RB1 alterations characterize poor prognostic subgroups in pediatric acute myeloid leukemia. | 0 |
| 35622005 | 2022 | AML with inv(16)/t(16;16) and high-risk cytogenetic abnormalities: atypical features and unfavorable outcome. | 0 |
| 38654658 | 2024 | Lower relapse incidence with haploidentical versus matched sibling or unrelated donor hematopoietic cell transplantation for core-binding factor AML patients in CR2: A study from the Global Committee and the Acute Leukemia Working Party of the European Society for Blood and Marrow Transplantation. | 0 |
| 12139724 | 2002 | Minimal residual disease quantification in patients with acute myeloid leukaemia and inv(16)/CBFB-MYH11 gene fusion. | 0 |
| 38220149 | 2023 | [Acute myeloid leukemia with type I CBFB::MYH11 fusion gene not detected by screening test for leukemia-related chimeric genes]. | 0 |
| 33162510 | 2020 | [Clinical significance of MRD in AML]. | 0 |
| 28475434 | 2017 | Enhancing acute myeloid leukemia therapy - monitoring response using residual disease testing as a guide to therapeutic decision-making. | 0 |
| 38084551 | 2023 | Therapy-related acute myeloid leukemia with CBFB/MYH11 in a patient with ovarian cancer after exposure to chemotherapy. | 0 |
| 8630430 | 1996 | Novel CBFB-MYH11 fusion transcripts or reverse transcription-polymerase chain reaction artifacts? | 0 |
| 12788791 | 2003 | Rapid identification of CBFB-MYH11-positive acute myeloid leukemia (AML) cases by one single MYH11 real-time RT-PCR. | 0 |
| 35184217 | 2022 | 3'CBFB deletion in CBFB-rearranged acute myeloid leukemia retains morphological features associated with inv(16), but patients have higher risk of relapse and may require stem cell transplant. | 0 |
| 27512765 | 2016 | Wilms’ Tumor Gene (WT1) Expression and Minimal Residual Disease in Acute Myeloid Leukemia. | 0 |
| 16502584 | 2006 | Diagnosis and monitoring of CBFB-MYH11-positive acute myeloid leukemia by qualitative and quantitative RT-PCR. | 0 |
| 31399807 | 2019 | BCR-ABL1- and CBFB-MYH11-positive chronic myeloid leukemia presenting with primary blast crisis and marrow fibrosis. | 0 |
| 20625124 | 2010 | Prognostic impact of minimal residual disease in CBFB-MYH11-positive acute myeloid leukemia. | 0 |
| 16193088 | 2005 | Quantification of CBFB-MYH11 fusion gene levels in paired peripheral blood and bone marrow samples by real-time PCR. | 0 |
| 35613501 | 2022 | Jumping translocation involving chromosome 13q in a patient with Crohn's Disease and inv(16)(p13.1q22)/CBFB-MYH11 acute myeloid leukemia. | 0 |
| 23878140 | 2013 | Exome sequencing identifies recurring FLT3 N676K mutations in core-binding factor leukemia. | 0 |
| 19901261 | 2010 | Strikingly different molecular relapse kinetics in NPM1c, PML-RARA, RUNX1-RUNX1T1, and CBFB-MYH11 acute myeloid leukemias. | 0 |
| 29517504 | 2018 | Somatic Mutations and Intratumoral Heterogeneity of MYH11 Gene in Gastric and Colorectal Cancers. | 0 |
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 2779289 | 1989 | Acute nonlymphocytic leukemia with marrow eosinophilia and chromosome 16 abnormality: a report of 18 cases. | Bernard P et al |
| 1453770 | 1992 | Abnormalities of chromosome 16q in myeloid malignancy: 14 new cases and a review of the literature. | Betts DR et al |
| 2069909 | 1991 | Chromosome 16 abnormalities associated with myeloid malignancies. | Campbell LJ et al |
| 8268905 | 1993 | Cloning the breakpoint cluster region of the inv(16) in acute nonlymphocytic leukemia M4 Eo. | Dauwerse JG et al |
| 8338941 | 1993 | Identification of yeast artificial chromosomes containing the inversion 16 p-arm breakpoint associated with acute myelomonocytic leukemia. | Liu P et al |
| 7727763 | 1995 | Molecular pathogenesis of the chromosome 16 inversion in the M4Eo subtype of acute myeloid leukemia. | Liu PP et al |
| 7780153 | 1995 | Heterogeneity in CBF beta/MYH11 fusion messages encoded by the inv(16)(p13q22) and the t(16;16)(p13;q22) in acute myelogenous leukemia. | Shurtleff SA et al |
| 9927211 | 1999 | Genomic acute myeloid leukemia-associated inv(16)(p13q22) breakpoints are tightly clustered. | van der Reijden BA et al |
Summary
Fusion gene
CBFB/MYH11 CBFB (16q22.1) MYH11 (16p13.11) M ins(16)(q22p13p13) ins(16;16)(q22;p13p13) inv(16)(p13q22) t(16;16)(p13;q22)|CBFB/MYH11 CBFB (16q22.1) MYH11 (16p13.11) TIC
Note
the three chromosome anomalies are variants of each other, and they share identical clinical features and genetic pathogenesis
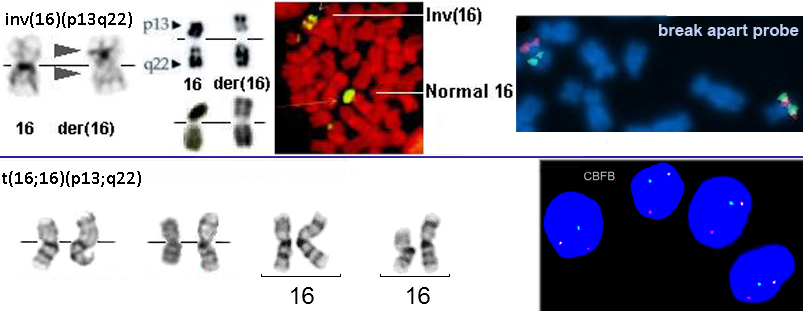
TOP: inv(16)(p13q22) G- banding (left) - Courtesy Jean-Luc Lai and Alain Vanderhaegen; R- banding center bottom: - Courtesy Christiane Charrin, center top and FISH (middle right) - Courtesy Pascale Cornillet-Lefebvre and Stephanie Struski; FISH (right) - Courtesy Hossein Mossafa; commercial FISH probes, split in the inv(16). BOTTOM: t(16;16)(p13;q22) G-banding - left two - Courtesy Diane H. Norback, Eric B. Johnson, and Sara Morrison-Delap, UW Cytogenetic Services; middle two and FSIH Vysis LSI CBFB Dual Color, Break Apart Rearrangement Probe ?nuc ish(CBFBx2)(5CBFB sep 3CBFBx1) - Courtesy Ian Brooks and Jackie Challis, Oncology Cytogenetics, Victorian Clinical Genetics Services, Australia.
Citation
Jean-Loup Huret
inv(16)(p13q22) CBFB/MYH11
t(16;16)(p13;q22) CBFB/MYH11
del(16)(q22) CBFB/MYH11
Atlas Genet Cytogenet Oncol Haematol. 1999-06-01
Online version: http://atlasgeneticsoncology.org/haematological/1036/inv(16)(p13q22)
Historical Card
1997-11-01 inv(16)(p13q22) CBFB/MYH11
t(16;16)(p13;q22) CBFB/MYH11
del(16)(q22) CBFB/MYH11 by Jean-Loup Huret
Affiliation
Genetics, Dept Medical Information, University of Poitiers, CHU Poitiers Hospital, F-86021 Poitiers, France
